* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Molybdenum cofactor-deficient mice resemble the phenotype of
Dominance (genetics) wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Human–animal hybrid wikipedia , lookup
Frameshift mutation wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Gene nomenclature wikipedia , lookup
Saethre–Chotzen syndrome wikipedia , lookup
Neuronal ceroid lipofuscinosis wikipedia , lookup
Gene therapy wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
History of genetic engineering wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
Designer baby wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Microevolution wikipedia , lookup
Point mutation wikipedia , lookup
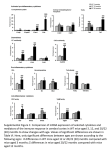
# 2002 Oxford University Press Human Molecular Genetics, 2002, Vol. 11, No. 26 3309–3317 Molybdenum cofactor-deficient mice resemble the phenotype of human patients Heon-Jin Lee1, Ibrahim M. Adham1, Günter Schwarz2, Matthias Kneussel3, Jörn O. Sass4, Wolfgang Engel1 and Jochen Reiss1,* 1 Institut für Humangenetik der Universität Göttingen, Göttingen, Germany, 2Institut für Pflanzenbiologie der Technischen Universität Braunschweig, Braunschweig, Germany, 3Max-Planck-Institut für Hirnforschung, Frankfurt, Germany and 4Zentrum für Kinderheilkunde und Jugendmedizin, Universitätsklinikum Freiburg, Freiburg, Germany Received August 22, 2002; Revised and Accepted October 17, 2002 Human molybdenum cofactor deficiency is a rare and devastating autosomal-recessive disease for which no therapy is known. The absence of active sulfite oxidase—a molybdenum cofactor-dependent enzyme— results in neonatal seizures and early childhood death. Most patients harbor mutations in the MOCS1 gene, whose murine homolog was disrupted by homologous recombination with a targeting vector. As in humans, heterozygous mice display no symptoms, but homozygous animals die between days 1 and 11 after birth. Biochemical analyis of these animals shows that molydopterin and active cofactor are undetectable. They do not possess any sulfite oxidase or xanthine dehydrogenase activity. No organ abnormalities were observed and the synaptic localization of inhibitory receptors, which was found to be disturbed in molybdenum cofactor deficient-mice with a Gephyrin mutation, appears normal. MOCS1/ mice could be a suitable animal model for biochemical and/or genetic therapy approaches. INTRODUCTION Molybdenum cofactor (MoCo) deficiency (MIM 252150) in human results in untreatable neonatal seizures and other neurological symptoms identical to those of sulfite oxidase deficiency (MIM 272300) (1). The biosynthetic pathway of MoCo has been discovered in humans and disease-causing mutations have been identified in the conserved genes MOCS1 (2,3) and MOCS2 (4,5) as well as in the gene for gephyrin (6), a protein that, besides its role in MoCo biosynthesis (7), has an additional function in neurotransmitter receptor clustering (8,9). Gephyrin-deficient mice resemble the phenotype of humans with hereditary MoCo deficiency and additionally display a failure in inhibitory neurotransmission (10). This latter defect prevents suckling and the homozygous animals die within the first day after birth. In co-cultivation experiments with fibroblasts from different MoCo-deficient patients two complementation groups (A and B) have been found (11). The majority of MoCo-deficient humans belong to complementation group A and harbor mutations in the MOCS1 gene (7). This bicistronic gene (2) encodes the two MoCo biosynthesis enzymes, MOCS1A and MOCS1B, in two consecutive open reading frames and their expression involves a conserved splicing pattern leading to a functional MOCS1A protein without the MOCS1B domain and activity and a fusion protein with MOCS1B activity and a non-functional MOCS1A domain (12,13). To create an animal model suitable for testing therapy approaches for the hitherto incurable MoCo deficiency, we have removed exon 3 of the murine MOCS1 gene by homologous recombination with a targeting vector. Homozygous / mice show a severe phenotype comparable to that of human MoCo-deficient patients. RESULTS Generation of MOCS1-deficient mice A murine MOCS1 cDNA was used to identify a genomic cosmid clone containing the murine MOCS1 gene. This revealed an exon/intron structure identical to that in the human genome (3). We chose to eliminate exon 3 of the murine MOCS1 gene by homologous recombination in embryonic stem cells, since it encodes several amino acid residues which are highly conserved in homologous proteins of mammals, plants and a variety of prokaryotes (2). Substitution of this exon by a neo-cassette should at least abolish the MOCS1A activity and—depending on the stability of the resulting mRNA—the MOCS1B activity could be affected as well (2,12,13). The targeting strategy is outlined in Figure 1. *To whom correspondence should be addressed at: Institut für Humangenetik der Universität Göttingen, Heinrich-Düker-Weg 12, D-37073 Göttingen, Germany. Tel: þ49 5513912926; Fax: þ49 551399303; Email: [email protected] 3310 Human Molecular Genetics, 2002, Vol. 11, No. 26 Figure 1. Disruption of the murine MOCS1 gene. (A) Targeting strategy. Wild-type MOCS1 gene (top), targeting vector (middle), and mutant locus (bottom) are shown. Sites of primers for PCR genotyping (arrows) and external probe for Southern blot (‘probe’) are indicated. (B) Southern blot analysis of ES cell clones. Genomic DNA extracted from ES clones was digested with HindIII and probed with the 50 external probe shown in panel (A). Homologous recombination events yielded a 9.4 kb hybridizing band detected in heterozygous (þ/) cell lines, while the wild-type (þ/þ) cell line showed a 8.7 kb band. (C) PCR genotyping of progeny. Wild-type (þ/þ), heterozygous (þ/) and homozygous (/) mice were identified by the amplication of PCR products specific for either the MOCS1 wild-type allele (650 bp) or the targeted allele (500 bp). As a disruption vector the plasmid pPNT (14) was used. This vector contains a neomycin phosphotransferase II resistance gene driven by a PGK promoter (pgk-neo), which should substitute for exon 3 of the MOCS1 gene and a herpes simplex virus thymidine kinase gene (tk) cassette outside the recombination region, which allows for positive and negative selection. Insertion of two genomic fragments (4.6 and 4.4 kb) resulted in the targeting vector (Fig. 1A), which was linearized with HindIII and transfected into RI ES cells (strain 129 Sv) (15). Drug-resistant clones were selected, from which DNA was isolated and screened by Southern blot analysis using a probe upstream of the targeting vector (Fig. 1B). This external probe detected the wild-type allele as an 8.7 kb fragment and the recombinant allele as a 9.4 kb fragment. One of two MOCS1þ/ ES clones was injected into C57BL/6J blastocytes, which gave rise to four male chimeric mice that transmitted the MOCS1 mutation in the germline. Chimeric mice were intercrossed to C57BL/6J and 129/Sv females, respectively, to establish the MOCS1-disrupted allele on a C57BL/6J 129/Sv hybrid and on a 129/Sv inbred genetic background. Heterozygous animals were identified by PCR analysis of DNA from mouse tails (Fig. 1A and C). In both backgrounds, male and female mice heterozygous for the MOCS1 mutation appeared normal and fertile. Heterozygous animals were mated and 25% of the offspring were homozygous for the mutant allele. Homozygous (/) animals appeared normal at birth and suckled. However, they failed to thrive (Fig. 2A and C) and died between days 1 and 11 after birth (Fig. 2B) with a mean life span of 7.5 days. MOCS1-deficient animals from the mixed background strain (129/Sv C57BL6J) showed no phenotypic differences as compared with those from the inbred line (129/ Sv) and were used for all subsequent analyses. Northern blot analysis with a 30 flanking probe (corresponding to exon 10) showed absence of MOCS1 RNA in the / animals, indicating that, owing due to the integration of the neo cassette, the expression of both the MOCS1A and the MOCS1B proteins is hampered (Fig. 3). Apart from their smaller size in comparison to their littermates and curly whiskers, the homozygous animals displayed no morphological abnormalities. Ataxia, convulsions and ectopic ocular lenses were not observed. Since loss of white matter has repeatedly been reported for MoCo-deficient human patients (5,16,17), nuclear staining of neuronal tissue was performed (Fig. 4). Hippocampus (Fig. 4A), spinal cord (Fig. 4B) and the cortex (Fig. 4C) of homozygous animals showed no macroscopic abnormalities in cell number or the development of cellular layers. For the MoCo-deficient Gephyrin knockout mouse, a disturbed postsynaptic clustering of inhibitory receptors has been reported (9). Immunostainings of MOCS1-deficient mice, in contrast, showed normal clusters for both Gephyrin and glycine receptors (Fig. 5). Biochemical analysis of MOCS1-deficient mice Molybdopterin was determined using the very sensitive nit-1 reconstitution assay. Figure 6A shows that molybdopterin (free Human Molecular Genetics, 2002, Vol. 11, No. 26 3311 Figure 2. MoCo-deficient mice. (A) MOCS1/ mouse (left) and healthy littermate (right) on day 8 after birth. (B) Survival of homozygous MoCo-deficient mice in days. No MOCS1/ mouse so far has survived beyond d11. (C) Weight development of þ/þ, þ/ and / mice during the first 5 days post partum. or in active cofactor form) in control animals is in the range 30–60 units/mg protein and virtually absent in the homoyzgous animals. Owing to the loss of MoCo all molybdenumdependant enzymes are inactive, as demonstrated for sulfite oxidase and xanthine dehydrogenase by direct measurement of enzyme activity from liver extracts (Fig. 6B and C). In control animals sulfite oxidase activity is increased in an agedependent manner. Because of the total loss of molybdopterin, 3312 Human Molecular Genetics, 2002, Vol. 11, No. 26 Figure 3. Northern blot analysis of liver RNA from the three genotypes. Hybridization with the murine MOCS1 exon 10 probe (top) revealed a 3.6 kb mRNA prominent in MOCS1þ/þ, reduced in MOCS1þ/, and absent in MOCS1/ mice. Rehybridization with human elongation factor (hEF, bottom) confirms equal amounts of mRNA. residual sulfite oxidase activities in homozygous mice are due to unspecific cytochrome c reductase activities in crude liver extracts (18). Semiquantitative determination of sulfite (Merckoquant) in fresh urine confirmed elevated sulfite levels in MOCS1/ animals as compared to wild-type and heterozygotes (data not shown). Amino acid analysis showed elevated taurin levels as compared to other amino acid concentrations in / mice. All urine samples revealed a signal coeluting with authentic sulfocystein reference compound, which was impaired by interfering components. This peak was higher in urine of / animals than in urine of wild-type or heterozygous animals. Cystin was clearly elevated in MoCo-deficient mice (Fig. 7A). The loss of xanthine dehydrogenase activity results in an elevation of the substrate xanthine and undetectable levels of the product uric acid in the urine (Fig. 7B and C). Wild-type as well as heterozygous mice show similar levels of xanthine oxidase educt and product. Hypoxanthine was undetectable in all samples. DISCUSSION In most MoCo-deficient patients severe mutations such as frameshift mutations or substitutions of conserved positions are found in the genes MOCS1, MOCS2 or Gephyrin (7). These mutations result in undetectable levels of molybdopterin, MoCo and sulfite oxidase activity with the consequence of considerable neurological damage and mental retardation. A few milder cases of isolated sulfite oxidase deficiency have been described (19,20), and Johnson et al. (5) have reported the first mutation in an unusual mild case of molybdenum cofactor deficiency. One of the two mutations identified in the MOCS2 gene apparently results in residual MoCo activity, thus leading to a milder phenotype. In this study, we have set out to create an animal model for the more common form of the disease, which is devastating and untreatable by any known therapy. We therefore eliminated a complete exon of the MOCS1 gene, removing a conserved domain of the MOCS1A ORF. Additionally, MOCS1B expression is hampered by the integration of the neo cassette. The presented data corroborate our assumption that thereby the synthesis of MoCo is completely abolished. As in humans, this results in a complete inactivation of the MoCo-dependant enzymes sulfite oxidase and xanthine dehydrogenase. It has been firmly established that sulfite oxidase deficiency alone is responsible for the severe neurological damages in MoCo deficiency (1). The phenotypic variance of the MoCo-deficient mice, as expressed in a lifespan of 1–11 days, is very similar to that observed in human patients, who show a survival of 1 week up to 10 years, but usually do not reach adolescence. The absence of any phenotypic differences between knockout mice with either a genetic background from the inbred strain 129/Sv or the mixed background C57BL/6J 129/Sv excludes any additional genetic elements in the mouse with a potentially compensating or modifying effect. As in humans, murine MoCo (MOCS1) deficiency is inherited as a monogenetic autosomal-recessive trait, for which our statistical data indicate complete penetrance. Various brain dysmorphologies have been described in MoCo-deficient patients and explained by both sulfite toxicity to the CNS and sulfate deficiency leading to neuronal loss (1). Because the here described MoCo-deficient animals die within their first days in life without any significant changes in CNS morphology, our observations put more emphasis on the critical role of sulfite toxicity. The precise mechanisms of this toxicity are not completely understood. The reaction of sulfite with disulfide bonds or with sulfhydryl groups would be a general process without pronounced sensitivity of the brain. Graf et al. (17) already pointed out that one effect of elevated sulfite levels might be the depletion of glutathione, which itself is necessary for cell membrane resistance to oxidative injury. Salman et al. (21) investigated a deceased MoCo-deficient patient in detail and concluded that the sulfite-induced toxic insult to the brain causes excitotoxic neuronal injury in the presence of excess magnesium. The connection of MoCo biosynthesis and postsynaptic receptor clustering, the two roles of the bifunctional protein Gepyhrin, remains elusive for the time being. One might speculate that these two biological processes interact with each other and that a MoCo deficiency other than Gephyrin deficiency also disturbs the postsynaptic architecture. We therefore examined the distribution of Gephyrin itself as well as that of the inhibitory glycine receptor in MOCS1-deficient mice. However, we did not find any abnormalities at the molecular level (Fig. 5) and these findings are in accordance with the absence of motor defects as described for the Gephyrin knockout mouse (10). The phenotype of the MoCo-deficient mice described in this study corresponds to that found in the majority of human patients, a MOCS1 mutation abolishing the formation of precursor Z. However, no morphological changes in brain tissue and no ectopic lenses were observed. These symptoms usually manifest after the neonatal period in human patients and a maximal life span of 11 days of the homozygous animals might be too short to develop such gross abnormalities. Although we did not observe any signs of convulsions or ataxia, it is difficult to exclude these symptoms in mice of such limited age. Clearly, further work is required here to determine the myelinization state of neurons. The homozygous animals truly reflect the biochemical abnormalities typical and diagnostic of MoCo deficiency, which finally result in premature death. These animals therefore can now be used for testing of therapeutical concepts for this hitherto incurable disease. Two options are available to this end. Human Molecular Genetics, 2002, Vol. 11, No. 26 3313 Figure 4. Nuclear staining of neuronal tissue derived from MOCS1A-deficient mice. The hippocampus (A), the spinal cord (B) and a cortical detail (C) are shown. As compared to wild-type tissue sections no macroscopic abnormalities in cell number or the development of cellular layers are detectable upon the loss of MOCS1A protein. One is the purification of precursor Z, which is the product of the MOCS1-encoded proteins and more stable than molybdopterin or active MoCo, and its delivery in different therapeutical regimes. The second option is gene therapy, delivering one or two types of cDNAs to various organs by the use of viral or plasmid vectors. Although these protocols in attempts to cure other diseases often lack a sufficient expression level to reverse or at least improve the clinical phenotype, the apparently very low level of expression of all MOCS genes (2,22) renders MoCo deficiency a promising candidate for this approach. A correction of the biochemical abnormalities of MoCo deficiency by these means will not necessarily translate into a reversal of neurological symptoms. The described murine model will therefore also be helpful to determine how early an effective treatment regime will have to start. MATERIALS AND METHODS Construction and transfection of the targeting vector A murine MOCS1 cDNA (GenBank AF214016) was isolated by reverse transcription followed by PCR amplification and used to screen a murine cosmid library (RZGPD). A genomic clone (MPMGc121C15173Q2) was identified and further characterized by restriction mapping and sequence analysis. A 4.6 kb XbaI fragment containing the 50 -flanking region of the MOCS1 gene exon 3 (including exon 2) was isolated and ligated with XbaI-digested pPNT vector (clone MOCS1/1). A 4.4 kb fragment containing a 30 -flanking region of the MOCS1-gene exon 3 (including exons 4–7) was amplified by high-fidelity PCR (Clontech), subcloned into pGEM-Teasy (Promega) and isolated again using two NotI sites flanking the cloning site. This 30 -flanking region was inserted into the NotI digested clone MOCS1/1 and the resulting targeting construct (Fig. 1A) was linearized with BamHI before transfection. Colonies resistant to G418 (400 mg/ml) and ganciclovir (GANC) (2 mM) were selected, genomic DNA extracted, digested with HindIII and blotted onto Hybond N membrane (Amersham). PCR genotyping Genomic DNA was extracted from mouse tails and PCR was carried out for 35 cycles using the following conditions: 45 s at 94 C, 45 s at 60 C and 75 s at 72 C. The following primers were used to discriminate wild-type and mutant alleles: 1(MOCS1 antisense), 50 -GGCAGAGGCTGTTCAACATGG-3; 2(MOCS1 sense), 50 -CTGGGTTCCTGTGCCATCTAG-3; 3(Pgk sense), 50 -TCTGAGCCCAGAAAGCGAAGG-30 . The amplification products were analysed on 1.5% agarose gels. A 500 bp fragment of the mutant allele was amplified with primers 1 and 3 whereas primers 1 and 2 amplified 650 bp wild-type product with template DNA from both heterozygous and wild-type animals (Fig. 1). 3314 Human Molecular Genetics, 2002, Vol. 11, No. 26 Figure 5. Postsynaptic immunoreactivities in spinal cord sections derived from wildtype (þ/þ) and MOCS1 knockout (/) mice. Immunostainings using antibodies specific for the postsynaptic clustering protein gephyrin (A–D) and the inhibitory glycine receptor (E and F) are shown. Sections derived from MOCS1deficient mice display normal gephyrin clusters in number and size as compared to wild-type sections (A and B). At higher magnification gephyrin immunoreactivity is detectable in dendritic neuronal processes of both genotypes (C and D). Glycine receptor immunoreactive hotspots are concentrated along spinal dendrites of both genotypes as shown in representative projections (E and F). Northern blot analysis Total RNA (40 mg) was extracted from mouse liver using ‘Total RNA Reagent’ according to a protocol provided by the manufacturer (Biomol). RNA samples were electrophoresed on 1% denaturing agarose gels containing formaldehyde (5%) and subsequently transferred onto nitrocellulose membrane (Hybond-C extra, Amersham). This filter was hybridized with 32 P-dCTP-labeled (3000 Ci/mmol) MOCS1 exon 10 fragment. To check for integrity and equal amounts of RNA, the filter was Human Molecular Genetics, 2002, Vol. 11, No. 26 3315 Figure 6. Biochemical analysis of MOCS1-deficient mice. (A) Molybdopterin content (including active molybdenum cofactor) of wild-type and MOCS1-deficient mice at various days after birth. (B) Sulfite oxidase activity of the same animals as investigated in (A). (C) In gel detection of xanthine dehydrogenase activity. As positive control (C) 0.75 mU of bovine xanthine oxidase were used. rehybridized with a human elongation factor-2 (hEF) cDNA probe (23). Histology Cryostat sections were fixed for 20 min in 4% (w/v) paraformaldehyde and stained with 0.1% (w/v) cresylviolet for 10 min to visualize nuclei. Sections were washed in water twice for 2 min followed by the sequential incubation in 70% (v/v), 90% (v/v), 100% (v/v), 100% (v/v) ethanol for 2 min each. For delipidation sections were then incubated in xylol twice for 2 min before being covered in entellan mounting solution (Merck). Stained sections were visualized using a Zeiss Axiovert microscope equipped with a cooled CCD camera and the imaging software MetaMorph (Visitron Systems). Immunochemistry and confocal microscopy Cryostat sections were fixed for 5 min in 4% (w/v) paraformaldehyde and processed for immunofluorescence as previously described (9). In brief, cells were permeabilized in 0.5% (v/v) Triton-X-100, blocked in 5% (v/v) goat serum for 20 min and processed for immunofluorescence. Gephyrin and glycine receptor were visualized using the monoclonal antibodies mAb7 and mAb4, respectively (24). As secondary antibodies Alexa 488/ 594 were used (Molecular Probes). Confocal microscopy was performed using a Leica TCS-SP confocal laser scanning microscope equipped with Leica TCS-NT image software. Molybdopterin analysis Neurospora crassa nit-1 extract was prepared as described (25) and desalted prior to use by gelfiltration using Nick columns 3316 Human Molecular Genetics, 2002, Vol. 11, No. 26 Figure 7. Metabolic analysis of MOCS1-deficient mice. (A) Cystin concentration of healthy (þ/þ), heterozygous (þ/) and homozygous (/) animals. Urine of at least three animals was collected on day 3 or 4 post partum and pooled. (B) Xanthine concentration in the urine of wild-type, heterozygous and / mice at day 5 post partum. (C) Uric acid levels in the urine of the same animals as investigated in (B). (Amersham/Pharmacia). Crude protein extracts from mouse liver were prepared using 2 vol nit-1 buffer (50 mM sodium phosphate, 200 mM NaCl, 5 mM EDTA, pH 7.2), sonication and centrifugation. According to the linear range of the assay 2 ml of different dilutions of liver extracts were co-incubated with 20 ml of nit-1 extract containing 5 mM sodium molybdate and 2 mM reduced gluthatione. Complementation was carried out overnight at 4 C under anaerobic conditions. After addition of 20 mM NADPH for 10 min, reconstituted NADPH-nitrate reductase activity was determined. One unit of molybdopterin activity is defined as reconstituted nit-1 nitrate reductase sufficient to produce an increase at 540 nm of 1.0 absorbance units per 20 min reaction time. Activity is expressed as units per mg protein of the crude extract. increase of 1.0 absorbance at 550 nm per min at 25 C. Activity is expressed as units per mg protein of the crude extract. Sulfite oxidase assay Metabolite analysis Sulfite oxidase activity was determined according to Johnson et al. (26). Crude protein extracts from mouse liver were prepared using one volume of 0.1 M Tris–HCl, 0.1 mM EDTA, pH 8.5 by sonication. Enzyme activity was assayed in a reaction volume of 300 ml containing 0.5 M Tris/HCl, 0.16 mM sodium deoxycholic acid, 2 mM potassium cyanide, 0.25 mg cytochrome c and 1 mM sodium sulfite. One unit sulfite oxidase activity is defined as enzyme activity needed to produce an Concentrations of the purines uric acid, xanthine and hypoxanthine in urine were determined by reversed-phase HPLC using a modification of a method described by Simmonds et al. (29). In brief, separation of purines was achieved by gradient elution on a Spherisorb 5 mm ODS-2 analytical column (250 4.6 mm; Phenomenex, Aschaffenburg, Germany) at a flow rate of 1.3 ml/min. Eluent A consisted of 40 mM ammonium acetate in HPLC grade water adjusted Xanthine dehydrogenase assay Xanthine dehydrogenase activity was determined based on the methods of Mendel and Müller (27) and Koshiba et al. (28). Crude protein extracts from mice liver were prepared by sonication in 3 vol 100 mM potassium phosphate, 2.5 mM EDTA, 5 mM dithiothreitol, pH 7.5. After centrifugation protein concentrations was determined and 280 mg protein were loaded on each lane of a 6% native polyacrylamide gel. Xanthine dehydrogenase activity was detected by activity staining using 300 mM hypoxanthine as substrate. As positive control 0.75 mU XO from cow buttermilk (Sigma) was used. Human Molecular Genetics, 2002, Vol. 11, No. 26 to pH 5.00 with acetic acid and eluent B of 80% methanol:10% acetonitrile:10% tetrahydrofuran (v/v/v). Urine was diluted with a 9-fold volume of eluent A. Fifty microliters of the diluted urine were directly injected into the HPLC system. Purines were quantitated based on peak area units obtained by UV detection at 254 nm using calibration with authentic standard compounds. Peak identity was confirmed by comparison of both the retention times and of the spectra obtained with a photodiode array detector between 220 and 320 nm. The determination of the creatinine concentration was based on the Jaffé reaction (30). Amino acid concentrations in urine were determined with a Biotronik LC 3000 aminoacid analyzer using two detection wavelenghts (440 and 570 nm). ACKNOWLEDGEMENTS This work was supported by the Deutsche Forschungsgemeinschaft (Re768/5-4). We thank Ina Bartnik, Sigrid GrossHardt, Tanja Otte, Ilona Paprotta, H. Riedesel, Monika Schindler and Stefan Wolf with his blue angels for excellent technical assistance. REFERENCES 1. Johnson, J.L. and Duran, M. (2001) Molybdenum cofactor deficiency and isolated sulfite oxidase deficiency. In Scriver, C.R., Beaudet, A.L., Sly, W.S. and Valle, D. (eds), The Metabolic and Molecular Bases of Inherited Disease. McGraw-Hill, New York, pp. 3163–3177. 2. Reiss, J., Cohen, N., Dorche, C., Mandel, H., Mendel, R.R., Stallmeyer, B., Zabot, M.T. and Dierks, T. (1998) Mutations in a polycistronic nuclear gene associated with molybdenum cofactor deficiency. Nat. Genet., 20, 51–53. 3. Reiss, J., Christensen, E., Kurlemann, G., Zabot, M.T. and Dorche, C (1998) Genomic structure and mutational spectrum of the bicistronic MOCS1 gene defective in molybdenum cofactor deficiency type A. Hum. Genet., 103, 639–644. 4. Reiss, J., Cohen, N., Stallmeyer, B., Mendel, R.R. and Dorche, C. (1999) Human molybdopterin synthase gene: genomic structure and mutations in molybdenum cofactor deficiency type B. Am. J. Hum. Genet., 64, 706–711. 5. Johnson, J.L., Coyne, K.E., Rajagopalan, K.V., Van Hove, J.L.K., Mackay, M., Pitt, J. and Boneh A. (2001) Molybdopterin synthase mutations in a mild case of molybdenum cofactor deficiency. Am. J. Med. Genet., 104, 169–173. 6. Reiss, J., Gross-Hardt, S., Christensen, E., Schmidt, P., Mendel, R.R. and Schwarz, G. (2001) A mutation in the gene for the neurotransmitter receptor-clustering protein Gephyrin causes a novel form of molybdenum cofactor deficiency. Am. J. Hum. Genet., 68, 208–213. 7. Reiss, J. (2000) Genetics of molybdenum cofactor deficiency. Hum. Genet., 106, 157–163. 8. Kirsch, J., Wolters, I., Triller, A. and Betz, H. (1993) Gephyrin antisense oligonucleotides prevent glycine receptor clustering in spinal neurons. Nature, 366, 745–748. 9. Kneussel, M., Brandstätter, J.H., Laube, B., Stahl, S., Müller, U. and Betz, H. (1999) Loss of postsynaptic GABAA receptor clustering in Gephyrin-deficient mice. J. Neurosci., 19, 9289–9297. 10. Feng, G., Tintrup, H., Kirsch, J., Nichol, M.C., Kuhse, J., Beth, H. and Sanes, J.R. (1998) Dual requirement for Gephyrin in glycine receptor clustering and molybdoenzyme activity. Science, 282, 1321–1324. 11. Johnson, J.L., Wuebbens, M.M., Mandell, R. and Shih, V.E. (1989) Molybdenum cofactor biosynthesis in humans. J. Clin. Invest., 83, 897–903. 3317 12. Gray, T.A. and Nicholls, R.D. (2000) Diverse splicing mechanisms fuse the evolutionary conserved MOCS1A and MOCS1B open reading frames. RNA, 6, 928–936. 13. Hänzelmann, P., Schwarz, G. and Mendel, R.R. (2002) Functionality of alternative splice forms of the first enzymes involved in human molybdenum cofactor deficiency. J. Biol. Chem., 277, 18303–18312. 14. Tybulewicz, V.L., Grawford, C.E., Jackson, P.K., Bronson, R.T. and Mulligan, R.C. (1991) Neonatal lethality and lymphopenia in mice with a homozygous disruption of the c-abl protooncogene. Cell, 65, 1153–1163. 15. Joyner, A.L. (ed.) (1993) Gene Targeting: a Practical Approach. IRL Press, Oxford. 16. Endres, W., Shin, Y.S., Günther, R., Ibel, H., Duran, M. and Wadman, S.K. (1988) Report on a new patient with combined deficiencies of sulphite oxidase and xanthine dehydrogenase due to molybdenum cofactor deficiency. Eur. J. Pediatr., 148, 246–249. 17. Graf, W.D., Oleinik, O.E., Jack, R.M., Weiss, A.H. and Johson, J.L. (1998) Ahomocysteinemia in molybdenum cofactor deficiency. Neurology, 51, 860–862. 18. Johnson, J.L. and Rajagopalan, K.V. (1976) Human sulfite oxidase deficiency. J. Clin. Invest., 58, 551–556. 19. Barbot, C., Martins, E., Vilarinho, L., Dorche, C. and Cardoso, M.L. (1995) A mild form of infantile isolated sulfite oxidase deficiency. Neuropediatrics, 26, 322–324. 20. Van der Klei-van Moorsel, J.M., Smit, L.M.E., Brockstedt, M., Jakobs, C., Dorche, C. and Duran, M. (1991) Infantile isolated sulfite oxidase deficiency: report of a case with negative sulfite test and normal sulphate excretion. Eur. J. Pediatr., 150, 196–197. 21. Salman, M.S., Ackerley, C., Senger, C. and Becker, L. (2002) New insights into the neuropathogenesis of molybdenum cofactor deficiency. Can. J. Neurol. Sci., 29, 91–96. 22. Stallmeyer, B., Drugeon, G., Reiss, J., Haenni, A.L. and Mendel, R.R. (1999) Human Molybdopterin synthase gene: identification of a bicistronic transcript with overlapping reading frames. Am. J. Hum. Genet., 64, 698–705. 23. Rapp, G., Klaudiny, J., Hagendorff, G., Luck, M.R. and Scheit, K.H. (1989) Complete sequence of the coding region of human elongation factor 2 (EF-2) by enzymatic amplication of cDNA form human ovarian granulose cells. Biol. Chem. Hoppe-Seyler, 370, 1071–1075. 24. Pfeiffer, F., Simler, R., Grenningloh, G. and Betz, H. (1984) Monoclonal antibodies and peptide mapping reveal structural similarities between the subunits of the glycine receptor of rat spinal cord. Proc. Natl Acad. Sci. USA, 81, 7224–7227. 25. Nason, A., Lee, K.Y., Pan, S.S., Ketchum, P.A., Lamberti, A. and DeVries, J. (1971) In vitro formation of assimilatory reduced nicotinamide adenine dinucleotide phosphate: nitrate reductase from a Neurospora mutant and a component of molybdenum-enzymes. Proc. Natl Acad. Sci. USA, 68, 3242–3246. 26. Johnson, J.L., Rajagopalan, K.V., Lanman, J.T., Schutgens, R.B., van Gennip, A.H., Sorensen, P. and Applegarth, D.A. (1991) Prenatal diagnosis of molybdenum cofactor deficiency by assay of sulphite oxidase activity in chorionic villus samples. J. Inherit. Metab. Dis., 14, 932–937. 27. Mendel, R.R. and Müller, A.J. (1976) A common genetic determinant of xanthine dehydrogenase and nitrate reductase in Nicotiana tabacum. Biochem. Physiol. Pflanzen, 70, 538–541. 28. Koshiba, T., Saito, E., Ono, N., Yamamoto, N. and Sato, M. (1996) Purification and properties of flavin-and molybdenum-containing aldehyde oxidase from coleoptyles of maize. Plant Physiol., 110, 781–789. 29. Simmonds, H.A., Duley, J.A. and Davies, P.M. (1991) Analysis of purines and pyrimidines in blood, urine, and other physiological fluids. In: Hommes F. A. (ed.), Techniques in Diagnostic Human Biochemical Genetics: A Laboratory Manual. Wiley-Liss, New York, pp. 397–424. 30. Jaffé, M. (1886) Ueber den Niederschlag, welchen Pikrinsäure in normalem Harn erzeugt und über eine neue Reaction des Kreatinins. Z. Physiol. Chem., 10, 391–400.