* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Principles of Protein Structure
Point mutation wikipedia , lookup
Genetic code wikipedia , lookup
Signal transduction wikipedia , lookup
Gene expression wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Peptide synthesis wikipedia , lookup
Expression vector wikipedia , lookup
Magnesium transporter wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Bimolecular fluorescence complementation wikipedia , lookup
Protein purification wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Metalloprotein wikipedia , lookup
Biochemistry wikipedia , lookup
Structural alignment wikipedia , lookup
Homology modeling wikipedia , lookup
Western blot wikipedia , lookup
Interactome wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Principles of Protein
Structure
Amino Acids - Basic Building
Blocks of Proteins
Amino Acids Have
Molecular Chirality
Molecular Chirality
Proteins are Polymers
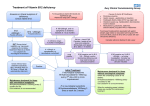
Cis Proline in the Active Site of
HCV NS2
Cis Pro 164
Cys
184
Glu 163
3.0 Å
4.1 Å
3.0 Å
3.2 Å
3.1 Å
3.1 Å
His 143
Leu 217
(C-term)
Nature 2006 vol. 442 (7104) pp. 831-5
Cis trans Proline Isomerase
• Cyclophilins are a family of proteins that
catalyze the isomerization of peptide
backbones at a proline
• Critically important for protein folding
• Cyclospronine is a potent
immunosuppressant and an inhibitor of
cyclophilins
• Commonly used after organ transplant
Four Levels of Protein
Structure
Four Levels of Protein Structure
• Primary, 1o!
– Amino acid sequence; Covalent bonds"
• Secondary, 2o!
– Local conformation of main-chain atoms (Φ and Ψ
angles); non-covalent interactions (h-bonds)"
• Tertiary, 3o!
– 3-D arrangement of all the atoms in space (mainchain and side-chain); non-covalent interactions"
• Quaternary, 4o!
– 3-D arrangement of subunit chains, non-covalent
interactions
Four Levels of Protein
Four Levels
of Protein Structure
Structure
• Primary, 1o "
TPEEKSAVTALWGKV"
• Secondary, 2o!
• Tertiary, 3o!
• Quaternary, 4o!
α2
α1
β2
β1
β2
Side Chain
Conformation
Protein Backbone Conformation
α Helices
α Helix
• If N-terminus is at bottom,
then all peptide N-H bonds
point “down” and all peptide
C=O bonds point “up”.
• N-H of residue n is H-bonded
to C=O of residue n+4.
• a-Helix has:
– 3.6 residues per turn
– Rise/residue = 1.5 Å
– Rise/turn = 5.4Å
Secondary Structure: Alpha Helices
Right handed
Left handed
310 and π Helices
310 helix H-bonds n and n+3
π helices H-bonds n and n+5
α Helix
• R-groups in α-helices:
– extend radially from the core,
– shown in helical wheel diagram.
– Can have varied distributions
Polar
Hydrophobic
Amphipathic
Secondary Structure: Beta Sheet
Antiparallel beta sheet
Parallel beta sheet
β Sheet
• Stabilized by H-bonds
between N-H & C=O
from adjacent
stretches of strands
• Peptide chains are fully
extended pleated
shape because
adjacent peptides
groups can’t be
coplanar.
β Sheet - 2 Orientations
Parallel
Not optimum Hbonds; less stable
Anti-parallel
Optimum H-bonds;
more stable
Beta Turn – 2 Conformations
Only Difference
Tertiary Structure
• Charge based interactions
– 62R:163E
– 55E:170R
• Hydrophobic interactions
– 189V
– 201L
– 213I
– 215L
– 266L
• Disulfide bond
– 203C:259C
1HSA
Peptide bound to
Class I MHC
Quaternary Structure
α2
α1
β2
β1
PDB File
Inter-atomic
Internal coordinates: bond lengths
distances:
(distance between bonded atoms)
Interatomic Distances
Atomic coordinates
a1 = (a1x, a1y, a1z)
a2 = (a2x, a2y, a2z)
Bond length
b12 = ((a2x–a1x )2+(a2y–a1y )2 +(a2z–a1z )2)1/2
a2
b12
a1
PDB file 3e7l
Bonded distances don’t change much:
Bond Lengths Remain Constant
lengths: Chain A backbone
1.459±0.003 Å
1.527±0.003 Å
1.329±0.001 Å
Bond or “valence”
angles:
Internal
coordinates: valence angles
(complement of angle formed by successive chemical bonds)
Bond Angles
Atomic coordinates
a1 = (a1x, a1y, a1z)
a2 = (a2x, a2y, a2z)
a3 = (a3x, a3y, a3z)
Bond vectors
b12 = (a2x–a1x, a2y–a1y, a2z–a1z)
b23 = (a3x–a2x, a3y–a2y, a3z–a2z)
Scalar product
b12"b23
= (a2x–a1x)(a3x–a2x)+(a2y–a1y)(a3y–a2y)+(a2z–a1z)(a3z–a2z)
a2
b12
a1
!123
b23
b12"b23 = b12b23 cos("–!123)
a3
cos!123 = –b12"b23/(b12b23)
cos("–!123) = cos" cos!123 + sin" sin!123 = – cos!123
Distribution of Backbone Angles
PDB file 3e7l
Distribution of backbone angles:
Valence angles: Chain A backbone
112.0±1.5°
116.4±0.5°
121.7±0.7°
Distribution of B-factors
Motifs, Topologies and Folds: β Sheet
The arrangement of secondary structure elements that give rise to a
folded entity
Motifs, Topologies and Folds: β Sheet
Motifs, Topologies and Folds: βαβ
Motifs, Topologies and Folds: α-helical
Motifs, Topologies and Folds: α-helical
C
N
Middle Domain of eIF4G - scaffold protein for translation initiation factors.
Protein Domains
• A folded unit of protein
• normally formed around a hydrophobic
core
• Tend to be globular
• 150-250 amino acids
Protein Domains
Many Eukaryotic proteins consist of multiple domains
•
Limited proteolysis can be used to experimentally
determine domain limits.
Limited Trypsin Digestion in the
Absence and Presence of dsRNA
Domain Swapping
Location of His 143, Glu 163, Cys 184
Glu
His
Cys
Cys
35Å
Glu
His
HIV Protease
Retroviral aspartic protease - dimerization forms a single active site
Structural Comparison
• Structural comparison requires a 3-Dimension
search
• Minimize the search by using just secondary
structural elements - Potential problem?
• DALI server
http://ekhidna.biocenter.helsinki.fi/dali_server/
• PDBe/fold
http://www.ebi.ac.uk/msd-srv/ssm/cgi-bin/
ssmserver
• VAST
http://www.ncbi.nlm.nih.gov/Structure/VAST/
SCOP:Structural Classification
Of Proteins
SCOP database classifies domains not entire proteins
CATH:Class, Architecture, Topology,
Homologous Superfamily
How to classify proteins with same general arrangement of
secondary structure but are connected differently?














































![pernicious%20anemia[1].](http://s1.studyres.com/store/data/000994580_1-27dd9ff09344a89075b2176563e85f6e-150x150.png)