* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download endosymbiosis
Genetic engineering wikipedia , lookup
Transposable element wikipedia , lookup
Non-coding DNA wikipedia , lookup
Signal transduction wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Community fingerprinting wikipedia , lookup
Genomic library wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Gene expression wikipedia , lookup
Gene regulatory network wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
Mitochondrion wikipedia , lookup
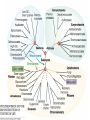
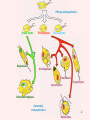
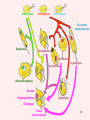
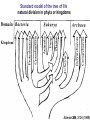
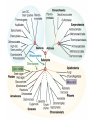
Recent discoveries about the molecular evolution of the three domains of life (Bacteria, Archaea, Eukaryota) Manolo Gouy Laboratoire de Biométrie & Biologie Evolutive - CNRS / Univ. Lyon 1 January 2009 1 Today the 3 domains of life LUCA First cell Origin(s) of life Third age : Post-luca cellular world Second age : Pre-luca cellular world First age : Pre-cellular world LUCA: last universal common ancestor 2 Discovery in 1977 of the three domains of life 3 SAB : similarity score between fragments of 2 rRNA molecules. SAB scores are high within each of the 3 groups and low between groups. 4 5 Search of the root of the universal phylogeny Ancestral duplication Ancestral duplication Ancestral duplication E: eucaryotes; B: bacteria; A: archea LUCA: Last Universal Common Ancestor 6 7 Current hypotheses about the origin of the three domains 1) The most widely accepted one, based of analyses of a few pre-LUCA gene duplicates. 2) Iconoclastic hypothesis proposed by some authors: the prokaryotic state (simple) is seen as resulting from a simplification rather than as ancestral. 4) The eukaryotic cell is seen as resulting from an archae + bacterium 8 fusion. Several scenarios have been proposed. Evolutionary history of the mitochondrial endosymbiosis - what was the donor organism ? - was the endosymbiosis unique or repeated ? - when did it occur ? what eukaryotic lineages received it ? 9 The unique endosymbiotic origin of mitochondria from an ancestral -proteobacterium -proteobacteria mitochondria Concatenation of amino acid sequences of respiratory chain proteins apocytochrome b (Cob) and cytochrome oxidase subunits 1 to 3 (Cox1-3). 10 Conservation of gene order between mitochondria and -proteobacteria 11 The gene-richest mitochondrial genome known as of today. 12 Evolutionary history of the mitochondrial endosymbiosis - what was the donor organism ? - was the endosymbiosis unique or repeated ? - when did it occur ? what eukaryotic lineages received it ? 13 Phylogenetic analysis of small subunit ribosomal RNA Mitochondrial symbiosis? Amitochondrial eucaryotes: « Archezoa » 14 Cavalier-Smith & Chao (1996) J Mol Evol 43:551 Eucaryotes from before the birth of mitochondria Tom Cavalier-Smith (1987) Nature 326:332 “It is a widespread fallacy that mitochondria are found in all eukaryotic cells.” “It is not the mitochondria, but the nucleus, endomembrane system and cytoskeleton that are the true hallmarks of the eukaryote cell.” “The idea that some protozoa are the living relics of the earliest phase of eukaryote cell evolution and diverged from our ancestors before the symbiotic origin of mitochondria is given strong support by DNA sequence studies.” 15 Microsporidia spore polar tube sporoplasm (nucleus + cytoplasm) spore of Nosema algerae (Undeen 1997) • > 1000 species • unicellular eukaryotes of very small size • obligate intracellular parasites • amitochondriate, aperoxysomal • evolutionary origin subject of much debate 16 Phylogenetic analysis of b-tubulin 17 Edlind et al. (1996) Mol. Phyl. Evol. 5:359. Phylogenetic analysis of RNA polymerase II large subunit Hirt et al. (1999) Proc.Natl.Acad.Sci. USA 96:580 18 Is it possible to reconcile ribosomal RNAs, tubulins and RNA polymerases ? MICROSPORIDIA ? Amitochondrial eucaryotes 19 The Long Branch Attraction artifact [ Felsenstein (1978) Syst Zool 27:401 ] 20 Philippe et al. (2000) Proc. Royal Soc. Lond. B 267:1213. Distance-based analysis of 42 LSU rRNA sequences from microsporidia and other eukaryotes. Distances were corrected for site-to-site rate variation. So this analysis uses a more realistic model of molecular evolution. 21 Van de Peer et al. (2000) Gene 246:1 Conclusion at this stage: The nice correspondence, for microsporidia, between - absence of mitochondria and - early origin among eucaryotes does not hold anymore. 22 Diversity of (anaerobe) ‘amitochondrial’ protists • no detectable‘mitochondrial’ organelle (microsporidia, diplomonads, Entamoeba,…) • hydrogenosomes: genomeless organelles producing ATP and H2 (some ciliates, anaerobe fungi, Parabasalia (ex: Trichomonas)) 23 from Roger & Silberman (2002) Nature 418:827. Gene shuffling during evolution of mitochondria Ancestral -proteobacterial endosymbiont transfer to nucleus loss stay A gene of mitochondrial evolutionary origin can be carried by 24 the nuclear genome of a eucaryote. Discovery in the genome of microsporidion Encephalitozoon cuniculi of several genes of mitochondrial evolutionary origin Example: IscS gene Homo-mt Mus-mt Caenorhabditis-mt 100 Candida albicans-mt 85 100 Candida maltosa-mt 99 Saccharomyces-mt 38 Schizosaccharomyces-mt 86 Encephalitozoon 33 Arabidopsis-mt Zygomonas 37 84 Rickettsia 37 Synechocystis Escherichia 100 Haemophilus 100 Rhodobacter capsulatus 97 Rhodobacter sphaeroides Rhizobium 100 100 Klebsiella Enterobacter Azospirillum 100 39 Azotobacter chroococcum 100 Azotobacter vinelandii 100 99 Anabaena azollae Anabaena sp 100 Helicobacter Thermotoga Lactobacillus 69 Archaeoglobus Aquifex Deinococcus Bacillus Methanobacterium Pyrococcus Mycoplasma sp. 25 100 Mycoplasma genitalium Aeropyrum 100 0.2 subst./site 55 48 48 59 75 100 100 100 100 Identification of 6 putative proteins that are closer to their mitochondrial or bacterial than to their eukaryotic homologues. These E. cuniculi proteins are very probably of mitochondrial evolutionary origin. Yeast homologues of these 6 E. cuniculi proteins : ATM1 (mitochondrial transporter ) ISU1 et ISU2 NFS1 (IscS cysteine desulfurase) SSQ1 (Heat Shock Protein 70 homologue) YAH1 PDB1 (pyruvate dehydrogenase E1 component b subunit) 5 of them are involved in the assembly of Fe-S clusters, co-factors of several mitochondrial and cytoplasmic enzymes. 26 Katinka et al. (2001) Nature 414:450. The microsporidian mitosome predicted by genome analysis: evolutionarily, it derives from a mitochondrion 27 Vivarès et al. (2002) Current Opinion in Microbiology 5:499. Detection of doublemembraned organelles by anti-HSP70 antibodies 28 The mitosome of the amitochondrial parasite Entamoeba histolytica Phylogeny of CPN60 mitosome Identification by cellular mapping of the CPN60 protein in Entamoeba histolytica Tovar et al. (1999) Mol. Microbiol. 32:1013. 29 Clark & Roger (1995) PNAS 92:6518. The mitosome of the amitochondrial parasite Giardia intestinalis Identification in Giardia of genes coding for mitochondrially targeted proteins in other eukaryotes : Cpn60, Hsp70, IscS (cysteine desulfurase) One example: IscS Tachezy et al. (2001) Mol. Biol. Evol. 18:1919. 30 double membrane Immunofluorescence mapping of IscS and IscU in Giardia trophozoites. 31 The missing link between mitochondrion and hydrogenosome Discovery [Akhmanova et al. (1998) Nature 396:527] and partial sequencing of the hydrogenosomal genome of the ciliate Nyctothermus ovalis 14,027 bp fragment of the hydrogenosomal genome - Several putative proteins of the hydrogenosomal genome group with their mitochondrial homologues of aerobic ciliate . - Identification of several nuclear genes coding for components of the mitochondrial proteome (pyruvate dehydrogenase, complex 32 II). Phylogenetic analyses of 2 hydrogenosomal genome genes nad7 (s.u. 49 kDa complex I) 12S (SSU) rRNA 33 Boxma et al. (2005) Nature 434:74. Death of the concept of primitively amitochondrial eucaryotes Mitochondrial symbiosis Amitochondrial eukaryotes « Archezoa » 34 Eukaryotic distribution of mitochondrial-derived organelles 35 Roger & Silberman (2002) Nature 418:827. Evolutionary history of the chloroplastic endosymbiosis - what was the donor organism ? - was the endosymbiosis unique or repeated ? - what eukaryotic lineages received it ? 36 37 Demonstration of the unique origin of primary photosynthetic eukaryotes 50 plastid proteins 143 nuclear proteins 38 Model of primary chloroplastic endosymbiosis 39 Secondary chloroplastic endosymbiosis 40 Secondary chloroplastic endosymbiosis in the cryptophyte Guillardia theta 41 Primary endosymbiosis Green plants Rhodophytes Glaucophytes ? Euglenozoa Dinoflagellates Heterokonts Cryptophytes Apicomplexa Chlorarachniophytes Secondary endosymbioses 42 Haptophytes Green plants Rhodophytes Glaucophytes Secondary endosymbioses Euglenozoa Euglenozoa Dinoflagellates Dinoflagellates Dinoflagellates Heterokonts Cryptophytes Apicomplexa Chlorarachniophytes Chlorarachniophytes Secondary plastid replacement Karenia (Lepidodinium) Kryptoperidinium Haptophytes Dinophysis Tertiary endosymbioses 43 State of the art about phylogenetic knowledge at the scale of each domain of the tree of life - Much debate for bacterial and archeal domains. Is the concept of phylogenetic tree adequate ? - Eukaryotic domain phylogeny : after much confusion, some structure begins to emerge. 44 Bacterial domain phylogeny. In the « classical » vision, a natural division in phyla exists. 45 Archaeal domain phylogeny. Recent discovery of a new phylum. 46 Standard model of the tree of life natural division in phyla or kingdoms 47 Alternative model: horizontal gene transfers between prokaryotes are so frequent that the notion of natural phyla does not apply. 48 Eukaryotic domain phylogeny. Emerging consensus for the identification of five superphyla. Relationships between them remain very uncertain. 49 Models of the origin of the eukaryotic cell - the fusion hypotheses - how to test these hypotheses 50 A Two-step scenario - creation by an bacteria/archaea fusion of an amitochondrial eukaryotic cell then - endosymbiosis with an -proteobacterium B Simultaneous creation of the eukaryotic nucleus and of the mitochondrion. The bacterial partner is always an proteobactérium. The archaeal partner varies according to theories (here a methanogen). 51 Models of the origin by fusion of the eukaryotic cell a-d: 2-step models e-g: mitochondrial origin = eukaryotic cell creation These hypotheses are testable: each predict similarities between part of the eukaryotic genome and some bacteria, and between the rest of this genome and some archaea. 52 53 Homogeneous model Heterogeneous model Recent evidence for a link between eukaryotes and a specific archaeal lineage “The archaebacterial origin of eukaryotes.” 54