* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Regulatory mechanisms that control T-follicular helper and
Lymphopoiesis wikipedia , lookup
Molecular mimicry wikipedia , lookup
Cancer immunotherapy wikipedia , lookup
Adaptive immune system wikipedia , lookup
Innate immune system wikipedia , lookup
Psychoneuroimmunology wikipedia , lookup
Polyclonal B cell response wikipedia , lookup
Immunosuppressive drug wikipedia , lookup
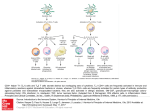
Immunology and Cell Biology (2014) 92, 34–39 & 2014 Australasian Society for Immunology Inc. All rights reserved 0818-9641/14 www.nature.com/icb REVIEW Regulatory mechanisms that control T-follicular helper and T-helper 1 cell flexibility Amy S Weinmann Following antigenic stimulation, CD4 þ T cells have the potential to differentiate into a number of specialized effector cell subtypes. To date, much progress has been made in defining the basic molecular mechanisms that regulate initial helper T-cell differentiation decisions. Emerging research in the field is now uncovering more complexity in the series of events that control helper T-cell commitment decisions than was previously appreciated. During the commitment process, helper T cells need to integrate both signals derived from the T-cell receptor and from the surrounding microenvironment. These external signals are then translated into internal changes in gene expression potential to ultimately define the functional characteristics of the cell. In this review, this topic will be discussed from the perspective of T-follicular helper (Tfh) and T-helper type 1 (Th1) cell differentiation. The focus will be on examining how the cytokine environment is perceived by signaling through signal transducer and activator of transcription (STAT) family proteins to initiate fate choices. The activities of STAT proteins are then in turn translated into changes in the molecular balance between B-cell lymphoma 6 (Bcl-6) and T-box expressed in T cells (T-bet), the helper T-cell lineage-specifying transcription factors that regulate Tfh and Th1 effector cell differentiation, respectively. Collectively, the knowledge of the molecular pathways that regulate Tfh and Th1 commitment have provided insight into the relationship between these two specialized helper T-cell subtypes and the potential for flexibility in their gene programs. Immunology and Cell Biology (2014) 92, 34–39; doi:10.1038/icb.2013.49; published online 1 October 2013 Keywords: Tfh; Th1; T-bet; Bcl-6; STAT; helper T cells In multicellular organisms, progenitor cells have the capacity to differentiate into specialized cell fates. Conserved lineage-specifying transcription factor families are required to precisely regulate gene expression program transitions to initiate unique cell fate choices.1 Developmentally controlled signaling events can tip the balance of these factors to promote commitment towards an individual cell fate choice at the expense of an alternative lineage.2–4 In the immune system, multipotential progenitor cell populations commit along individual fate pathways as environmental conditions signal the cell to change the composition of lineage-specifying transcription factors, ultimately resulting in the development of the full complement of the unique cell types that make up a functional immune system. The mechanistic principles that mediate developmental transitions continue to operate when lineage committed cells, such as CD4 þ T cells, further differentiate into functional effector fates. In contrast to the stable commitment pathways that mediate lineage differentiation decisions, emerging research suggests that the transitional events that promote more refined functional effector states retain a greater degree of flexibility than is observed with a true developmental decision.5–8 This is an especially important new area of research exploration in the field of CD4 þ T-cell differentiation, because the degrees to which specialized CD4 þ T-cell subtypes remain responsive to their microenvironment impacts how we approach immunotherapy and vaccination strategies.6,9 To date, much effort has been invested into defining the mechanisms that regulate the commitment of CD4 þ helper T cells into specialized functional subtypes.10–14 The emphasis of research efforts are now shifting towards uncovering the mechanistic reasons why these commitment events allow specialized CD4 þ T cells to remain flexible and respond to changing environmental conditions.7,15–17 In this review, this topic will be examined from the perspective of the molecular mechanisms that translate changing environmental conditions into specialized T-follicular helper (Tfh) and T-helper type 1 (Th1) gene programs and how these mechanisms provide insight into the potential for helper T-cell flexibility. ACTIVATION OF NAIVE CD4 þ HELPER T CELLS It is important to first highlight the type of events that create the milieu of regulatory factors that will be present in the cell and will have the potential to impact the commitment decision of naive CD4 þ helper T cells towards a specialized effector fate. The first event that occurs, which changes the composition of regulatory pathways activated in a cell, is the antigen-dependent engagement of the T-cell receptor (TCR) along with costimulation. Signaling through the TCR and costimulation activates a number of regulatory pathways, including the induction of NFAT and AP-1.18 Current research Department of Immunology, University of Washington, Seattle, WA, USA Correspondence: Dr AS Weinmann, Department of Immunology, University of Washington, Box 358059, 750 Republican Street, Seattle, WA 98195, USA. E-mail: [email protected] Received 12 August 2013; revised 28 August 2013; accepted 29 August 2013; published online 1 October 2013 Tfh and Th1 flexibility AS Weinmann 35 efforts are focused on defining how the TCR repertoire contributes to the differentiation potential of specialized helper T-cell subtypes. Recent studies have suggested that TCR intrinsic properties predispose the cell towards initiating either a Th1 or Tfh program.19,20 Interestingly, the strength of TCR signaling, as determined by the duration of TCR engagement with peptide–major histocompatibility complex II (MHCII) complexes, pushes a cell towards a fate choice, with strong TCR signaling promoting Tfh differentiation.19 Taken together, this means that a clonal population of cells is predisposed towards initiating a Th1 or Tfh program based on the characteristics of the TCR, but that the pathogen burden can also influence cellular differentiation as well. This is because antigen concentrations can change the dwell time between the peptide– MHCII complexes and the TCR, which will impact the strength of TCR signaling.19 Mechanistically, it is currently unclear which series of molecular events that occur downstream of TCR signaling are required to differentially induce the specialized effector cell programs. This will be an exciting new avenue of research to pursue in future studies. REGULATORY PATHWAYS CONTROLLED BY ENVIRONMENTAL CONDITIONS Following antigen-dependent activation, the differentiation of a CD4 þ T cell into an effector fate is next dependent on the environmental conditions the CD4 þ T cell encounters after activation. Most often, the encounter with a specific type of pathogen will cause the innate immune cells in the environment to initiate a somewhat prototypic cytokine response to the pathogen.21–23 For instance, the conserved components of bacteria and viruses (for example, pathogen-associated molecular patterns) are recognized by pattern recognition receptors, such as the Toll-like receptors, that cause innate immune cells to secrete a specific complement of cytokines and chemokines.21,24 The cytokine environment created by the innate immune response thus represents the first dominant microenvironment that influences CD4 þ T-cell effector differentiation. Therefore, depending on the characteristics of the pathogen, this initial microenvironment will usually induce a defined effector cell program designed to combat a specific type of pathogenic insult. However, the environmental milieu that a CD4 þ T cell is exposed to is more complex than merely accounting for the factors secreted by the initial ‘hard-wired’ response of innate immune cells to the invading pathogen, especially when considering the functional lifespan of effector and memory cells.25,26 For instance, the microenvironment can be influenced by competing factors such as another infection or a chronic condition that results in prolonged systemic changes in the environment. The temporal changes during the course of an immune response also influence specialized CD4 þ T cells. Collectively, this means that as long as the cells remain responsive to an environmental signal, the new microenvironments that the cells encounter throughout the course of an infection will have the potential to change the functional characteristics of the CD4 þ T cells. These are particularly interesting points to think about in the context of the potential for flexibility in the Tfh-like cells that are found in circulation versus germinal center (GC) Tfh cells located in the follicle, or in regards to effector versus memory Th1 cells circulating in different anatomical locations.27–30 REGULATING THE EXPRESSION PATTERNS OF LINEAGE-SPECIFYING TRANSCRIPTION FACTORS An important aspect of how specialized CD4 þ T-cell gene expression programs are established is through the regulatory activities of developmental lineage-specifying transcription factor families. Lineage-specifying transcription factors have evolved conserved mechanistic activities that promote the transition from one cell program to another. Many times these transcription factors utilize mechanistic activities that alter the epigenetic state of select target genes.31–33 In addition, they also possess the dual ability to either activate lineage-specific genes or, alternatively, repress genes involved in opposing cellular states.34,35 In true developmental lineages, the lineage-specifying transcription factors promote cellular transitions that are permanent, resulting in the development of a new cell fate.36 Previously, the paradigm in the field was that the differentiation of CD4 þ T cells into specialized subtypes was a stable developmental fate choice.37,38 However, experimental observations have discovered a much more complicated picture, uncovering complex and flexible gene expression patterns in specialized helper T cells that are inconsistent with a solely stable fate model.6,7,39 This concept will be highlighted by examining how changing environmental signals, mediated via signal transducer and activator of transcription (STAT) proteins, modulate the molecular balance between the opposing helper T-cell lineage-specifying transcription factors T-box expressed in T cell (T-bet) and B-cell lymphoma 6 (Bcl-6), and how this in turn regulates flexibility in the Th1 and Tfh gene programs. SENSING THE ENVIRONMENT THROUGH THE STAT FAMILY IN TFH AND TH1 DIFFERENTIATION Many of the cytokines responsible for the differentiation decisions that promote specialized CD4 þ T-cell effector subtypes modulate the activity of STAT proteins.40,41 This transcription factor family serves to translate the external cytokine environment into internal events that control many aspects of cellular differentiation, including regulating the expression of helper T-cell lineage-specifying transcription factors.41–43 The helper T-cell lineage-specifying transcription factors then orchestrate another series of events, often still in cooperation with STAT proteins, to create a predominantly specialized effector cell program. Detailing how the STAT family mechanistically regulates aspects of Tfh and Th1 gene programs powerfully illustrates the relationship between these two specialized subtypes and the events that contribute to the potential for flexibility. The complexity of how the STAT proteins translate the cytokine environment into functional changes in specialized CD4 þ T cells is reflected by the nature of the specific versus redundant pathways that upregulate their activities, and then also by the specificity and redundancy in the downstream gene expression programs that are regulated by each family member.40 This concept is quite evident in studies examining Tfh cell differentiation.44 Here a number of elegant studies have defined the roles for STAT1, STAT3 and STAT4 in positively regulating aspects of Tfh cell differentiation in response to interleukin-6 (IL-6), IL-21 and IL-12, as well as an inhibitory role for STAT5 in response to IL-2.44–48 These studies are clarifying the environmental conditions that promote Tfh differentiation and the role for STAT proteins in this process. It is now clear that there is functional redundancy between cytokine signaling pathways and the STAT family member activities that promote Tfh differentiation, leading to the concept that multiple signaling pathways have the potential to initiate a Tfh response.44 Another intriguing aspect of these studies is that the same STAT signaling pathways that promote Tfh differentiation are also involved in the development of alternative specialized CD4 þ helper T-cell subtypes in other situations.44,49 This means that the cellular context for STAT family member activity is important for defining the specialized CD4 þ T-cell gene expression program that will be established in response to an environmental signal. Immunology and Cell Biology Tfh and Th1 flexibility AS Weinmann 36 These concepts become evident when comparing the role for individual STAT proteins in Tfh versus Th1 differentiation, as well as in examining mouse versus human Tfh cells. Recently, several comprehensive reviews were published addressing how STAT proteins regulate Bcl-6 induction and Tfh differentiation.44,50,51 The focus here will be on highlighting the STAT-protein-regulated events that have the potential to influence aspects of either Tfh or Th1 differentiation. SENSING IL-6 AND INTERFERON-c THROUGH STAT1 The early research surrounding the STAT proteins focused on defining how individual STAT family members are activated by different cytokine microenvironments.49 This information was then integrated into models highlighting the potential for unique STAT proteins to predominantly drive the differentiation of individual specialized helper T-cell responses.49 Recent research has provided a much more complex view of this topic, with individual STAT family members having roles in what had previously been thought of as opposing helper T-cell fate decisions.44 An example of this is in the role for STAT1 in both Th1 and Tfh cell differentiation. It has long been recognized that the interferon-g (IFNg)-dependent upregulation of STAT1 induces the expression of the Th1 lineage-specifying transcription factor T-bet to promote the differentiation of the Th1 gene program.52–54 However, recent studies have revealed that IL-6 also activates STAT1, but in this context STAT1 promotes the induction of Bcl-6 and the development of a Tfh phenotype.44,47 Thus, STAT1 is able to integrate a set of diverse environmental signals to tip the balance of the cell towards either a Th1 or Tfh phenotype. One of the interesting questions the field now faces is defining mechanistically how STAT1 serves as a positive regulator for both the Th1 and Tfh gene programs. Current data indicate that STAT1 is able to promote the upregulation of either T-bet or Bcl-6 depending upon the cellular context.47,53 A plausible hypothesis for how STAT1 can achieve specificity in the differential regulation of T-bet or Bcl-6 is that unique IFNg- or IL-6-dependent signaling events will induce a specific composition of additional regulatory factors in the cell that influences the final outcome of STAT1 activity. In this scenario, one complement of factors induced in response to IFNg signaling will favor the STAT1-dependent enhancement of T-bet expression, whereas IL-6 signaling will favor a different complement of factors to promote Bcl-6 expression. It is plausible to envision the possibility for flexibility between the Th1 or Tfh pathways if different sets of environmental conditions change the expression patterns of the complementary factors present in the cell. In other words, given that STAT1 serves as a conserved determinant for both T-bet and Bcl6 expression, dynamically altering the composition of the other regulatory factors involved in selecting the lineage-specifying factor will influence the downstream decision between the expression of the Th1 and Tfh gene programs. IL-12 SIGNALING INITIATES THE STAT4-DEPENDENT REGULATION OF T-BET AND BCL-6 A series of studies in both mice and humans have shown that the IL12-dependent upregulation of STAT4 can promote the induction of both T-bet and Bcl-6.48,55–57 Of note, this is in contrast to the integration of different environmental signals (that is, IFNg or IL-6) mediated by STAT1. In mice, immediately following the activation of CD4 þ T cells, exposure to environmental IL-12 causes the simultaneous induction of T-bet and Bcl-6 in a STAT4-dependent manner.48 The co-expression of T-bet and Bcl-6 results in the dual expression of IFNg and IL-21 by CD4 þ T cells. Interestingly, T-bet eventually outcompetes Bcl-6 when IL-12 signaling persists, resulting Immunology and Cell Biology in the downregulation of Bcl-6 expression to low levels. This allows for the predominance of a Th1 gene program at the expense of the Tfh-like state.48 Although the resolution of the STAT4-dependent coexpression between T-bet and Bcl-6 favors T-bet in environments with robust IL-12 conditions, it is possible that pathogenic insults that induce a more blunted IL-12 response could have the capacity to favor Bcl-6 over T-bet. In addition, if there are unique environmental signaling events that contribute to the decision of whether STAT4 predominantly activates either T-bet or Bcl-6 expression, encountering variations in these environmental conditions will likely have the potential to create flexibility between the Th1 and Tfh gene expression profiles later in the lifespan of the cells. The studies associated with IL-12 signaling in human cells suggest an even more prominent role for IL-12 in Tfh potential, creating a more complicated picture concerning the dichotomy between Th1 and Tfh cells in the human setting.55–58 The current data indicate a role for the IL-12-dependent induction of STAT3 and STAT4 in promoting the sustained expression of IL-21 and Tfh cell differentiation in human cells.56,58 This is in contrast to the transient induction of IL-21-producing CD4 þ T cells in response to environmental IL-12 observed in the murine setting.48 In addition, data from patients with mutations in IL-12 receptor b1 have a pronounced defect in Tfh cell development, also supporting the conclusion that there is a substantial role for IL-12 signaling in the generation of Tfh cells in the human setting.55 Intriguingly, a high proportion of IL-21-producing cells also co-express IFNg, likely reflecting the prominent role for IL-12 signaling in both IL-21 and IFNg expression in the human setting.56 Together, these data raise the possibility that human Th1 and Tfh cells may have an even greater degree of interrelatedness and flexibility between their gene programs due to the overlapping role for IL-12–STAT4 signaling in both specialized effector cell outcomes.50 These findings also suggest that additional regulatory factors that are not yet defined will have a prominent role in influencing whether IL-12–STAT4 signaling will be translated into a more T-bet-dominated Th1 gene program or a more Bcl-6-centric Tfh gene program in human immune responses. Further knowledge about the mechanisms that occur downstream of IL-12 signaling to tip the balance between T-bet and Bcl-6 to favor either the Th1 or Tfh gene programs will provide new insights into the types of environmental changes that will have the potential to influence these outcomes. IL-21 SIGNALING ENHANCES TFH DIFFERENTIATION There are often feedback loops that reinforce gene expression programs to create more stability in their phenotypes.38 IL-21 is an important cytokine involved in the germinal center B-cell response, in part, through its role in creating a positive feedback loop promoting the GC response.59,60 Tfh cells secrete IL-21 and, in turn, IL-21 signaling in Tfh cells also promotes the expression of genes associated with the Tfh phenotype.61,62 IL-21 signaling in Tfh cells predominantly upregulates the activity of STAT3 and, to an extent, STAT1, which then promotes the induction of Tfh-associated genes.40,44,45 Intriguingly, both IL-6 and IL-21 promote the activation of similar STAT family members and both have a role in Tfh development.45 Functional redundancy has been a common theme in the studies examining Tfh differentiation, with a large degree of redundancy observed in both the cytokine milieu and in the STAT family members that translate these environmental signals into the induction of the Tfh gene program.45 It has been hypothesized that this redundancy is advantageous because of the critical Tfh and Th1 flexibility AS Weinmann 37 importance for the antibody response in controlling diverse pathogenic insults.44 IL-2 SIGNALING MODULATES THE BALANCE BETWEEN STAT3 AND STAT5 IN CD4 þ T-HELPER CELLS STAT3 and STAT5 have been shown to have opposing roles in regulating some classes of target genes.63 In the case of Bcl-6 expression, STAT3 has a role in activating Bcl-6 expression, while STAT5 appears to be inhibitory towards Bcl-6 expression.15,46,64 This is an important point because in cells maintained in Th1 polarizing conditions, changes in environmental IL-2 have been shown to modulate the relative association of STAT3 and STAT5 with the Bcl-6 promoter.15 Notably, several independent studies have shown that high levels of environmental IL-2, which promotes STAT5 activation, inhibits Bcl-6 expression and Tfh differentiation.15,46,65,66 Interestingly, effector and memory Th1 cells remain responsive to environmental IL-2 conditions to create a scenario in which IL-2sensitive Bcl-6 expression changes have the potential to regulate flexibility between the Th1- and Tfh-like gene programs.15 COMMONALITIES IN STAT FAMILY MEMBERS IN TH1 AND TFH DIFFERENTIATION The high degree of overlap between the STAT family members that have roles in Tfh and Th1 differentiation is striking (Figure 1). As just highlighted, recent research has shown that the STAT family members once thought to have unique roles in Th1 development are also involved in generating a Tfh gene program.44 This is quite evident when considering the well-established required roles for STAT1 and STAT4 in the induction and maintenance of T-bet expression to promote Th1 cell development.49 As discussed above, several recent studies now indicate that STAT1 and STAT4 also contribute to the Tfh gene program.47,48,56 At least part of this role appears to be in regulating Bcl-6 expression.44 In addition, it also appears that STAT4 directly contributes to the induction of a subset of Tfh signature genes.48 Thus, STAT1 and STAT4 have roles in both the Th1 and Tfh gene programs, with a prominent aspect of this in regulating T-bet and Bcl-6 expression. At present, it is not entirely clear how the context of activation influences the activity of STAT proteins and the overall decision within the cell to predominantly initiate a Th1 or Tfh gene program. IFN IL-6 STAT1 T-bet Th1 IL-21 IL-12 STAT3 STAT4 IL-2 STAT5 Bcl-6 Bcl-6 T-bet Bcl-6 Tfh Tfh Th1 Tfh Figure 1 Cytokine signaling pathways that regulate Th1 and Tfh differentiation. Regulatory pathways for each cytokine are highlighted with different colors and the relative strength of the pathway is indicated by the thickness of the arrow. The schematic illustrates that there is a significant degree of overlap between the cytokine and STAT signaling pathways that regulate both the Th1 and Tfh gene programs. The milieu of other regulatory proteins present in the cell will have a role in the commitment and stabilization of the effector cell phenotypes between Tfh and Th1 cells. In this line of thought, studies have shown that basic leucine zipper transcription factor ATF-like (Batf) and c-Maf are important for the differentiation of Tfh cells, whereas H2.0-like homeobox (Hlx) has a role in Th1 cell responses.67–71 Thus, the environmental context for the activation of these regulatory factors will influence the commitment decision. It will be important to determine what series of events are required for the expression of these, and other, regulatory factors, because this may represent one of the key selective events that define the phenotype of the cells. Another intriguing possibility is that the cellular context, or the receptor signaling strength leading to STAT protein activation, will contribute to the selection of the downstream Th1 or Tfh gene expression pathways that are activated in different circumstances. The quality of STAT protein activation may be influenced by either the exposure to different cytokines, or the relative strength of cytokine signaling, impacting the overall composition of STAT protein activity or family member selection. An example for how signaling strength controls diverse outcomes is with the graded regulation of Bcl-6 expression in Th1 cells exposed to variable environmental IL-2 conditions.15 Here, the strength of IL-2 signaling is translated through the relative induction of STAT3 versus STAT5 activity, which then determines the expression levels for Bcl-6 in Th1 cells. Therefore, changes in cytokine concentrations in the environment have the potential to alter the balance between competing STAT family members activities, which can then cause graded changes in lineage-specifying transcription factors. The relative competing activities between STAT3 and STAT5 are also likely to differentially regulate the expression of additional target genes important in establishing either the Tfh or Th1 phenotypes as well. MOLECULAR BALANCE BETWEEN T-BET AND BCL-6 IN TH1–TFH FLEXIBILITY Modulating the expression of lineage-specifying transcription factors, such as T-bet and Bcl-6, represents a powerful way to integrate realtime changes in the microenvironment, because these transcription factors have the capacity to alter the underlying gene expression program of the antigen-specific effector cells. In part, the nature of the interplay between T-bet and Bcl-6 regulates both the commitment and the potential for flexibility between specialized Th1 and Tfh subtypes. Mechanistically, T-bet and Bcl-6 can physically interact to form a complex in helper T cells.15,34 The properties of the interaction between T-bet and Bcl-6 create a scenario in which T-bet is able to control Bcl-6 activity when the balance of the two proteins favors T-bet expression. This is because the DNA-binding zinc fingers of Bcl6 are required for the interaction with T-bet, whereas, in contrast, the C-terminal domain of T-bet, but not its centrally located DNAbinding domain, is required for complex formation with Bcl-6.15 Thus, T-bet–Bcl-6 complex formation will mask the DNA-binding domain of Bcl-6, but the T-bet DNA-binding domain will still be exposed.15 This means that when the conditions in the cell favor T-bet–Bcl-6 complex formation, it will prevent Bcl-6 from accessing its own target genes. However, as the DNA-binding domain of T-bet remains available, T-bet can target the repressive capability of Bcl-6 to a subset of its target genes to downregulate the expression of genes involved in alternative helper T-cell fates.34,35 Mechanistically, this also means that T-bet activity is dominant over Bcl-6, especially when there is a higher ratio of T-bet to Bcl-6 in the cell. As noted above, both T-bet and Bcl-6 expression is controlled by the activities of similar STAT family members that are activated in response to changes in the Immunology and Cell Biology Tfh and Th1 flexibility AS Weinmann 38 cytokine microenvironment of the cell. Therefore, the molecular balance between T-bet and Bcl-6 can be fine tuned by changes in the composition of STAT protein activities to effectively sense the microenvironment and create new functional capabilities for the cell. CONFLICT OF INTEREST The author declares no conflict of interest. ACKNOWLEDGEMENTS INTEGRATING THE COMPONENTS THAT TRANSLATE THE ENVIRONMENT INTO TH1 AND TFH EFFECTOR STATES The emerging data in the field examining Th1 and Tfh differentiation is providing new insight into the concept of either defining specialized helper T-cell fates as stable lineages, akin to the T- and B-cell fates, versus viewing them from the opposite end of the spectrum as completely flexible cell populations that are defined by the environment. The physiological answer appears to lie somewhere in between these two extremes. Insight into this topic is being extrapolated from the recent series of studies highlighted in this review that have shown there is a similar composition of STAT family members, and in some cases the same cytokine environmental conditions, which are involved in regulating the expression of T-bet and Bcl-6. This raises the possibility that the underlying gene expression programs of Th1 and Tfh cells have a degree of relatedness in their origin and also will remain responsive to similar environmental perturbations. Therefore, the decision for the functional direction of the specialized helper T cell will involve the integration of the conserved STAT signaling events, along with the composition of other regulatory factors concomitantly activated in the cell, to alter the balance between T-bet and Bcl-6. Notably, the cell maintains the ability to dynamically respond to changes in the environment because of the nature of these regulatory mechanisms. One important question that arises from the recent findings surrounding flexibility is how these same mechanisms can also account for the stability in the specialized helper T-cell subtypes that is often observed in the natural setting of the immune response.72 Here it is helpful to view the stability versus flexibility of the cell as a gradient of possibilities that is determined by the underlying mechanisms that regulate these processes. For example, stability will be promoted by any mechanistic event that makes it more difficult for a cell to respond to changes in the microenvironment. This may include downregulating receptors for cytokines and chemokines, altering the composition of additional regulatory proteins that are found in the cell, or establishing repressive epigenetic states surrounding the genes that are important for alternative phenotypes.39,66,73 If many layers that promote stability are built up in the cell, this may effectively ensure a highly stable specialized functional subtype that becomes relatively impervious to environmental changes. In contrast, the less layers of stability that are embedded in the cellular phenotype, the more likely it is that the cell will remain highly dynamic to respond to changes in the microenvironment. Therefore, the observations characterizing both stability and flexibility in specialized helper T-cell phenotypes in different circumstances do not represent a contradiction, but rather likely represent the number of barriers that are found within the cell population at any given point in natural immune responses. As discussed in this review, much information on this topic is already available with regards to Tfh and Th1 cells, and this is guiding new concepts in this area of study. In moving forward in the field, it will be important to define more completely the mechanisms that create these gradients of stability/flexibility, especially in relationship to the Tfh and Th1 gene programs. In the context of Tfh cell activity, understanding these principles may help with designing more efficacious vaccination strategies by creating conditions that promote the generation of stable Tfh cells and more robust antibody responses. Immunology and Cell Biology I would like to thank Ken Oestreich and members of the Weinmann lab for helpful discussions. Research in the author’s laboratory is supported by grants from the NIAID (AI061061) and the American Cancer Society (RSG-09-04501-DDC). 1 Miller SA, Weinmann AS. Common themes emerge in the transcriptional control of T helper and developmental cell fate decisions regulated by the T-box, GATA and ROR families. Immunology 2009; 126: 306–315. 2 Naiche LA, Harrelson Z, Kelly RG, Papaioannou VE. T-box genes in vertebrate development. Annu Rev Genet 2005; 39: 219–239. 3 Beaulieu AM, Sant’Angelo DB. The BTB-ZF family of transcription factors: key regulators of lineage commitment and effector function development in the immune system. J Immunol 2011; 187: 2841–2847. 4 Siggs OM, Beutler B. The BTB-ZF transcription factors. Cell Cycle 2012; 11: 3358–3369. 5 Hirahara K, Vahedi G, Ghoreschi K, Yang XP, Nakayamada S, Kanno Y et al. Helper T-cell differentiation and plasticity: insights from epigenetics. Immunology 2011; 134: 235–245. 6 O’Shea JJ, Paul WE. Mechanisms underlying lineage commitment and plasticity of helper CD4 þ T cells. Science 2010; 327: 1098–1102. 7 Lu KT, Kanno Y, Cannons JL, Handon R, Bible P, Elkahloun AG et al. Functional and epigenetic studies reveal multistep differentiation and plasticity of in vitro-generated and in vivo-derived follicular T helper cells. Immunity 2011; 35: 622–632. 8 Zhou L, Chong MM, Littman DR. Plasticity of CD4 þ T cell lineage differentiation. Immunity 2009; 30: 646–655. 9 Streeck H, D’Souza MP, Littman DR, Crotty S. Harnessing CD4( þ ) T cell responses in HIV vaccine development. Nat Med 2013; 19: 143–149. 10 Zheng W, Flavell RA. The transcription factor GATA-3 is necessary and sufficient for Th2 cytokine gene expression in CD4 T cells. Cell 1997; 89: 587–596. 11 Szabo SJ, Kim ST, Costa GL, Zhang X, Fathman CG, Glimcher LH. A novel transcription factor, T-bet, directs Th1 lineage commitment. Cell 2000; 100: 655–669. 12 Zhou L, Lopes JE, Chong MM, Ivanov II, Min R, Victora GD et al. TGF-beta-induced Foxp3 inhibits T(H)17 cell differentiation by antagonizing RORgammat function. Nature 2008; 453: 236–240. 13 Yu D, Rao S, Tsai LM, Lee SK, He Y, Sutcliffe EL et al. The transcriptional repressor Bcl-6 directs T follicular helper cell lineage commitment. Immunity 2009; 31: 457–468. 14 Johnston RJ, Poholek AC, DiToro D, Yusuf I, Eto D, Barnett B et al. Bcl6 and Blimp-1 are reciprocal and antagonistic regulators of T follicular helper cell differentiation. Science 2009; 325: 1006–1010. 15 Oestreich KJ, Mohn SE, Weinmann AS. Molecular mechanisms that control the expression and activity of Bcl-6 in TH1 cells to regulate flexibility with a TFH-like gene profile. Nat Immunol 2012; 13: 405–411. 16 Oestreich KJ, Weinmann AS. Master regulators or lineage-specifying? Changing views on CD4( þ ) T cell transcription factors. Nat Rev Immunol 2012; 12: 799–804. 17 Hegazy AN, Peine M, Helmstetter C, Panse I, Frohlich A, Bergthaler A et al. Interferons direct Th2 cell reprogramming to generate a stable GATA-3( þ )T-bet( þ ) cell subset with combined Th2 and Th1 cell functions. Immunity 2010; 32: 116–128. 18 Muller MR, Rao A. NFAT, immunity and cancer: a transcription factor comes of age. Nat Rev Immunol 2010; 10: 645–656. 19 Tubo NJ, Pagan AJ, Taylor JJ, Nelson RW, Linehan JL, Ertelt JM et al. Single naive CD4 þ T cells from a diverse repertoire produce different effector cell types during infection. Cell 2013; 153: 785–796. 20 Fazilleau N, McHeyzer-Williams LJ, Rosen H, McHeyzer-Williams MG. The function of follicular helper T cells is regulated by the strength of T cell antigen receptor binding. Nat Immunol 2009; 10: 375–384. 21 Beutler B. Inferences, questions and possibilities in Toll-like receptor signalling. Nature 2004; 430: 257–263. 22 Keating SE, Baran M, Bowie AG. Cytosolic DNA sensors regulating type I interferon induction. Trends Immunol 2011; 32: 574–581. 23 Saenz SA, Noti M, Artis D. Innate immune cell populations function as initiators and effectors in Th2 cytokine responses. Trends Immunol 2010; 31: 407–413. 24 Iwasaki A, Medzhitov R. Regulation of adaptive immunity by the innate immune system. Science 2010; 327: 291–295. 25 Kaech SM, Cui W. Transcriptional control of effector and memory CD8 þ T cell differentiation. Nat Rev Immunol 2012; 12: 749–761. 26 Murphy KM, Stockinger B. Effector T cell plasticity: flexibility in the face of changing circumstances. Nat Immunol 2010; 11: 674–680. 27 Morita R, Schmitt N, Bentebibel SE, Ranganathan R, Bourdery L, Zurawski G et al. Human blood CXCR5( þ )CD4( þ ) T cells are counterparts of T follicular cells and contain specific subsets that differentially support antibody secretion. Immunity 2011; 34: 108–121. Tfh and Th1 flexibility AS Weinmann 39 28 Vinuesa CG, Cook MC. Blood relatives of follicular helper T cells. Immunity 2011; 34: 10–12. 29 Vinuesa CG, Cyster JG. How T cells earn the follicular rite of passage. Immunity 2011; 35: 671–680. 30 Mueller SN, Gebhardt T, Carbone FR, Heath WR. Memory T cell subsets, migration patterns, and tissue residence. Annu Rev Immunol 2013; 31: 137–161. 31 Miller SA, Huang AC, Miazgowicz MM, Brassil MM, Weinmann AS. Coordinated but physically separable interaction with H3K27-demethylase and H3K4-methyltransferase activities are required for T-box protein-mediated activation of developmental gene expression. Genes Dev 2008; 22: 2980–2993. 32 Miller SA, Mohn SE, Weinmann AS. Jmjd3 and UTX play a demethylase-independent role in chromatin remodeling to regulate T-box family member-dependent gene expression. Mol Cell 2010; 40: 594–605. 33 Basso K, Dalla-Favera R. Roles of BCL6 in normal and transformed germinal center B cells. Immunol Rev 2012; 247: 172–183. 34 Oestreich KJ, Huang AC, Weinmann AS. The lineage-defining factors T-bet and Bcl-6 collaborate to regulate Th1 gene expression patterns. J Exp Med 2011; 208: 1001–1013. 35 Oestreich KJ, Weinmann AS. T-bet employs diverse regulatory mechanisms to repress transcription. Trends Immunol 2012; 33: 78–83. 36 Rothenberg EV. T cell lineage commitment: identity and renunciation. J Immunol 2011; 186: 6649–6655. 37 Mosmann TR, Cherwinski H, Bond MW, Giedlin MA, Coffman RL. Two types of murine helper T cell clone. I. Definition according to profiles of lymphokine activities and secreted proteins. J Immunol 1986; 136: 2348–2357. 38 Yamane H, Paul WE. Memory CD4 þ T cells: fate determination, positive feedback and plasticity. Cell Mol Life Sci 2012; 69: 1577–1583. 39 Wei G, Wei L, Zhu J, Zang C, Hu-Li J, Yao Z et al. Global mapping of H3K4me3 and H3K27me3 reveals specificity and plasticity in lineage fate determination of differentiating CD4 þ T cells. Immunity 2009; 30: 155–167. 40 Rochman Y, Spolski R, Leonard WJ. New insights into the regulation of T cells by gamma(c) family cytokines. Nat Rev Immunol 2009; 9: 480–490. 41 O’Shea JJ, Plenge R. JAK and STAT signaling molecules in immunoregulation and immune-mediated disease. Immunity 2012; 36: 542–550. 42 O’Shea JJ, Lahesmaa R, Vahedi G, Laurence A, Kanno Y. Genomic views of STAT function in CD4 þ T helper cell differentiation. Nat Rev Immunol 2011; 11: 239–250. 43 Yamane H, Paul WE. Cytokines of the gamma(c) family control CD4 þ T cell differentiation and function. Nat Immunol 2012; 13: 1037–1044. 44 Choi YS, Yang JA, Crotty S. Dynamic regulation of Bcl6 in follicular helper CD4 T (Tfh) cells. Curr Opin Immunol 2013; 25: 366–372. 45 Eto D, Lao C, DiToro D, Barnett B, Escobar TC, Kageyama R et al. IL-21 and IL-6 are critical for different aspects of B cell immunity and redundantly induce optimal follicular helper CD4 T cell (Tfh) differentiation. PloS One 2011; 6: e17739. 46 Johnston RJ, Choi YS, Diamond JA, Yang JA, Crotty S. STAT5 is a potent negative regulator of TFH cell differentiation. J Exp Med 2012; 209: 243–250. 47 Choi YS, Eto D, Yang JA, Lao C, Crotty S. Cutting edge: STAT1 is required for IL-6mediated Bcl6 induction for early follicular helper cell differentiation. J Immunol 2013; 190: 3049–3053. 48 Nakayamada S, Kanno Y, Takahashi H, Jankovic D, Lu KT, Johnson TA et al. Early Th1 cell differentiation is marked by a Tfh cell-like transition. Immunity 2011; 35: 919–931. 49 Adamson AS, Collins K, Laurence A, O’Shea JJ. The current STATus of lymphocyte signaling: new roles for old players. Curr Opin Immunol 2009; 21: 161–166. 50 Cannons JL, Lu KT, Schwartzberg PL. T follicular helper cell diversity and plasticity. Trends Immunol 2013; 34: 200–207. 51 Liu X, Nurieva RI, Dong C. Transcriptional regulation of follicular T-helper (Tfh) cells. Immunol Rev 2013; 252: 139–145. 52 Lighvani AA, Frucht DM, Jankovic D, Yamane H, Aliberti J, Hissong BD et al. T-bet is rapidly induced by interferon-gamma in lymphoid and myeloid cells. Proc Natl Acad Sci USA 2001; 98: 15137–15142. 53 Afkarian M, Sedy JR, Yang J, Jacobson NG, Cereb N, Yang SY et al. T-bet is a STAT1induced regulator of IL-12R expression in naive CD4 þ T cells. Nat Immunol 2002; 3: 549–557. 54 Yang Y, Ochando JC, Bromberg JS, Ding Y. Identification of a distant T-bet enhancer responsive to IL-12/Stat4 and IFNgamma/Stat1 signals. Blood 2007; 110: 2494–2500. 55 Schmitt N, Bustamante J, Bourdery L, Bentebibel SE, Boisson-Dupuis S, Hamlin F et al. IL-12 receptor beta1 deficiency alters in vivo T follicular helper cell response in humans. Blood 2013; 121: 3375–3385. 56 Schmitt N, Morita R, Bourdery L, Bentebibel SE, Zurawski SM, Banchereau J et al. Human dendritic cells induce the differentiation of interleukin-21-producing T follicular helper-like cells through interleukin-12. Immunity 2009; 31: 158–169. 57 Ma CS, Suryani S, Avery DT, Chan A, Nanan R, Santner-Nanan B et al. Early commitment of naive human CD4( þ ) T cells to the T follicular helper (T(FH)) cell lineage is induced by IL-12. Immunol Cell Biol 2009; 87: 590–600. 58 Ma CS, Avery DT, Chan A, Batten M, Bustamante J, Boisson-Dupuis S et al. Functional STAT3 deficiency compromises the generation of human T follicular helper cells. Blood 2012; 119: 3997–4008. 59 Vinuesa CG, Linterman MA, Goodnow CC, Randall KL. T cells and follicular dendritic cells in germinal center B-cell formation and selection. Immunol Rev 2010; 237: 72–89. 60 Linterman MA, Beaton L, Yu D, Ramiscal RR, Srivastava M, Hogan JJ et al. IL-21 acts directly on B cells to regulate Bcl-6 expression and germinal center responses. J Exp Med 2010; 207: 353–363. 61 Luthje K, Kallies A, Shimohakamada Y, Belz GT, Light A, Tarlinton DM et al. The development and fate of follicular helper T cells defined by an IL-21 reporter mouse. Nat Immunol 2012; 13: 491–498. 62 Spolski R, Leonard WJ. IL-21 and T follicular helper cells. Int Immunol 2010; 22: 7–12. 63 Yang XP, Ghoreschi K, Steward-Tharp SM, Rodriguez-Canales J, Zhu J, Grainger JR et al. Opposing regulation of the locus encoding IL-17 through direct, reciprocal actions of STAT3 and STAT5. Nat Immunol 2011; 12: 247–254. 64 Reljic R, Wagner SD, Peakman LJ, Fearon DT. Suppression of signal transducer and activator of transcription 3-dependent B lymphocyte terminal differentiation by BCL-6. J Exp Med 2000; 192: 1841–1848. 65 Ballesteros-Tato A, Leon B, Graf BA, Moquin A, Adams PS, Lund FE et al. Interleukin-2 Inhibits germinal center formation by limiting T follicular helper cell differentiation. Immunity 2012; 36: 847–856. 66 Pepper M, Pagan AJ, Igyarto BZ, Taylor JJ, Jenkins MK. Opposing signals from the Bcl6 transcription factor and the interleukin-2 receptor generate T helper 1 central and effector memory cells. Immunity 2011; 35: 583–595. 67 Bauquet AT, Jin H, Paterson AM, Mitsdoerffer M, Ho IC, Sharpe AH et al. The costimulatory molecule ICOS regulates the expression of c-Maf and IL-21 in the development of follicular T helper cells and TH-17 cells. Nat Immunol 2009; 10: 167–175. 68 Kroenke MA, Eto D, Locci M, Cho M, Davidson T, Haddad EK et al. Bcl6 and Maf cooperate to instruct human follicular helper CD4 T cell differentiation. J Immunol 2012; 188: 3734–3744. 69 Mullen AC, Hutchins AS, High FA, Lee HW, Sykes KJ, Chodosh LA et al. Hlx is induced by and genetically interacts with T-bet to promote heritable T(H)1 gene induction. Nat Immunol 2002; 3: 652–658. 70 Crotty S. Follicular Helper CD4 T Cells (T(FH)). Annu Rev Immunol 2011; 29: 621–663. 71 Betz BC, Jordan-Williams KL, Wang C, Kang SG, Liao J, Logan MR et al. Batf coordinates multiple aspects of B and T cell function required for normal antibody responses. J Exp Med 2010; 207: 933–942. 72 Amsen D, Spilianakis CG, Flavell RA. How are T(H)1 and T(H)2 effector cells made? Curr Opin Immunol 2009; 21: 153–160. 73 Crotty S. The 1-1-1 fallacy. Immunol Rev 2012; 247: 133–142. Immunology and Cell Biology