* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Cotyledon organogenesis - Journal of Experimental Botany
Gene therapy of the human retina wikipedia , lookup
Epigenetics in stem-cell differentiation wikipedia , lookup
Ridge (biology) wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Genetic engineering wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Genomic imprinting wikipedia , lookup
Minimal genome wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Genome evolution wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Genome (book) wikipedia , lookup
History of genetic engineering wikipedia , lookup
Gene expression programming wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Mir-92 microRNA precursor family wikipedia , lookup
Gene expression profiling wikipedia , lookup
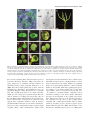
Journal of Experimental Botany, Vol. 59, No. 11, pp. 2917–2931, 2008 doi:10.1093/jxb/ern167 Advance Access publication 8 July, 2008 REVIEW ARTICLE Cotyledon organogenesis John W. Chandler* Department of Developmental Biology, University of Cologne, Gyrhofstrasse 17, D-50923 Cologne, Germany Received 15 April 2008; Revised 15 May 2008; Accepted 19 May 2008 Abstract The cotyledon represents one of the bases of classification within the plant kingdom, providing the name-giving difference between dicotyledonous and monocotyledonous plants. It is also a fundamental organ and there have been many reports of cotyledon mutants in many species. The use of these mutants where they have arisen in Arabidopsis has allowed us to unravel some of the complexities of embryonic patterning and cotyledon development with a high degree of resolution. The cloning of genes involved in cotyledon development from other species, together with physiological work, has supported the hypothesis that there exists a small number of orthologous gene hierarchies, particularly those involving auxin. The time is therefore appropriate for a summary of the regulation of cotyledon development gleaned from cotyledon mutants and regulatory pathways in the model species Arabidopsis and what can be inferred from cotyledon mutants in other species. There is an enormous variation in cotyledon form and development throughout the plant kingdom and this review focuses on debates about the phylogenetic relationship between mono- and dicotyledony, discusses gymnosperm cotyledon development and pleiocotyly in natural populations, and explores the limits of homology between cotyledons and leaves. Key words: Arabidopsis, auxin, cotyledon, dicots, evolution, monocots, natural variation, phylogeny. Introduction The patterning mechanisms responsible for generating any plant organ are complex and involve many gene hierarchies, and co-ordinated spatially and temporally regulated programmes of cell divisions, gene expression, and hormone function. The developmental regulation of leaf and root growth has been extensively described and reviewed (Fleming, 2005; Hochholdinger and Zimmermann, 2008), as have the patterning processes during embryogenesis responsible for establishing the meristems, mainly in Arabidopsis (Laux et al., 2004; Jenik and Barton, 2005; Jenik et al., 2007; Nawy et al., 2008), and maize (Vernoud et al., 2005). However, the development of the cotyledon as an organ has been neglected, although cotyledons are often the first aerial organ to differentiate and their initiation is inherent in embryonic patterning processes. With the explosion in the use of cotyledon mutants as tools for gene cloning (see Table 1 for a summary of cotyledon mutants in Arabidopsis and other species), many genetic pathways have been elucidated, which are usually included in reviews on embryonic patterning. However, one alternative consideration which provides a framework for thinking about cotyledon development, is the similarity between leaf and cotyledon development. Although dicot cotyledons and leaves have different functions in terms of storage and photosynthesis and morphological differences such as trichomes and stipules on leaves, on the basis of comparative morphology, they can be considered homologous organs according to the following criteria: the homologous apical positioning of both sets of organs on the shoot, the presence of meristems at their bases during primordium growth, similar organ expansion programmes and mature morphology, and that intermediate structures exist between cotyledons and leaves (quoted in Kaplan and Cooke, 1997). In addition to morphological homology, cotyledon and leaf initiation share significant signalling determinants, most notably resulting from the asymmetric distribution of auxin; in a similar way to the pre-patterning of cotyledon primordia outgrowth, leaf primordia initiate at positions of maximum auxin concentration/perception resulting from polar PIN1 localization (Reinhardt et al., 2003; Reinhardt, 2005; Heisler et al., 2005; Jönsson et al., 2006). One important difference * To whom correspondence should be addressed. E-mail: [email protected] ª The Author [2008]. Published by Oxford University Press [on behalf of the Society for Experimental Biology]. All rights reserved. For Permissions, please e-mail: [email protected] 2918 Chandler Table 1. Loci, mutations and redundant gene families affecting cotyledon organogenesis or pleiocotyly in various species Species Locus or mutant Cotyledon phenotype Reference Arabidopsis ALTERED MERISTEM PROGRAM1 ASYMMETRIC LEAVES1 DORNRÖSCHEN/DORNRÖSCHEN-LIKE EXTRA COTYLEDON1,2 FACKEL GURKE HYDRA1 laterne (pinoid, enhancer of pinoid) LEC/FUSCA Supernumery cotyledons Distorted cotyledons Fused cotyledons, variable cotyledon number Homeotic transformation of leaves into cotyledons Malformed cotyledons Cotyledons reduced or absent Pleiocotyly Cotyledons lacking Homeotic transformation of cotyledons into leaves Cotyledon fusion variable cotyledon number Single cotyledons Fused cotyledons, Pleiocotyly Partially fused cotyleons Temperature-dependent cotyledon defects Variable cotyledon number Chaudhury et al., 1993 Byrne et al., 2000 Chandler et al., 2007a Conway and Poethig, 1997 Schrick et al., 2000 Torres-Ruiz et al., 1996 Topping et al., 1997 Treml et al., 2005 Meinke et al., 1994; Keith et al., 1994 Furutani et al., 2007 MACCHI-BOU4 ENHANCER OF PINOID MONOPTEROS PINOID SHOOTMERISTEMLESS TOPLESS TWIN1 Antirrhinum Petunia Tomato Pea TIR family CUP-SHAPED COTYLEDON (CUC) family HD-ZIP Class III family KANADI/YABBY families Single cotyledons or absence Altered cotyledon differentiation PINFORMED1 family YUCCA family CONNATA CUPULIFORMIS PLEIOCOTYLEDONEA NO APICAL MERISTEM DEFECTIVE EMBRYO AND MERISTEM LANCEOLATE POLYCOTYLEDON Single and fused cotyledons Single cotyledons Cotyledon fusion Cotyledon fusion Pleiocotyly Partial cotyledon fusion Pleiocotyly Cotyledons sometimes lacking Pleiocotyly SINGLE COTYLEDON Cotyledon fusion Single cotyledons Fused cotyledons between cotyledon and leaf initiation resides in phyllotaxis: dicot cotyledons arise opposite each other, but leaves are subsequently initiated in many different phyllotactic patterns, however, the similarities provide an analogous framework for comparing the developmental genetic mechanisms in each case. With this analogy as a background, this review consolidates what is known about the initiation and outgrowth of cotyledons in the model eudicot Arabidopsis thaliana and surveys mutants known to affect cotyledon development in this and other species. The formal division of angiosperm species based on morphological characteristics into Dicotyledonae and Monocotyledonae classes was first created by John Ray in his Methodus Platarum Nova in 1682. Cotyledon number is one of the characteristics involved in this division, with the Monocotyldonae (monocots) having a single cotyledon and containing approximately 50 000 lineages and the Dicotyledonae comprising about 200 000 species. Similarly for leaves and flowers, the cotyledon as an organ displays huge variation in morphology and developmental programmes throughout the plant kingdom and the evolutionary origin of the cotyledon will also be Berleth and Jürgens, 1993 Bennet et al., 1995 Aida et al., 1999 Long et al., 2002 Vernon et al., 2001; Vernon and Meinke, 2004 Dharmasiri et al., 2005b Aida et al., 1997, 1999; Vroeman et al., 2003 Prigge et al., 2005 Izhaki and Bowman, 2007; Eshed et al., 2001; Siegfried et al., 1999 Friml et al., 2003 Cheng et al., 2007a Stubbe, 1966 Weir et al., 2004 Stubbe, 1966 Souer et al., 1996 Keddie et al., 1998 Avasarala et al., 1996 Al Hammadi et al., 2003; Madishetty et al., 2006 Liu et al., 1999 reviewed here, together with considerations of natural variation in cotyledon number and development. Cotyledon diversity In dicots, the cotyledons are lateral organs and the shoot apical meristem (SAM) is at a central position, whereas in monocots the cotyledon is terminal and the SAM at a lateral position. Cotyledons are formed during embryogenesis and are defined as either epigeal, if they emerge above ground, expand following germination and become photosynthetic, or hypogeal, if they remain below ground, do not expand, and remain non-photosynthetic. Cotyledons can be ephemeral structures, only persisting for a few days following emergence, or can be retained by the plant throughout its vegetative life. They may have petioles, or be sessile. For most dicot species, the number of cotyledons is invariate at two (Fig. 1A), however, exceptions include some species of Acer, Juglans, and Coffea which have three cotyledons (Eames, 1961; Duke, 1969) as well as Raphanus (Dube et al., 1981) and Sesamum (Pillai and Goyal, 1983), some members of the Ranales which have Cotyledon development and evolution 2919 Fig. 1. Variation in cotyledon morphology amongst the plant kingdom. A typical eudicot seedling; Raphanus sativus (A). A monocot seedling; a transverse section through a Zea mays (maize) kernel, showing the coleoptile (col), the scutellum, considered to be the single monocot cotyledon (sc) and the endosperm (end) (B). A gymnosperm seedling: Pinus sylvestris with five cotyledons (C). An anisocotyledonous plant; Monophyllaea horsfieldii showing the gradual development of unequally sized cotyledons: isocotyls 1 week after germination (D), the start of the expansion of a single cotyledon 2 weeks after germination (E), and obvious anisocotyly after 3 weeks’ growth (F). Arabidopsis cotyledons in wild type (G) and in the dornröschen (drn) mutant: a pseudo monocot (H), fused cotyledons (I), tricots (J), cup-shaped cotyledons (K), and a complete absence of cotyledon in the drn drnl double mutant (L). All scale bars represent 1 mm. three or four cotyledons (Datta, 1988) and some species of Persoonia (Protaceae) (Fletcher, 1909). Conversely, the genus Pinguicula is anomalous amongst eudicots for sometimes having a single cotyledon (Degtjareva et al., 2006) and some parasitic plants such as those from the Orobanchaceae, Santalales, and Rafflesiales have no cotyledons (Burgher, 1998). Monocots have a scutellum (Fig. 1B), considered by many to be a single cotyledon, which does not emerge above ground and can therefore only be shown in a seedling cross-section (Fig. 1B). As the scutellum remains underground it does not photosynthesize and therefore is not green, in contrast to most epigeal dicot cotyledons. Diversity exists in monocot cotyledon number, with some species such as Commelina, Dioscorea, Tamus, and Tinantia having a second vestigial cotyledon (Datta, 1988). Gymnosperms, distinct from angiosperm cotyledon classification, have variable cotyledon numbers between species, ranging from two, to 24 for Pinus maximartinezii; the largest number of cotyledons known for a plant (Farjon and Styles, 1997). Cotyledon number is also flexible within many gymnosperm species, for example, 5–7 for Pinus radiata and 7–13 for Pinus jeffreyi (Mirov, 1967), or 5 for Pinus sylvestris (Fig. 1C). In addition to cotyledon number, cotyledons can be classified according to size; dicots usually demonstrate isocotyly: the development of two equally sized cotyledons. Anisocotyly, however, is the prolonged growth of one cotyledon over the other, to give a large macrocotyledon and a small microcotyledon. This is demonstrated by Claytonia virginica (Cook), Capparis species (Franceschini and Tressens, 2004) and members of the Gesneriaceae including Streptocarpus rexii (Mantegazza 2920 Chandler et al., 2007) and Monophyllaea horsfieldii (Tsukaya, 1997; Fig. 1D–F), where the macrocotyledon develops from a basal meristem to resemble a large foliage leaf up to 30 cm long, with no additional leaves being initiated before floral induction. Syncotyly relates to partial or complete fusion of cotyledons, as exists in species of Calophyllum, Swietenia, Guarea, and Carapa, where cotyledons are distally fused (Datta, 1988). The corollary of syncotyly is schizocotyly, applied to cotyledon bifurcation or lobing which can result in supernumery cotyledons. Schizocotyly occurs in seedlings of the Chenopodiaceae, where one of two cotyledons may be cleft, to give hemitrocotyly, or one cotyledon of a tricotyledonous plant bifurcated to give a hemitetracotyl (Karschon, 1973). Various types of cotyledon syncotyly or pleiocotyly are demonstrated by some developmental mutants in Arabidopsis such as dornröschen and dornröschen-like, different alleles of which show pleiocotyly and syncotyly at low penetrance (Fig. 1G–L). The large diversity in cotyledon form and morphology throughout the plant kingdom argues against a single conserved ontogenetic programme. The evolution of cotyledony Cotyledons exist across the plant kingdom in gymnosperms, monocots, and eudicots. A distinction should be made here between dicots as a group, containing all plants with two cotyledons, where the term dicot is used as a form of morphology only, and the eudicots, a monophyletic phyllogenetic group containing most dicot species. The Angiosperm Phylogeny Group website has consolidated current views of land plant phylogeny (http://www.mobot.org/MOBOT/research/APweb/) and it is believed that the gymnosperms diverged about 385 MYA (Zimmer et al., 2007) from eudicot progenitors and the monophyletic monocot group about 200 MYA (Zimmer et al., 2007) (Fig. 2). There has been a longstanding debate as to the origin of monocotyledony, but since the dicot state exists across all eudicot groups, including the Gnetopsids, it is accepted to be ancestral to monocotyly (Dean, 2002), suggesting that the monocot state is derived (is an apomorphy). The exact position of the Gnetopsids is also a subject of debate, with the anthophyte hypothesis placing them as a sister group to the angiosperms, but most molecular evidence places them as a sister group to the conifers (Chaw et al., 2000; Fig. 2). Dicotyly in diverse groups could also be the result of independent coevolution and the debate as to whether mono- or dicotyly represents the more primitive developmental programme is ongoing, with several alternative theories: a single monocot cotyledon could have derived from the fusion of two cotyledons, or dicotyly might represent an evolutionary organ duplication or cotyledon splitting event from the monocot state. Alternatively, the Fig. 2. Schematic phylogeny of the plant kingdom, showing the branching of the monocots and gymnosperms from eudicot ancestors about 200 and 385 million years’ ago (MYA), respectively. The dicotyledonous synapomorphy and monocot and gymnosperm cotyledon apomorphies are represented by circles on the cladogram. The debated position of the Gnetales is also shown, according Chaw et al. (2000). monocot cotyledon state could have resulted from the loss or suppression of both cotyledons and subsequent novel apomorphic innovation, or by functional sub-specification, whereby one cotyledon became retarded to form the first plumular leaf and the other retained cotyledon position and function, i.e. heterocotyly. The taxonomic allocation into monocots or dicots has never been entirely clear for some species of paleoherbs such as Nymphaea (Tillich, 1990), which has a single cotyledon with two lobes, which could also be interpreted as two fused cotyledons. It has been suggested that Nymphaea is a monocot (Titova and Batygina, 1986), although molecular studies confirm a dicot association (Nickrent and Soltis, 1995). Alternatively, the cotyledon syncotyly of Nymphaea may represent a transitional cotyledon form. Some members of the Hydatellaceae have a sheathing structure which is similar to both a single monocot cotyledon and to the two fused cotyledons of Nymphaeales. The recent reclassification of the Hydatellaceae from the monocot order Poales to the Nymphaeales (Sokoloff et al., 2007) illustrates how unclear morphological characteristics defining the monocot–dicot categories can be and Hydatellaceae may demonstrate how a monocotyledonous condition could have arisen from the dicotyly present in water lilies. The phenomenon of dicotyledonous embryo formation in the monocot Agapanthus and the observation of stable transitional forms between mono- and dicotyledonous embryos supports the hypothesis of homology between monocot and dicot cotyledons, and Titova (2003) has advanced the hypothesis that, at least in this species, monocotyly derived from the dicotylous situation via congenital fusion of two cotyledons and subsequent suppression of the growth of one of them. The question whether dicotyly is plesiomorphic to monocotyly depends on the fundamental question of Cotyledon development and evolution 2921 homology between both, i.e. a continuous evolutionary origin should give rise to homologous structures (Burgher, 1998). Burgher claims that monocot and dicot cotyledons are not homologous for the following reasons: firstly, both sets of organs do not share equivalent positions, with the terminal cotyledon of monocots not equivalent to the lateral dicot organs. The second criterion is homology of structure; dicot cotyledons are morphologically distinct from succeeding leaves, whereas monocot cotyledons share homology with succeeding leaves. Assuming homology, intermediate or transitional structures should be found between both sets of cotyledons. Despite occasional instances of unstereotypic cotyledons amongst both monocots and dicots, these are considered by Burgher to be exceptions and the structures in question do not completely resemble the respective mono- or dicot ontogeny. Finally, an assumption of homology is that mono- and dicot cotyledons cannot co-occur in one plant at the same developmental stage. It has been interpreted that the third leaf in Nymphaea corresponds to the monocot cotyledon (references quoted in Burgher, 1998), and therefore, that dicot and monocot characteristics cooccur in Nymphaea. These lines of evidence against a shared homology between mono- and dicot cotyledons still leaves the question of the origin of both sets of organs unresolved. The situation is confused since dicot plants are not a monophyletic group (Soltis and Soltis, 2004) and not all monocots possess a single cotyledon (Tillich, 1995). With respect to homology between monocot and dicot cotyledons, there is debate as to whether the scutellum or coleoptile represents the monocot cotyledon. The grass scutellum and dicot cotyledon are functionally equivalent in being reserve lipid and protein storage organs, but to what extent the grass scutellum represents the whole cotyledon, part of it or an alternative structure is open (Vernoud et al., 2005). The coleoptile protects the plumule during germination and has also been considered by some to be equivalent to the cotyledon or part of it (Satoh et al., 1999). This debate is also related to whether the coleoptile is part of the scutellum, the first leaf, or a novel organ (reviewed in Vernoud et al., 2005) and is influenced by the observation that the development of the scutellum and coleoptile are genetically distinct (Satoh et al., 1999). The evolution of gymnosperm cotyledons is not clear, but they probably arose as a derived or apomorphic character via cotyledon duplication or schizocotyly within an ancestral angiosperm. Homology between cotyledons and leaves Despite developmental similarities between cotyledons and leaves, mutants exist which completely fail to develop cotyledons, but correctly initiate leaf primordia. These include several mutant combinations in Arabidopsis in genes involved in auxin biosynthesis or PIN1 expression, such as the laterne phenotype, caused by mutations in both PINOID and MACCHI-BOU 4/ENHANCER OF PINOID (Treml et al., 2005; Furutani et al., 2007) or DORNRÖSCHEN (DRN) and DORNRÖSCHEN-LIKE (DRNL) (Chandler et al., 2007), NON-PHOTOTROPHIC HYPOCOTYL3 and PINOID or a loss of YUCCA1, YUCCA4, and PINOID (Cheng et al., 2007a, b). In addition, some mutants fail to establish a SAM, such as stm in Arabidopsis, no apical meristem in Petunia (Souer et al., 1996), and leaf pattern mutants in pea (Marx, 1987), but produce normal cotyledons. This genetic uncoupling of cotyledon and leaf development demonstrates that they must each possess at least partly independent developmental programmes. Transformation of cotyledons into leaves and leaves into cotyledons Several mutants in Arabidopsis inform a discussion as to whether leaves and cotyledons are homologous structures or sui generis organs. These mutant phenotypes either demonstrate partial transformation of leaves into cotyledons or cotyledons into leaves, respectively. The LEAFY COTYLEDON (LEC) class of genes, LEC1 and LEC2, and FUSCA3 (FUS3) provide functions required for normal seed development. When mutated, cotyledons display features usually associated with vegetative leaves, such as trichomes, storage products, desiccation tolerance, and more complex vasculature (West et al., 1994; Meinke et al., 1994; Keith et al., 1994; Stone et al., 2001), and over-expression results in seedlings with embryonic characteristics (Stone et al., 2001). The lec cotyledon phenotype has been interpreted as representing partial homeosis (Meinke et al., 1994), or a disturbance in heterochrony (Keith et al., 1994), leading to an altered timing of developmental programmes. Similar to lec mutants, mutants in the HYDRA1 or HYDRA2/FACKEL genes in Arabidopsis have cotyledons with trichomes, thereby having a mixed phase phenotype (Topping et al., 1997; Jang et al., 2000; Schrick et al., 2000). Supplementation is defined as when the first true leaf is cotyledonary in character and three such mutations have been characterized in Arabidopsis (Conway and Poethig, 1997); extra cotyledon1 (xtc1), xtc2, and altered meristem program1 (amp1). These mutants produce leaves which phyllotactically develop perpendicular to the cotyledons, but possess partial or complete cotyledonary characteristics such as few or no trichomes, a simple venation pattern, and protein, lipid, and starch storage bodies. Some amp mutants show true polycotyly, with a true extra cotyledon being produced (Chaudhury et al., 1993), but the basis for the supplementation phenotypes is the precocious initiation of leaves during embryogenesis, and the mutations 2922 Chandler are therefore heterochronic, rather than homeotic. However, the simultaneous presence of cotyledon- and leafspecific characteristics in xtc1, xtc2, amp, and the lec mutants demonstrates that both cotyledons and leaves must at least share some common regulatory programmes. Phase change Phase-change from embryonic to vegetative development programmes has been little studied, but the question has been addressed in Brassica napus, which has non-dormant embryos (Fernandez, 1997). Precociously germinating embryos either produce extra cotyledons in a spiral phyllotaxy, possessing stipules, both leaf developmental characteristics, or chimeric organs with sectors of leaf tissue, characterized by trichomes and cotyledon tissue. Expression of molecular storage protein markers, normally restricted to the embryo, showed that the fate of a particular primordium was determined by the identity of the cells immediately surrounding it, implicating short range signalling pathways in primordium fate determination. As organ identity is related to the age of the embryo in Brassica napus, homeosis of cotyledons and leaves is a consequence of heterochrony, i.e. differentiation is dependent on temporal events in cellular environments. In anisocotylous plants such as Streptocarpus wendlandii, growth of the macrocotyledon is accompanied by trichome initiation in the basal region, which is characteristic for leaves (Nishii et al., 2004). Cytokinin application can convert the microcotyledon into a macrocotyledon in this species (Nishii et al., 2004), suggesting that endogenous cytokinin may play a role in cotyledon to leaf phase change. Do cotyledons arise from the SAM? If cotyledons and leaves share homology, the question arises whether they develop from the SAM or independently from it (Kaplan and Cooke, 1997; Barton and Poethig, 1993; Bowman and Eshed, 2000). If cotyledons originate from stem cell precursors in the SAM, what causes the phase-change switch to the initiation of leaf primordia in a different phyllotaxy to cotyledons? and if cotyledon development is independent of the SAM, what are the instructive signals determining cotyledon precursor cell fate? The SAM is usually defined in relation to its position between the cotyledons and, although in Arabidopsis, fully mature SAM histology becomes apparent only after cotyledon outgrowth, in some species, such as Pinus strobus, the SAM is visible before cotyledon initiation (Spurr, 1949). In addition, changes in gene expression domains which pre-pattern cotyledon outgrowth and SAM initiation in Arabidopsis are already established in the globular embryo, and clonal analyses cannot alone define the presence and activity of the SAM (Kaplan and Cooke, 1997). Long and Barton (1998) argue against the origin of both leaves and cotyledons from the SAM, on the basis that most cotyledon cells have never expressed STM, in contrast to those of leaf primordia, and that full SAM structure and activity is established following cotyledon outgrowth. In anisocotylous species such as Streptocarpus wendlandii (Nishii et al., 2004), the macrocotyledon enlarges by basal meristem activity, but as no leaves are subsequently produced, only an inflorescence, the meristem cannot be considered as a classical SAM. The question of the origin of the cotyledons can not be solved by histology alone, partly because of the huge developmental variation between species, and the danger of using Arabidopsis as a single paradigm, but only at the resolution of transcriptional cell profiling in single cells to identify the cell fate determinants of the cotyledon precursor cell(s). Cotyledon morphogenesis in Arabidopsis Cotyledon initiation and outgrowth The initiation and outgrowth of cotyledons in eudicots as exemplified by Arabidopsis is morphologically visible at the late globular stage and represents a symmetry-breaking event, taking the embryo from a radially symmetrical embryo to one having two planes of bilateral symmetry (Chandler et al., 2008b). This re-definition of symmetry creates two domains within the apical embryo region which are each characterized by diagnostic gene expression patterns: an apical central domain, not to be confused with the embryo central domain situated between apical and basal domains, and an apical peripheral domain, with a central-peripheral axis linking them (Fig. 3A). The differential expression of several genes in these domains leads to the establishment of the SAM in the central apical domain. A feature of cotyledon outgrowth is the necessity to maintain separate boundaries between the central embryo domain and the enlarging cotyledon primordium and between the two primordia and the role of the cupshaped cotyledon (CUC) genes is fundamental in this process: Arabidopsis possesses three redundant NAC domain transcription factors, CUC1, CUC2, and CUC3. Double cuc1 cuc2 and triple cuc1 cuc2 cuc3 mutants show complete cotyledon fusion to create cup-shaped cotyledons and seedlings lack a SAM, demonstrating that CUC gene functions are upstream of STM (Aida et al., 1999; Takada et al., 2001; Vroeman et al., 2003). The expression of CUC genes in the medial domain, between apical central and peripheral regions, in the heart stage embryo represses cell proliferation and differentiation in this domain, thereby creating a boundary which separates the developing cotyledons. In addition, CUC genes positively regulate STM expression in the central apical domain (Aida et al., 1999), where STM also represses Cotyledon development and evolution 2923 Fig. 3. Schematic overview of embryo domains and the genetic pathways involved in cotyledon initiation and outgrowth in Arabidopsis: a late globular/transition stage embryo at the point where the cotyledons begin to bulge out from the embryo flanks and the embryo changes from having radial symmetry to two planes of bilateral symmetry. The central apical domain is distinguished from the peripheral apical domain and the embryo apical, central, and basal domains are shown (A). Genetic interactions involved in cotyledon outgrowth (B). A heart stage embryo with cotyledon adaxial and abaxial sides distinguished (C). The genetic interactions involved in cotyledon polarization (D). CUC expression (Fig. 3B), leading to mutually exclusive expression domains (Aida et al., 1999). KNAT6 is also expressed at the boundary between the SAM and cotyledons and acts redundantly with STM in organ separation via CUC genes (Belles-Boix et al., 2006). The boundary of CUC gene expression is also regulated by miR164 (Laufs et al., 2004) and TCP transcription factors (Koyama et al., 2007) and the STM expression domain is also positioned by DRN/DRNL (Chandler et al., 2008a). A fundamental aspect of the transition phase of embryogenesis, when apical and basal organ fate is established, is the repression of root-promoting gene cascades in the apical embryo domain, which is provided by the activity of the TOPLESS (TPL) and TOPLESS-RELATED (TPR) proteins (Long et al., 2006), co-repressors which function together with AUX/IAA proteins in auxin-mediated transcriptional repression (Szemenyei et al., 2008). ASYMMETRIC LEAVES1 (AS1), a MYB domain transcription factor is required for normal cotyledon development and is involved in the control of cell differentiation. AS1 expression marks the sub-epidermal domain of the cotyledon initials and is later negatively regulated by STM in the developing SAM, leading to AS1 expression in a sub-epidermal domain of the developing cotyledons but is absent in the central domain (Byrne et al., 2000). Two redundant homeodomain proteins ATML1 and PDF2 which maintain the identity of L1 cells are necessary for cotyledon initiation (Abe et al., 2003), and the serine protease ALE1 and receptor like kinases ALE2 and ACR4 have overlapping roles in regulating protoderm-specific gene expression and cotyledon development (Tanaka et al., 2007). This provides some circumstantial evidence that appropriate protoderm differentiation in the embryo is a prerequisite for cotyledon formation. This is further supported by rpk1 toad2 double mutants, deficient in two receptor-like kinases, redundantly required for cotyledon development, which have defects in the central domain protoderm of globular embryos subjacent to cotyledon defects (Nodine et al., 2007; Nodine and Tax, 2008). How embryo central domain protoderm function is linked to cotyledon initiation is an open question, but it implicates an additional signalling mechanism involved in cotyledon initiation to those involving the apical central embryo domain. Establishing central and peripheral embryo domains and cotyledon differentiation Continuing the theme of the analogous development of cotyledons and leaves, polarity or sidedness is established in both organs with respect to the SAM, with the adaxial or upper side being that closest to the plant axis or SAM and deriving from the embryo apical central domain, and the abaxial side the opposite or underside generated by the peripheral embryo domain (Fig. 3C). The adaxial and abaxial surfaces of cotyledons are morphologically distinct with respect to cell size and stomatal density: the adaxial epidermis possesses cells of uniform size and has a flat surface with low stomatal density, whilst the abaxial surface has interspersed large and small cells, a rough surface and high stomatal density (Siegfried et al., 1999). This polarization during cotyledon differentiation, similarly to that of leaves, is established by three gene families: the class III HD-ZIP genes promote adaxial cell fate and KANADI and YABBY families promote abaxial cell fate (Fig. 3B). These roles are supported in each case by gene expression domains, loss-of-function and gain-offunction phenotypes. Of the five class III HD-ZIP gene family members, PHAVOLUTA (PHV) and PHABULOSA (PHB) contribute to a functional apical meristem and REVOLUTA (REV) towards vasculature development, but all are also responsible for specifying the adaxial domain of the leaves and cotyledons. PHV, PHB, and REV are expressed from the early globular stages, but as the cotyledon develops, its adaxial side derives from cells from the central domain, whilst the abaxial side results from cells from the peripheral domain and PHV and PHB transcripts localize to the adaxial side of the emerging and developing cotyledons (Emery et al., 2003; Prigge et al., 2005). The 2924 Chandler YABBY (YAB) genes FILAMENTOUS FLOWER and YAB3 are expressed from transition and heart stage embryos onwards, respectively, and subsequently at the abaxial side of the developing cotyledons, in cells derived from the peripheral domain (Sawa et al., 1999; Siegfried et al., 1999). Similarly, KAN1, KAN2, and KAN3, encoding GARP family plant-specific transcription factors (Eshed et al., 2001) are expressed in abaxial regions of developing embryos and cotyledons (Kerstetter et al., 2001; Eshed et al., 2004) and KAN4 expression is mainly ovule-specific (McAbee et al., 2006). Although the exact interaction between class III HD ZIP genes and KANADI/ YABBY genes is not clear, the mutually exclusive expression domains of class III HD-ZIP genes and YABBY and KANADI genes has led to models of mutual antagonism (Emery et al., 2003; Eshed et al., 2004, Fig. 3D). In addition, the boundaries of class III HD-ZIP gene expression are partly delineated by miRNA control (Williams et al., 2005) and via interaction with the class of LITTLE ZIPPER (ZPR) genes (Wenkel et al., 2007; Fig. 3D). Dominant gain-of-function alleles of PHV, PHB, and REV alleles cause the adaxialization of lateral organs (McConnell et al., 2001; Emery et al., 2003). Conversely, a triple phb phv rev mutant develops a single abaxialized cotyledon lacking bilateral symmetry (Emery et al., 2003), showing that the genes are necessary for adaxial cell fate and involved in hierarchies leading to the establishment of bilateral symmetry. Constitutive over-expression of KAN1, KAN2 or KAN3, or the YABBY genes FILAMENTOUS FLOWER1 (FIL) or YAB3 leads to constitutive wild-type abaxial characteristics in cotyledons (Eshed et al., 2001; Siegfried et al., 1999), suggesting that these genes promote abaxial cell fate. Constitutive expression of FIL or YAB3 (Siegfried et al., 1999) or the KAN genes (Eshed et al., 2001) also results in SAM arrest and together with the fact that phb-1d mutants have an enlarged meristem, shows that SAM formation correlates with alterations in expression boundaries of genes determining adaxial or abaxial cotyledon fate, and thereby, that SAM maintenance and cotyledon differentiation involves two-way signalling between them (Siegfried et al., 1999; Golz, 2006). Not all single mutant YABBY and KANADI genes display a mutant phenotype, but double fil yab3 mutants result in a loss of polarity in leaves, with a uniform mesophyll and the epidermis is comprised of mosaics of abaxial and adaxial tissue (Siegfried et al., 1999). Increasingly, severe polarity phenotypes are observed in higher order kan mutants (Eshed et al., 2004), such that kan1 kan 2 kan4 triple mutants exhibit radialized leaf-like outgrowths on the abaxial side of the developing cotyledons and hextuple kan1 kan2 kan4 fil yab3 yab5 mutants initiate three narrow cotyledons (Izhaki and Bowman, 2007). Polarity defects in higher order class III HD-ZIP or KANADI mutants (phb phv rev and kan1 kan 2 kan4) can be explained by an altered PIN1 expression pattern, leading to altered auxin maxima and, consequently, abnormal organ initiation (Izhaki and Bowman, 2007). The hormonal control of cotyledon development Auxin synthesis, transport and perception Auxin is a key regulator of cotyledon development as shown by mutant phenotypes of many of the loci in Table 1 which are involved with either auxin biosynthesis, polar transport or response, via the auxin receptor TIR1, or genes known to be up- or downstream of auxin response or sensitivity. In addition, many of these loci are members of redundant gene families, such as PINFORMED (PIN; Friml et al., 2003), YUCCA (Cheng et al., 2007a) and TIR (Dharmasiri et al., 2005b). Two lines of evidence demonstrate the importance of auxin synthesis or concentration in cotyledon organogenesis: the first is the use of the synthetic auxin-responsive DR5 promoter which has routinely been used to provide a read-out of auxin concentration (Ulmasov et al., 1997; Benkovà et al., 2003; Friml et al., 2003), although an absence of DR5 expression does not prove an absence of auxin maxima. DR5 expression concentrates in the tips of the incipient cotyledons in transition stage embryos and the tips of developing cotyledons (Cheng et al., 2007a; Fig. 4A) and correlates with an accumulation of IAA (Benkovà et al., 2003). Secondly, expression of the YUCCA1 and YUCCA4 genes, encoding flavin monooxygenase, a key enzyme in Trp-dependent auxin biosynthesis is strong in the central apical domain of the globular embryo and subsequently peaks in the tips of cotyledons (Cheng et al., 2007a). Auxin synthesized in the central apical domain is redirected by PIN1-mediated polar auxin transport (Friml et al., 2003). It has been known for many years that treatment of embryos with various polar auxin inhibitors inhibits cotyledon and SAM development in wheat (FischerIglesias et al., 2001), Brassica napus (Ramesar-Fortner and Yeung, 2001), Brassica juncea (Liu et al., 1993), Arabidopsis (Benkovà et al., 2003) and, more recently, in Norway Spruce (Larsson et al., 2008) and moss (Fujita et al., 2008). Auxin is transported in Arabidopsis embryos via the family of PIN auxin efflux transporters (Friml et al., 2003). PIN1 is polarly localized in the L1 layer of the developing embryo, such that the polarity correlates with auxin accumulation in cotyledon tips (Benkovà et al., 2003; Fig. 4B, C). Within the developing cotyledon, PIN1 becomes localized on the basal side of developing vasculature cells to direct auxin flow into the centre of the primordium and basipetally within the embryo (Benkovà et al., 2003). Thus polar auxin transport is responsible for creating auxin maxima necessary for Cotyledon development and evolution 2925 Fig. 4. Cotyledons develop at sites of auxin maxima: An Arabidopsis heart stage embryo with GFP expression under control of the DR5 auxinresponsive promoter, representing a maximum for auxin concentration, perception or response marking the tips of the developing cotyledons (A). PIN1–GFP expression under control of the PIN1 promoter in a transition stage (B) or heart stage Arabidopsis embryo (C), showing polar PIN1 localization to direct auxin flux in the L1 layer, towards the tip of the developing cotyledons. correct cotyledon specification. The activity of the PIN proteins is regulated by the phosphorylation state controlled by the PINOID serine/threonine protein kinase (Benjamins et al., 2001) and PP2A phosphatase (Michniewicz et al., 2007) and PIN1 polar localization is also regulated by ENHANCER OF PINOID/MACCHI BOU4 (Treml et al., 2005; Furutani et al., 2007). pin1 pid double mutants completely lack cotyledons and the embryos are characterized by an expansion in the expression domains of CUC1, CUC2, and STM, demonstrating that the negative regulation of the expression domains of these genes by PIN1 and PID is necessary for normal cotyledon initiation (Furutani et al., 2004). The auxin receptor has been identified as TIR1 (Kepinski and Leyser, 2005; Dharmasiri et al., 2005a), a member of a family of related AFB F-box proteins (Dharmasiri et al., 2005b). Auxin binds to a hydrophobic pocket within the TIR1 protein (Tan et al., 2007), thereby stabilizing the interaction between TIR1 and Aux/IAA proteins, targeting them for degradation and thereby relieving auxin response factor (ARF) transcription factors from transcriptional repression (Quint and Gray, 2006). Higher order mutants between TIR1 and AFB genes show cotyledon development defects (Dharmasiri et al., 2005b), demonstrating that TIR and AFB proteins regulate auxin responses during cotyledon development (Dharmasiri et al., 2005b). Arabidopsis possesses 23 ARFs (Guilfoyle and Hagen, 2007) which bind to auxin responsive elements (AREs) in target genes and lead to their activation or repression. The DRN promoter contains canonical AREs (Chandler et al., 2008a), providing further corroboration that auxin-controlled gene expression is relevant for cotyledon development. Cytokinins Some evidence implicates cytokinins in cotyledon development: in hybrid larch, exogeneous application of benzyladenine caused a decrease in cotyledon number (von Aderkas, 2002) and a monocotyledonous mutant of Catharanthus roseus could be restored to normal dicotyledony by the application of kinetin, suggesting that the mutant phenotype resulted from developmental-stagespecific cytokinin deficiency (Rai and Kumar, 2000). The Arabidopsis amp mutant has a pleiotropic phenotype, including polycotyly and considerably higher cytokinin levels than the wild type (Chaudury et al., 1993), showing that appropriate cytokinin levels play a role in normal cotyledon initiation. The DORNRÖSCHEN (DRN)/ ENHANCER OF SHOOT REGENERATION1 (ESR1) and DRN-LIKE/SUPPRESSOR OF PHYTOCHROMEB/ BOLITA/ESR2 genes have a role in cotyledon development, with mutants showing syncotyly and pleiocotyly (Ikeda et al., 2006; Chandler et al., 2007) and they have an additional role in cytokinin-independent shoot regeneration and are themselves regulated by cytokinins (Banno et al., 2001; Ikeda et al., 2006). Cotyledon development in other eudicots and monocots Although Arabidopsis has become the developmental paradigm for eudicot patterning processes, it must be reiterated that it is atypical amongst eudicot embryo development in having an inflexible hallmark cell division pattern. Embryo and cotyledon development in other species have not been extensively studied at the molecular level, only on a morphological descriptive basis (Wardlaw, 1955). Embryos of monocots and eudicots are basically morphologically indistinguishable at the early embryo stages and differences only become apparent at the globular stage when, in eudicots, the outgrowth of the cotyledons results in a heart-stage embryo and, in monocots, the embryo is more elongated as the cotyledon defines the primary axis (Burgher, 1998). Homologues and orthologues of several Arabidopsis genes involved in cotyledon organogenesis have been isolated from other species and show a conservation of function: CUPULIFORMIS (CUP) from Antirrhinum is a CUC orthologue 2926 Chandler (Weir et al., 2004), as is NO APICAL MERISTEM (NAM) in Petunia (Souer et al., 1996) and mutations in these genes lead to cotyledon phenotypes. In maize, ZmCUC3 is a CUC3 orthologue and ZmNAM1 and ZmNAM2 are CUC1/CUC2 orthologues (Zimmermann and Werr, 2005) and their expression pattern supports a function for ZmCUC3 in boundary specification and for ZmNAM1 and ZmNAM2 in SAM establishment, similar to CUC gene function in Arabidopsis. Demonstrating the orthologous function of Arabidopsis genes involved in cotyledon development in monocots is difficult, due to morphological differences. However, orthologues of STM exist in maize (Knotted1; Long et al., 1996) and rice (Sentoku et al., 1999) and the maize AS1 homologue is rough sheath2 (RS2; Timmermans et al., 1999; Tsiantis et al., 1999). barren inflorescence2 (bif2) in maize is an orthologue of Arabidopsis PINOID (McSteen et al., 2007) and bif2 homologues from other grass species and their comparison with PINOID shows that the sequence, expression, and function is conserved between monocots and eudicots. PIN1 orthologues have also been isolated from maize (Carraro et al., 2006), but as yet, not subject to mutagenesis. The DRN homologue in maize, ZmDRN is expressed in the prospective scutellum (Zimmermann and Werr, 2007), thereby showing an analogous expression pattern to that in Arabidopsis, where DRN expression prepatterns cotyledon development in the Arabidopsis globular embryo, and suggesting an orthologous function. There is, therefore, clear conservation of many transcription factor hierarchies controlling cotyledon development in Arabidopsis in divergent plant species, and involving auxin (Table 1). The function of these genes in monocot cotyledon development is often hard to elucidate, and even if they have no role in cotyledon development, their further characterization by heterologous complementation, together with the isolation of more heterologous genes will reveal the repertoire of ancestral pathways affecting cotyledon development and those which have sub-functionalized or diverged. An evolutionary developmental biology perspective should help to address this question. Cotyledon development in gymnosperms Studies on cotyledon development in gymnosperms are few and are mainly related to cotyledon number (Harrison and Von Aderkas, 2004) and limited to somatic embryos and morphological descriptions. However, a comparison of expressed sequence tags from loblolly pine embryos with those of Arabidopsis revealed a large conservation with angiosperm embryogenesis-related genes (Cairney et al., 2006). The variation in cotyledon number within a given gymnosperm species correlates with embryo size, which alters from year to year (Butts and Buchholz, 1940). There is a greater degree of variation in cotyledon number in somatic embryos compared with zygotic embryos. The ring-shaped initiation of multiple cotyledons in gymnosperms means that multiple axes of bilateral symmetry are established by cotyledon initiation and not just two axes as in eudicots. Pleiocotyly and natural variation in cotyledon number An abnormally large number of cotyledons within a species has been referred to in the literature as pleiocotyly or polycotyly. Pleiocotyly in natural populations has not been extensively studied, although widespread examples of spontaneous tricotyledony have been reviewed by Holtorp (1944) and Haskell (1954) in Populus (Rajora and Zsuffa, 1986), Prosopis cieraria (Nagesh et al., 2001a), Acacia mellifera (Nagesh et al., 2001b), Tectona grandis (Nagesh and Kardam, 2004), Sapindus emarginatus (Nagesh et al., 2004), and sunflower (Hu et al., 2005). The genetic basis for some of these examples is not known, but a tricotyledonous mutant of Catharanthus roseus was found to be controlled by a single recessive mutation (Rai and Kumar, 2001). In tomato, two independent loci causing polycotyledony were characterized: the polycotyledon (poc) mutation was mapped (Madishetty et al., 2006) and the mutant showed increased levels of polar auxin transport, suggesting that the cotyledon phenotype is a result of altered auxin distribution (Al-Hammadi et al., 2003). The defective embryo and meristems (dem) mutant also lacks a functional SAM and is caused by the mutation of a gene encoding a novel protein of unknown function (Keddie et al., 1998). In pea, three independent single cotyledon (sic) loci when mutated, give rise to single or fused cotyledons (Liu et al., 1999). The frequency of pleiocotyly has been studied as a complex trait in wild radish populations: Raphanus raphanistrum and Raphanus sativus show either more or fewer than two cotyledons at a frequency of between 0% and 5% (Conner and Agrawal, 2005). Paternal half-sibling families were compared to assess whether the low phenotypic penetrance was due to stabilizing selection or a lack of genetic variance for the trait. These alternatives could not be distinguished due to the small samples and low number of phenotypic plants. However, artificial selection has shown that genetic variance for pleiocotyly is not lacking: the frequency of tricotyledonous seedlings resulting from a cross between wild and cultivated sunflower lines increased from 2% in the F2 generation, to about 50% in the F5 generation (Hu et al., 2005), showing that additive genetic variation for polycotyly exists. Similarly, artificial selection for pleiocotyly in Antirrhinum majus was able to increase its frequency Cotyledon development and evolution 2927 from 5% to 64% (Haskell, 1954), which suggests that lack of genetic variation is not the reason for the usually invariant number of cotyledons. The variable numbers of cotyledons in gymnosperms has been addressed in clonal populations of somatic embryos of hybrid larch and was found to correlate directly with the diameter of the apical embryo surface (Harrison and Von Aderkas, 2004). Breeding for cotyledon number information to evolutionary developmental biology will increasingly reveal the extent of this conservation across the plant kingdom and distinguish pathways which have divergently evolved. It will also inform the debate on the evolution of cotyledon states. The similarities in the genetic regulation and auxindependent initiation and outgrowth of cotyledons and leaves demonstrates the homology between the structures. However, open questions such as what distinguishes leaf founder cells from cotyledon precursor cells and how cotyledon and leaf development differ, including determinants of phase change and the relationship of cotyledon initiation to the SAM, remain tantalizing foci of research. Natural variation in cotyledon number is underresearched and understanding the genetic basis of pleiocotyly could enable breeding agronomically important seed oil crops for increased productivity based on this trait. The possibility exists that additive variance in natural populations of some plants could allow the selection of pleiocotyly as a trait, although the potential adaptive value of variable cotyledon numbers has not been well studied and is dependent on understanding the genetic basis of natural variation in cotyledon number. The possible agricultural benefits of pleiocotyly or manipulating cotyledon size are greater yield and quality of oils in oil seed plants such as rape or sunflower, where oil is stored in the cotyledons rather than the endosperm. Additional cotyledons may even confer an enhancement in seedling establishment; a tricotyledonous sunflower mutation had no detrimental effect on growth and development and led to three leaves per node, increasing photosynthetic surface area and therefore potential productivity (Hu et al., 2005). JWC thanks Melanie Cole for PIN1 and DR5 Arabidopsis embryo pictures, Judith Nardmann for maize embryo photos, Siegfried Werth for photography, Klaus Menrath and Jürgen Hintzsche for providing plant material, and Wolfgang Werr for critical reading of this manuscript. JWC also gratefully acknowledges funding for this work from the Deutsche Forschungsgemeinschaft via SFB572. Conclusions References The Arabidopsis developmental paradigm has elucidated many gene hierarchies regulating cotyledon development and has demonstrated the key involvement of auxin. A further feature of the control of cotyledon development is redundancy amongst gene families such as the PIN (Friml, 2003), class III HD-ZIP (Prigge et al., 2005), YUCCA (Cheng et al., 2007a) or TIR/AFB (Dharmasiri et al., 2005b). Redundancy also exists between transcription factor pairs such as DRN/DRNL (Chandler et al., 2007) and TOAD2/RPK1 (Nodine et al., 2007). This genetic robustness and plethora of parallel pathways presumably reflects evolutionary adaptation to ensure buffering against negative mutations at a critical point in plant survival, thereby ensuring that the embryonic patterning programme proceeds successfully. It will be interesting to discover how conserved this redundancy is amongst other eudicots for which Arabidopsis is not representative. Despite the enormous diversity of cotyledon morphology and development within and between the monocots and eudicots, the existence of many cotyledon development genes and regulatory pathways are conserved, such as those involving CUC genes, STM, pathways involving auxin, although in many cases no monocot mutants are available to demonstrate how they affect cotyledon development. The increased ease of isolating gene homologues by heterologous cloning and applying the Abe M, Katsumata H, Komeda Y, Takahashi T. 2003. Regulation of shoot epidermal cell differentiation by a pair of homeodomain proteins in Arabidopsis. Development 130, 635– 643. Aida M, Ishida T, Tasaka M. 1999. Shoot apical meristem and cotyledon formation during Arabidopsis embryogenesis: interaction among the CUP-SHAPED COTYLEDON and SHOOTMERISTEMLESS genes. Development 126, 1563–1570. Aida M, Ishida T, Fukaki H, Fujisawa H, Tasaka M. 1997. Genes involved in organ separation in Arabidopsis: an analysis of the cup-shaped cotyledon mutant. The Plant Cell 14, 841–857. Al-Hammadi ASA, Sreelakshmi Y, Negi S, Seddiqi I, Sharma R. 2003. The polycotyledon mutant of tomato shows enhanced polar auxin transport. Plant Physiology 133, 113–125. Avasarala S, Yang J, Caruso JL. 1996. Production of phenocopies of the lanceolate mutant in tomato using polar auxin transport inhibitors. Journal of Experimental Botany 47, 709–712. Banno H, Ikeda Y, Niu QW, Chua NH. 2001. Overexpression of Arabidopsis ESR1 induces initiation of shoot regeneration. The Plant Cell 13, 2609–2618. Barton MK, Poethig RS. 1993. Formation of the shoot apical meristem in Arabidopsis thaliana: an analysis of development in the wild type and in the shoot meristemless mutant. Development 119, 823–831. Belles-Boix E, Hamant O, Witiak SM, Morin H, Traas J, Pautot V. 2006. KNAT6: an Arabidopsis homeobox gene involved in meristem activity and organ separation. The Plant Cell 18, 1900–1907. Benjamins R, Quint A, Weijers D, Hooykaas P, Offringa R. 2001. The PINOID protein kinase regulates organ development in Acknowledgements 2928 Chandler Arabidopsis by enhancing polar auxin transport. Development 128, 4057–4067. Benkovà E, Michniewicz M, Sauer M, Teichmann T, Seifertovà D, Jürgens G, Friml J. 2003. Local, effluxdependent auxin gradients as a common module for plant organ formation. Cell 115, 591–602. Bennet SRM, Alvarez J, Bossinger G, Smyth D. 1995. Morphogenesis in pinoid mutants of Arabidopsis thaliana. The Plant Journal 8, 505–520. Berleth T, Jürgens G. 1993. The role of the MONOPTEROS gene in organising the basal body region of the Arabidopsis embryo. Development 118, 575–587. Bowman JL, Eshed Y. 2000. Formation and maintenance of the shoot apical meristem. Trends in Plant Science 5, 110–115. Burger WC. 1998. The question of cotyledon homology in angiosperms. The Botanical Review 64, 356–371. Butts D, Buchholz JT. 1940. Cotyledon numbers in conifers. Transactions of the Illinois Academy of Science 33, 58–62. Byrne ME, Barley R, Curtis M, Arroyo JM, Dunham M, Hudson A, Martienssen RA. 2000. Asymmetric leaves1 mediates leaf pattering and stem cell function in Arabidopsis. Nature 408, 967–971. Cairney J, Zheng L, Cowels A, et al. 2006. Expressed sequence tags from loblolly pine embryos reveal similarities with angiosperm embryogenesis. Plant Molecular Biology 62, 485–501. Carraro N, Forestan C, Canova S, Traas J, Varotto S. 2006. ZmPIN1a and ZmPIN1b encode two novel putative candidates for polar auxin transport and plant architecture of maize. Plant Physiology 142, 254–264. Chandler JW, Cole M, Flier A, Grewe B, Werr W. 2007. The AP2 transcription factors DORNRÖSCHEN and DORNRÖSCHEN-LIKE redundantly control Arabidopsis embryo patterning via interaction with PHAVOLUTA. Development 134, 1653–1662. Chandler JW, Cole M, Werr W. 2008a. The role of DORNROESCHEN (DRN) and DRN-LIKE (DRNL) in Arabidopsis embryonic patterning. Plant Signaling and Behavior 3, 49–51. Chandler JW, Cole M, Werr W. 2008b. Plant development revolves around axes. Trends in Plant Science 13, 78–84. Chaudhury AM, Letham S, Craig S, Dennis E. 1993. amp1: a mutant with high cytokinin levels and altered embryonic pattern, faster vegetative growth, constitutive photomorphogenesis and precocious flowering. The Plant Journal 4, 907–916. Chaw S-M, Parkinson CL, Cheng Y, Vincent TM, Palmer JD. 2000. Seed plant phylogeney inferred from all three plant genomes: monophyly of extant gymnosperms and origin of Gnetales from conifers. Proceedings of the National Academy of Sciences, USA 97, 4086–4091. Cheng Y, Dai X, Zhao Y. 2007a. Auxin synthesized by the YUCCA flavin monooxygenases is essential for embryogenesis and leaf formation in Arabidopsis. The Plant Cell 19, 2430–2439. Cheng Y, Qin G, Dai X, Zhao Y. 2007b. NPY1, a BTB-NPH3-like protein, plays a critical role in auxin-regulated organogenesis in Arabidopsis. Proceedings of the National Academy of Sciences, USA 104, 18825–18829. Conner JK, Agrawal AA. 2005. Mechanisms of constraints; the contributions of selection and genetic variance to the maintenance of cotyledon number in wild radish. Journal of Evolutionary Biology 18, 238–242. Conway LJ, Poethig S. 1997. Mutations of Arabidopsis thaliana that transform leaves into cotyledons. Proceedings of the National Academy of Sciences, USA 94, 10209–10214. Datta SC. 1988. Systemic botany. New Delhi, India: Wiley Eastern Ltd. Dean E. 2002. Upcoming changes in flowering plant family names: those pesky taxonomists are at it again! Fremontia 30, 3–12. Degtjareva GV, Caspar SJ, Hellwig FH, Schmidt AR, Steiger J, Sokoloff DD. 2006. Morphology and nrITS phylogeny of the genus Pinguicula L. (Lentibulariaceae), with special attention to embryo evolution. Plant Biology 8, 778–790. Dharmasiri N, Dharmasiri S, Estelle M. 2005a. The F-box protein TIR1 is an auxin receptor. Nature 435, 441–445. Dharmasiri N, Dharmasiri S, Weijers D, Lechner E, Yamada M, Hobbie L, Ehrismann JS, Jürgens G, Estelle M. 2005b. Plant development is regulated by a family of auxin receptor F box proteins. Developmental Cell 9, 109–119. Dube VP, Awasthi DK, Singhal VP. 1981. Comparative anatomical observations on the dicotylous and tricotylous seedlings of Raphanus sativus L. (Brassicaceae). Acta Botanica Indica 9, 134–147. Duke JA. 1969. On tropical tree seedlings. Seeds, seedlings, systems and systematics. Annals of the Missouri Botanical Garden 56, 125–161. Eames AJ. 1961. Morphology of the angiosperms. New York: McGraw-Hill Company. Emery JF, Floyd SK, Alvarez J, Esched Y, Hawker NP, Izhaki A, Baum SF, Bowman JL. 2003. Radial patterning of Arabidopsis shoots by class III HD-ZIP and KANADI genes. Current Biology 13, 1768–1774. Eshed Y, Baum SF, Perea JV, Bowman JL. 2001. Establishment of polarity in laeral organs of plants. Current Biology 11, 1251– 1260. Eshed Y, Izhaki A, Baum SF, Floyd SK, Bowman JL. 2004. Asymmetric leaf development and blade expansion in Arabidopsis are mediated by KANADI and YABBY activities. Development 131, 2997–3006. Farjon A, Styles BT. 1997. Pinus (Pinaceae). Flora Neotropica Monograph 75, 221–224. Fernandez DE. 1997. Developmental basis of homeosis in precociously germinating Brassica napus embryos: phase change at the shoot apex. Development 124, 1149–1157. Fischer-Iglesias C, Sundberg B, Neuhaus G, Jones AM. 2001. Auxin distribution and transport during embryonic pattern formation in wheat. The Plant Journal 26, 115–129. Fleming A. 2005. The control of leaf development. New Phytologist 166, 9–20. Fletcher JJ. 1909. Illustrations of polycotyledony in the genus Persoonia, with some reference to Nuytsia. Proceedings of the Linnean Society of New South Wales 33, 867. Franceschini MC, Tressens SG. 2004. Morphology of fruits, seeds and embryos of Argentinian Capparis L. (Capparaceae). Botanical Journal of the Linnean Society 145, 209–218. Friml J. 2003. Auxin transport: shaping the plant. Current Opinion in Plant Biology 6, 7–12. Friml J, Vieten A, Sauer M, Weijers D, Schwarz H, Hamann T, Offringa R, Jürgens G. 2003. Efflux-dependent auxin gradients establish the apical-basal axis of Arabidopsis. Nature 426, 147–153. Fujita T, Sakaguchi H, Hiwatashi Y, Wagstaff SJ, Ito M, Deguchi H, Sato T, Hasebe M. 2008. Convergent evolution of shoots in land plants: lack of auxin polar transport in moss shoots. Evolution and Development 10, 176–186. Furutani M, Vernoux T, Traas J, Kato T, Tasaka M, Aida M. 2004. PIN-FORMED1 and PINOID regulate boundary formation and cotyledon development in Arabidopsis embryogenesis. Development 131, 5021–5030. Furutani M, Kajiwara T, Kato T, Treml BS, Stockum C, TorreRuiz RA, Tasaka M. 2007. The gene MACCHI-BOU 4/ ENHANCER OF PINOID encodes a NPH3-like protein and reveals similarities between oprganogenesis and phototropism at the molecular level. Development 124, 3849–3859. Cotyledon development and evolution 2929 Golz JF. 2006. Signalling between the shoot apical meristem and developing lateral organs. Plant Molecular Biology 60, 889–903. Guilfoyle TJ, Hagen G. 2007. Auxin response factors. Current Opinion in Plant Biology 10, 453–460. Harrison LG, von Aderkas P. 2004. Spatially quantitative control of the number of cotyledons in a clonal population of somatic embryos of hybrid larch Larix3leptoeuropaea. Annals of Botany 93, 423–434. Haskell G. 1954. Pleiocotyledony and differentiation within angiosperms. Phytomorphology 4, 140–152. Heisler M, Ohno C, Das P, Sieber P, Reddy GV, Long JA, Meyerowitz E. 2005. Patterns of auxin transport and gene expression during primordium development revealed by live imaging of the Arabidopsis inflorescence meristem. Current Biology 15, 1899–1911. Hochholdinger R, Zimmermann R. 2008. Conserved and diverse mechanisms in root development. Current Opinion in Plant Biology 11, 70–74. Holtorp HE. 1944. Tricotyledony. Nature 153, 13–14. Hu J, Miller JF, Chen J, Vick BA. 2005. Preliminary observation on a spontaneous tricotyledonous mutant in sunflower. Research Workshop Proceedings. 27th Sunflower Research Workshop, January 12–13, 2005, Fargo, ND. Ikeda Y, Banno H, Niu QW, Howell SH, Chua NH. 2006. The ENHANCER OF SHOOT REGENERATION 2 gene in Arabidopsis regulates CUP-SHAPED COTYLEDON 1 at the transcriptional level and controls cotyledon development. Plant and Cell Physiology 47, 1443–1456. Izhaki A, Bowman J. 2007. KANADI and Class III HD-Zip gene families regulate embryo patterning and modulate auxin flow during embryogenesis in Arabidopsis. The Plant Cell 19, 495–508. Jang J-C, Fujioka S, Tasaka M, Seto H, Takatsuto S, Ishii A, Aida M, Yoshida S, Sheen J. 2000. A critical role of sterols in embryonic patterning and meristem programming revealed by the fackel mutants of Arabidopsis thaliana. Genes and Development 14, 1485–1497. Jenik PD, Barton K. 2005. Surge and destroy: the role of auxin in plant embryogenesis. Development 132, 3577–3585. Jenik PD, Gillmor CS, Lukowitz W. 2007. Embryonic patterning in Arabidopsis thaliana. Annual Review of Cell and Developmental Biology 23, 207–236. Jönsson H, Heisler MG, Shapiro BE, Meyerowitz EM, Mjolsness E. 2006. An auxin-driven polarized transport model for phyllotaxis. Proceedings of the National Academy of Sciences, USA 103, 1633–1638. Kaplan DR, Cooke TJ. 1997. Fundamental concepts in the embryogenesis of dicotyledons: a morphological interpretation of embryo mutants. The Plant Cell 9, 1903–1919. Karschon R. 1973. Seedling morphology, and schizocotyly in Hammada salicordia (Moq.). Iljin. leaflet 46. Ilanot, Israel, Forestry Division, Agricultural Research Organisation. Keddie J, Carroll BJ, Thomas CM, Reyes ME, Klimyuk V, Holtan H, Gruissem W, Jones JDG. 1998. Transposon tagging of the Defective embryo and meristems gene of tomato. The Plant Cell 10, 877–887. Keith K, Kraml M, Dengler NG, McCourt P. 1994. fusca3: a heterochronic mutation affecting late embryo development in Arabidopsis. The Plant Cell 6, 589–600. Kepinski S, Leyser O. 2005. The Arabidopsis TIR1 protein is an auxin receptor. Nature 435, 446–451. Kerstetter RA, Bollman K, Taylor RA, Bomblies K, Poethig RS. 2001. KANADI regulates organ polarity in Arabidopsis. Nature 41, 706–709. Koyama T, Furutani M, Tasaka M, Ohme-Takagi M. 2007. TCP transcription factors control the morphology of shoot lateral organs via negative regulation of the expression of boundaryspecific genes in Arabidopsis. The Plant Cell 19, 473–484. Larsson E, Sitbon F, Ljung K, Von Arnold S. 2008. Inhibited polar auxin transport results in aberrant embryo develoment in Norway spruce. New Phytologist 177, 356–366. Laufs P, Peaucelle A, Morin H, Traas J. 2004. MicroRNA regulation of the CUC genes is required for boundary size control in Arabidopsis meristems. Development 131, 4311–4322. Laux T, Würschum T, Breuninger H. 2004. Genetic regulation of embryonic pattern formation. The Plant Cell 16, S190–S202. Liu C-M, Xu Z-H, Chua N-H. 1993. Auxin polar transport is essential for the establishment of bilateral symmetry during early plant embryogenesis. The Plant Cell 5, 621–630. Liu C-M, Johnson S, Di Gregorio S, Wang TL. 1999. single cotyledon (sic) mutants of pea and their significance in understanding plant embryo development. Developmental Genetics 25, 11–22. Long J, Barton MK. 1998. The development of apical embryonic pattern in Arabidopsis. Development 125, 3027–3035. Long JA, Moan EI, Medford JI, Barton MK. 1996. A member of the KNOTTED class of homeodomain proteins encoded by the STM gene of Arabidopsis. Nature 379, 66–69. Long JA, Woody S, Poethig S, Meyerowitz EM, Barton MK. 2002. Transformation of shoots into roots in Arabidopsis embryos mutant at the TOPLESS locus. Development 129, 2297–2306. Long JA, Ohno C, Smith Z, Meyerowitz E. 2006. TOPLESS regulates apical embryonic fate in Arabidopsis. Science 312, 1520–1523. Madishetty K, Bauer P, Sharada MS, Al-Hammadi ASA, Sharma R. 2006. Genetic characterization of the polycotyledon locus in tomato. Theoretical and Applied Genetics 113, 673–683. Mantegazza R, Möller M, Harrison CJ, Fior S, De Luca C, Spada A. 2007. Anisocotyledony and meristem initiation in an unorthodox plant, Streptocarpus rexii (Gesneriaceae). Planta 225, 653–663. Marx GA. 1987. A suite of mutants that modify pattern formation in pea leaves. Plant Molecular Biology Reports 5, 311–335. McAbee JM, Hill TA, Skinner DJ, Izhaki A, Hauser BA, Meister RJ, Venugopala Redy G, Meyerowitz EM, Bowman JL, Gasser CS. 2006. ABERRANT TESTA SHAPE encodes a KANADI family member linking polarity determination to separation and growth of Arabidopsis ovule integuments. The Plant Journal 46, 522–531. McConnell JR, Emery J, Eshed Y, Bao N, Bowman J, Barton MK. 2001. Role of PHABULOSA and PHAVOLUTA in determining radial patterning in shoots. Nature 411, 709–713. McSteen P, Malcomber S, Skirpan A, Lunde C, Wu X, Kellogg E, Hake S. 2007. barren inflorescence2 encodes a coorthologue of the PINOID serine/threonine kinase and is required for organogenesis during inflorescence and vegetative development in maize. Plant Physiology 144, 1000–1011. Meinke DW, Franzmann LH, Nickle TC, Yeung EC. 1994. Leafy cotyledon mutants of Arabidopsis. The Plant Cell 6, 1049–1064. Michniewicz M, Zago MK, Abas L, et al. 2007. Antagonistic regulation of PIN phosphorylation by PP2A and PINOID directs auxin flow. Cell 130, 1044–1056. Mirov NT. 1967. The genus Pinus. New York: Ronald Press Company. Nagesh K, Gaud GK, Raj MP, Rao PS. 2001a. Tricotyledony in Prosopis cineraria (L.) Druce. (Mimosaceae). Indian Forester 127, 591–592. Nagesh K, Reddy CS, Bhanja MR. 2001b. Tricotyledony in Acacia mellifera (Vahl) Benth. (Mimosaceae). Indian Journal of Forestry 23, 440–441. Nagesh K, Kardam TS. 2004. Tricotylous seedlings in Tectona grandis Linn.F. (Verbenaceae). Indian Forester 130, 819–820. 2930 Chandler Nagesh K, Ramesh M, Venkaiah K, Rao PS. 2004. Tricotylous seedlings in Sapindus emarginatus Vahl (Sapindaceae). Indian Forester 130, 463. Nawy T, Lukowitz W, Bayer M. 2008. Talk global, act local: patterning in the Arabidopsis embryo. Current Opinion in Plant Biology 11, 28–33. Nickrent DL, Soltis DE. 1995. A comparison of angiosperm phylogenies from nuclear 18S rDNA and rbcL sequences. Annals of the Missouri Botanic Gardens 82, 208–234. Nishii K, Kuwabara A, Nagata T. 2004. Characterization of anisocotylous leaf formation in Streptocarpus wendlandii (Gesneriaceae): significance of plant growth regulators. Annals of Botany 94, 457–467. Nodine MD, Yadegari R, Tax FE. 2007. RPK1 and TOAD2 are two receptor-like kinases redundantly required for Arabidopsis embryonic pattern formation. Developmental Cell 12, 943–956. Nodine MD, Tax FE. 2008. Two receptor-like kinases required together for the establishment of Arabidopsis cotyledon primordia. Developmental Biology 314, 161–170. Pillai A, Goyal SC. 1983. Developmental anatomy of some oilyielding plants. IV. Normal and tricotylous seddlings of Sesamum indicum L. Feddes Repert 94, 87–90. Prigge MJ, Otsuga D, Alonso JM, Ecker JR, Drews GN, Clarke S. 2005. Class III homeodomain-leucine zipper gene family members have overlapping, antagonistic and distinct roles in Arabidopsis development. The Plant Cell 17, 61–76. Quint M, Gray WM. 2006. Auxin signaling. Current Opinion in Plant Biology 9, 448–553. Rai SP, Kumar S. 2000. Induced mutation to monocotyledony in periwinkle, Catharanthus roseus, and suppression of mutant phenotype by kinetin. Journal of Genetics 79, 97–104. Rai SP, Kumar S. 2001. A tricotyledonous seedling mutant with Medelian inheritance in periwinkle Catharanthus roseus. Journal of Medicinal and Aromatic Plant Science 22, 267–268. Rajora OP, Zsuffa L. 1986. Atypical seedlings of Populus L.: their genetic significance and value in breeding. Silvae Genetica 35, 22–124. Ramesar-Fortner NS, Yeung EC. 2001. Tri-iodobenzoic acid affects shoot apical meristem formation and function in zygotic embryos of Brassica napus cv. Topas. Canadian Journal of Botany 79, 265–273. Reinhardt D. 2005. Regulation of phyllotaxis. International Journal of Developmental Biology 49, 539–546. Reinhardt D, Pesce ER, Steiger P, Mandel T, Baltensperger K, Bennett M, Traas J, Friml J, Kuhlemeier C. 2003. Regulation of phyllotaxis by polar auxin transport. Nature 426, 55–260. Satoh N, Hong S-K, Nishimura A, Matsuoka M, Kitano H, Nagato Y. 1999. Initiation of shoot apical meristem in rice: characterization of four SHOOTLESS genes. Development 126, 3629–3636. Sawa S, Watanabe K, Goto K, Kanaya E, Morita EH, Okada K. 1999. FILAMENTOUS FLOWER, a meristem and organ identity gene of Arabidopsis, encodes a protein with a zinc finger and HMG-related domains. Genes and Development 13, 1079–1088. Schrick K, Mayer U, Horrichs A, Kuhnt C, Bellini C, Dangl J, Schmidt J, Jürgens G. 2000. FACKEL is a sterol C-14 recuctase required for organized cell expansion in Arabidopsis embryogenesis. Genes and Development 14, 1471–1484. Sentoku N, Sato Y, Kurata N, Ito Y, Kitano H, Matsuoka M. 1999. Regional expression of the rice KN1-type homeobox gene family during embryo, shoot and flower development. The Plant Cell 11, 1651–1664. Siegfried KR, Eshed Y, Baum SF, Otsuga D, Drews G, Bowman J. 1999. Members of the YABBY gene family specify abaxial cell fate in Arabidopsis. Development 126, 4117–4128. Sokoloff DD, Remizowa MV, Macfarlane TD, Tuckett RE, Ramsay MM, Beer AS, Yadav SR, Rudall PJ. 2007. Seedling diversity in Hydatellaceae: implications for the evolution of angiosperm cotyledons. Annals of Botany 101, 153–164. Soltis PS, Soltis DE. 2004. The origin and diversification of angiosperms. American Journal of Botany 91, 1614–1626. Souer E, van Houweligen A, Kloos D, Mol J, Koes R. 1996. The no apical meristem gene of Petunia is required for pattern formation in embryos and flowers and is expressed at meristem and primordia boundaries. Cell 85, 159–170. Spurr AR. 1949. Histogenesis and organization of the embryo in Pinus strobus L. American Journal of Botany 36, 629–641. Stone S, Kwong L, Yee KM, Pelletier J, Fischer RL, Goldberg RB, Harada JJ. 2001. LEAFY COTYLEDON2 encodes a B3 domain transcription factor that induced embryo development. Proceedings of the National Academy of Sciences, USA 98, 11806–11811. Stubbe H. 1966. Genetik und Zytologie von Antirrhinum L. sect. Antirrhinum. Jona: VEB Gustav Fischer. Szemenyei H, Hannon M, Long JA. 2008. TOPLESS mediates auxin-dependent transcriptional repression during Arabidopsis embryogenesis. Science 319, 1384–1386. Takada S, Hibara K-I, Ishida T, Tasaka M. 2001. The CUPSHAPED COTYLEDON1 gene of Arabidopsis regulates shoot apical meristem formation. Development 128, 1127–1135. Tan X, Calderon-Villalobos LI, Sharon M, Zheng C, Robinson CV, Estelle M, Zheng N. 2007. Mechanism of auxin perception by the TIR1 ubiquitin ligase. Nature 446, 640–645. Tanaka H, Watanabe M, Sasabe M, Hiroe T, Tanaka T, Tsukaya H, Ikezaki M, Machida C, Machida Y. 2007. Novel receptor-like kinase ALE2 controls shoot development by specifying epidermis in Arabidopsis. Development 134, 1643–1652. Tillich HJ. 1990. The seedlings of Nymphaeaceae: monocotylar or dicotylar? Flora 184, 169–176. Tillich HJ. 1995. Seedling and systematics in monocotyledons. In: Rudall PJ, Cribb PJ, Cutler DF, Humphries CJ, eds. Monocotyledons: systematics and evolution. Kew, London, UK: Royal Botanic Gardens, 303–352. Timmermans MC, Hudson A, Becraft PW, Nelson T. 1999. ROUGH SHEATH2: a Myb protein that represses knox homeobox genes in maize lateral organ primordial. Science 284, 151–153. Titova GE. 2003. Algorithms of embryo morphogenesis in Agapanthus praecox Willd. (Alliaceae) in monocotyly, dicotyly and transitional forms. Acta Biologica Cracoviensia 45, 161–165. Titova GE, Batygina TB. 1986. Is the embryo of Nymphaealean plants (Nymphaeales s.l.) dicotyledonous? Phytomorphology 46, 171–190. Topping JF, May VJ, Muskett PR, Lindsey K. 1997. Mutations in the HYDRA1 gene of Arabidopsis perturb cell shape and disrupt embryonic and seedling morphogenesis. Development 124, 4415–4424. Torres-Ruiz RA, Lohner A, Jürgens G. 1996. The GURKE gene is required for the normal organization of the apical region in the Arabidopsis embryo. The Plant Journal 10, 1005–1016. Treml BS, Winderl S, Radykewicz R, Herz M, Schweizer G, Hutzler P, Glawischnig E, Ruiz RA. 2005. The gene ENHANCER OF PINOID controls cotyledon development in the Arabidopsis embryo. Development 132, 4063–4074. Tsiantis M, Schneeberger R, Golz JF, Freeling M, Langdale J. 1999. The maize rough sheath2 gene and leaf development programs in monocot and dicot plants. Science 284, 154–156. Tsukaya H. 1997. Determination of the unequal fate of cotyledons of a one-leaf plant, Monophyllaea. Development 124, 1275–1280. Ulmasov T, Murfett J, Hagen G, Guilfoyle TJ. 1997. Aux/IAA proteins repress expression of reporter genes containing natural Cotyledon development and evolution 2931 and highly active synthetic auxin response elements. The Plant Cell 9, 1963–1971. Vernon DM, Meinke DW. 1994. Embryonic transformation of the suspensor in twin, a polyembryonic mutant of Arabidopsis. Developmental Biology 165, 566–573. Vernon DM, Hannon MJ, Le M, Forsthoefel NR. 2001. An expanded role for the TWN1 gene in embryogenesis: defects in cotyledon pattern and morphology in the TWN1 mutant of Arabidopsis (Brassicaceae). American Journal of Botany 88, 570–582. Vernoud V, Hajduch M, Khaled A-S, Depège N, Rogowski P. 2005. Maize embryogenesis. Maydica 50, 469–483. Von Aderkas P. 2002. In vitro phenotypic variation in larch cotyledon number. International Journal of Plant Sciences 163, 301–307. Vroeman CW, Mordhurst AP, Albrecht C, Kwaaitaal MA, de Vries SC. 2003. THE CUP-SHAPED COTLEDON3 gene is required for boundary and shoot apical meristem formation in Arabidopsis. The Plant Cell 15, 1563–1577. Wardlaw CW. 1955. Embryogenesis in plants. London: Methuen. Weir I, Lu J, Cook H, Causier B, Schwarz-Sommer Z, Davies B. 2004. CUPULIFORMIS establishes lateral organ boundaries in Antirrhinum. Development 131, 915–922. Wenkel S, Emery J, Hou B-H, Evams MMS, Barton MK. 2007. A feedback regulatory module formed by LITTLE ZIPPER and HD-ZIPIII genes. The Plant Cell 19, 3379–3390. West M, Yee KM, Danao J, Zimmerman JL, Fischer RL, Goldberg RB, Harada JJ. 1994. LEAFY COTYLEDON1 is an essential regulator of late embryogenesis and cotyledon identity in Arabidopsis. The Plant Cell 6, 1731–1745. Williams L, Grigg SP, Xie M, Christensen S, Fletcher J. 2005. Regulation of Arabidopsis shoot apical meristem and lateral organ formation by microRNA miR166g and its AtHD-ZIP target genes. Development 132, 3657–3668. Zimmer A, Lang D, Richardt S, Frank W, Reski R, Rensing R. 2007. Dating the evolution of plants: detection and molecular clock analyses of orthologues. Molecular Genetics and Gemonics 278, 393–402. Zimmermann R, Werr W. 2005. Pattern formation in the monocot embryo as revealed by NAM and CUC3 orthologues from Zea mays L. Plant Molecular Biology 58, 669–685. Zimmermann R, Werr W. 2007. Transcription of the putative maize orthologue of the Arabidopsis DORNROSCHEN gene marks early asymmetry in the proembryo and during leaf initiation in the shoot apical meristem. Gene Expression Patterns 7, 158–164.