* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Structure, expression and phylogenetic analysis of the glycoprotein
Gene regulatory network wikipedia , lookup
Signal transduction wikipedia , lookup
Interactome wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
Metalloprotein wikipedia , lookup
Biosynthesis wikipedia , lookup
Magnesium transporter wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Plant virus wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Biochemistry wikipedia , lookup
Genetic code wikipedia , lookup
Homology modeling wikipedia , lookup
Gene expression wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Western blot wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Expression vector wikipedia , lookup
Protein structure prediction wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Proteolysis wikipedia , lookup
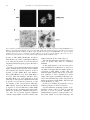
Virus Research 54 (1998) 197 – 205 Short communication Structure, expression and phylogenetic analysis of the glycoprotein gene of Cocal virus Resham S. Bhella 1,a, Stuart T. Nichol b, Essam Wanas a, Hara P. Ghosh a,* b a Department of Biochemistry, McMaster Uni6ersity, 1200 Main Street West, Hamilton, Ontario L8N 3Z5, Canada Special Pathogens Branch, Di6ision of Viral and Rickettsial Diseases, Centers for Disease Control and Pre6ention, Atlanta, GA 30333, USA Received 8 October 1997; received in revised form 4 December 1997; accepted 10 December 1997 Abstract A cDNA copy of the mRNA of the glycoprotein G of Cocal virus, a rhabdovirus, has been cloned, sequenced and expressed in mammalian cells. The deduced amino acid sequence shows a typical transmembrane glycoprotein, 512 amino acids in length, containing two potential N-linked glycosylation sites. The amino acid sequence showed a high degree of identity with that of the prototype vesicular stomatitis virus serotype Indiana [VSV (IND)] G protein. In addition, phylogenetic analysis of amino acid sequence differences among the G proteins of vesiculoviruses indicated that Cocal virus represents a distinct lineage within the VSV (IND) serotype. Expression of the cloned Cocal G gene in mammalian cells produced a glycoprotein of mol.wt 71000 which was not palmitylated but induced cell fusion at acid pH. © 1998 Elsevier Science B.V. All rights reserved. Keywords: Glycoprotein G; Cocal virus; Vesicular stomatitis; Rhabdovirus Glycoproteins of enveloped animal viruses have been used extensively to study biogenesis, transport and targeting of membrane glycoproteins (Ghosh, 1980; Garoff, 1985; Einfeld and Hunter, * Corresponding author. Tel.: +1 905 5259140, ext. 22451; fax: +1 905 5229033; e-mail: [email protected] 1 Present address: NRC Plant Biotechnology Research Institute, Saskatoon, Saskatchewan, Canada. 1991; Pettersson, 1991). The single glycoprotein G of vesicular stomatitis virus of the Indiana serotype [VSV (IND)], which is the prototype of the negative-stranded rhabdovirus family, has been used as a model in many of these studies. Since the G protein plays a central role in virus entry and assembly of infectious particles as well as antigenicity and host range determination, identification of the various functional domains of 0168-1702/98/$19.00 © 1998 Elsevier Science B.V. All rights reserved. PII S0168-1702(98)00006-9 198 R.S. Bhella et al. / Virus Research 54 (1998) 197–205 this protein is essential to understand the structure-function relationship of this protein (Wagner, 1990). A knowledge of the sequences of G proteins of other serological strains of VSV as well as temperature sensitive mutants, is also required to examine evolutionary relationship and antigenic diversity of these viruses (Bilsel and Nichol, 1990). The G protein genes of three vesiculoviruses other than VSV (IND), namely VSV New Jersey serotype [VSV (NJ)], Chandipura (CHP) and Piry have been sequenced (Rose and Gallione, 1981; Gallione and Rose, 1983; Masters et al., 1989, 1990; Brun et al., 1995). Cocal virus was originally isolated from mites in Trinidad (Jonkers et al., 1984), but infections have since been detected in Trinidad, Brazil and Argentina including isolation of the virus from cattle, horses, and insects (Karabatsos, 1985). The virus is serologically distinct but related to the VSV (IND) prototype strain (Jonkers et al., 1984; Travassos da Rosa et al., 1984). We report here, the complete sequence of the G protein gene of Cocal virus. The cloned gene expresses a biologically active G protein. Cocal virus was grown in monolayers of mouse L or baby hamster kidney (BHK-21) or HeLa cells. A full length cDNA clone of the G mRNA of Cocal virus was generated from poly A+ mRNA using the protocol of Gubler and Hoffman (1983). The cDNA was cloned in the EcoRI site of pGEM7Zf(+) and finally into the COS cell expression vector pXM (Yang et al., 1986). DNA sequence was determined by using dideoxy chain termination method using a modified T7 DNA polymerase (Sequenase) (Sanger et al., 1977) and nested deletion of nucleotides (Henikoff, 1984). The sequence was reconfirmed by sequencing the DNA of the gene cloned in pXM [pXMG(COC)] using synthetic oligonucleotide primers. Genomic RNA extracted from Cocal virions was also sequenced to determine sequences of nucleotides 1 – 300, 971 – 1140 and 1460 –1652 by using oligonucleotide primers and avian myeloblast virus reverse transcriptase (Bilsel and Nichol, 1990). The complete nucleotide sequence of the G gene was determined (Fig. 1) and deposited in the Gene Bank, EMBL. The Cocal G mRNA spans 1602 nucleotide excluding the poly(A) tail. It contains a single continuous open reading frame starting at the first ATG at 29 nucleotides from the 5%-end. The encoded protein contains 512 amino acids with a calculated mol.wt of 58036, which is in agreement with its observed mobility in SDS-polyacrylamide gels (Kotwal et al., 1983). In agreement with other vesiculovirus mRNAs, the G mRNA has the following terminal sequences: 5% AACAG(N)23AUG-A coding region - UGAUGA(N)28UAUGA7 –poly(A) 3% The sequence 3% UUGUC 5% represents the vesiculovirus conserved transcription initiation (capping) sequence (Banerjee and Barik, 1992), while 3% AUACU7 5% represents the transcription termination and polyadenylation signal in the viral genome (Banerjee and Barik, 1992). As in the case of VSV (IND) genome the G-L intragenic sequence of Cocal virus 5% UAUGA7CUAACAG 3%, did not contain any insertion of nucleotides (Rose, 1980). The 5% AACAG 3% at the 3% end of G mRNA denotes the transcription initiation signal for the L gene. The translation initiator methione codon AUG like the other three vesiculovirus G proteins, also contains a strong eukaryotic ribosomal initiation context (Kozak, 1989). The deduced amino acid sequence of Cocal G protein (Fig. 2) was compared with the amino acid sequence of the N-terminal 24 residues of the mature glycoprotein isolated from Cocal virions or from the infected cells as well as with the amino acid sequence of the primary translation product of the G mRNA (Kotwal et al., 1983). The results showed identity between the deduced sequence with the sequence determined in both the signal sequence and in the sequence of the processed mature G protein. The predicted signal sequence of 17 residues was identical to the signal peptide determined from direct sequencing except for two residues: a Lys and Leu were determined at positions − 16 and − 11, respectively, instead of Asn and Thr residues predicted in the corresponding position. The N-terminal 24 residues were identical except for the fact that at position 22 the virion protein contained Tyr while the deduced sequence predicted His. These differences could be accounted for by one or two base R.S. Bhella et al. / Virus Research 54 (1998) 197–205 199 Fig. 1. Complete nucleotide sequence of glycoprotein G gene of Cocal virus. The conserved vesiculovirus transcription signals and transcription termination and polyadenylation signals (Banerjee and Barik, 1992) are underlined. The translation initiation and termination codons are indicated by arrowheads. changes occurring either during the cloning of the cDNA or at the virus genome as a result of random mutations produced during serial passages (Wagner, 1990). In order to ascertain the biological activity of the cloned G gene, COS cells were transfected with the plasmid pXMG(COC), an eukaryotic expression vector containing the complete coding 200 R.S. Bhella et al. / Virus Research 54 (1998) 197–205 Fig. 2. Alignment of the deduced amino acid sequences of the G proteins of Cocal, VSV Indiana serotype (IND) (Rose and Gallione, 1981), VSV New Jersey serotype (NJ) (Gallione and Rose, 1983), Chandipura (CHP) (Masters et al., 1989, 1990; Brun et al., 1995) and Piry (Brun et al., 1995) viruses. Dashed lines represent gaps introduced to maximize matching of amino acid residues. Residues not shown and shaded are identical to those of Cocal G protein. Shaded residues indicate conservative changes. The arrow indicates the signal peptide cleavage site. The asterisk above N indicates consensus site for N-linked glycosylation. The hydrophobic membrane anchoring domain is underlined. The conserved Cys, Pro and Trp residues are indicated with a circle. R.S. Bhella et al. / Virus Research 54 (1998) 197–205 201 Fig. 3. Expression, glycosylation and fatty acid acylation of Cocal G protein from cloned gene. (A) COS cells were transfected with pXMG(IND), pXMG(COC) or no DNA and labelled with [35S]methionine. The cell lysates were immunoprecipitated with anti-VSV Indiana G for pXMG(IND) transfected cells and anti-Cocal G antibody for cells transfected with pXMG(COC) or no DNA and analyzed by SDS-polyacrylamide gel electrophoresis (Li et al., 1993). (B) N-linked glycosylation of expressed Cocal G proteins was determined by the acquisition of endo H resistance (Li et al., 1993). Proteins expressed in cells transfected with pXMG(COC) were labeled with [35S]methionine for 15 min, chased with excess of non-radioactive methionine for a period of 0 min or 60 min, and immunoprecipitated with anti-Cocal G antibody. One half of the sample was treated with endo H ( +), and the other half was not (−). Samples were analyzed by SDS-polyacrylamide gel electrophoresis. (C) COS cells transfected with pXMG(COC) or pXMG(IND) were labeled with [3H]palmitic acid (Kotwal and Ghosh, 1984; Masters et al., 1989), immunoprecipitated with anti-Cocal G or anti-VSV Indiana G antibody, respectively, and analyzed on SDS-polyacrylamide gel. region of Cocal G gene. Transfected cells were labelled with [35S]methionine and labelled proteins were immunoprecipitated with anti-Cocal G antiserum. As shown in Fig. 3A, the antibody recognized a protein of about 71000 Da, the expected size of Cocal G protein (Kotwal et al., 1983). This protein was not present in an immunoprecipitate of COS cells transfected with the parent vector pXM. The enzyme endoglycosidase H (endo H) hydrolyzes the oligomannose moiety of N-linked glycoproteins containing unprocessed glycosyl residues, while the mature glycoproteins containing processed and complex oligosaccharide residues are resistant to endo H digestion (Kornfeld and Kornfeld, 1985). Digestion of the expressed protein with endo H shows that the immature G protein is sensitive to endo H but the mature G protein produced after 1-h chase is resistant to endo H digestion (Fig. 3B) indicating that Cocal G protein expressed from cloned G contained N-linked oligosaccharide. This is in agreement with the predicted Cocal G protein amino acid sequence, which shows the presence of two N-linked glycosylation sites at residues 180 and 337. Similar to the results obtained with Cocal virus infected cell (Kotwal and Ghosh, 1984), no palmitylation of the expressed G protein was also observed in the transfected COS cells, which is consistent with the absence of any Cys residues in the membrane anchor or cytoplasmic domains of Cocal G protein (Fig. 3C). The G(IND) protein was, as reported earlier, labeled with palmitic acid. As expected, the expressed G protein was also transported to the cell surface as determined by indirect immunofluorescence (Fig. 4A). Glycoprotein G of VSV expressed from cloned gene can induce membrane fusion at acidic pH in the absence of other viral gene products (Florkiewitz and Rose, 1984; Reidel et al., 1984). When COS cells expressing Cocal G protein were briefly exposed to a buffer of pH 5.6, polykaryons containing more than 20 nuclei were observed (Fig. 4B). This demonstrated that, like the G 202 R.S. Bhella et al. / Virus Research 54 (1998) 197–205 Fig. 4. Cell surface localization and polykaryon formation by Cocal G protein. (A) COS cells transfected with pXMG(COC) or no DNA were fixed with paraformaldehyde and reacted with rabbit anti-Cocal G antiserum and fluorescein isothiocynate-conjugated anti-rabbit immunoglobulin G (Zhang and Ghosh, 1994). (B) COS cells transfected with pXMG(COC) were exposed to fusion medium at pH 7.4 or 5.6 for 60 s, incubated in regular medium for 2 h, re-exposed to fusion medium at pH 7.4 or 5.6 for 60 s, incubated in regular medium for 1.5 h, fixed, stained and photographed (Zhang and Ghosh, 1994). protein of VSV (IND) (Florkiewitz and Rose, 1984; Reidel et al., 1984) or Chandipura (Masters et al., 1989), the G protein of Cocal virus showed low-pH dependent fusogenic property (Kotwal et al., 1983). In order to locate functionally homologous and unique domains in the G protein, we have aligned and compared the deduced sequences of the G proteins of VSV (IND) (Rose and Gallione, 1981), CHP (Masters et al., 1989, 1990; Brun et al., 1995), VSV (NJ) (Gallione and Rose, 1983), and Piry (Brun et al., 1995) with that of Cocal virus (Fig. 2). The sequence alignment shows that G protein of Cocal virus has about 48, 35 and 38% identity to that of G proteins of VSV (NJ), CHP and Piry, respectively. However, the identity of sequence of Cocal G with that of VSV (IND) G protein was 72%, and when conservative substitutions are included the similarity increased to 77%. Also, the G protein sequence alignment contains a large number of blocks of amino acids identical between the five viruses (Fig. 2). The following important structural features of Cocal G protein may thus be tentatively identified. (i) The signal sequence of Cocal G was previously determined by us (Kotwal et al., 1983) to be one amino acid longer than the signal sequence of G protein of VSV (IND) or VSV (NJ). The predicted signal sequence and the site of cleavage were identical to those determined by direct amino acid sequence determination (Kotwal et al., 1983), and N-terminal Lys residue was present in mature G proteins of VSV (IND), VSV (NJ) and Cocal (Kotwal et al., 1983). (ii) The two N-linked glycosylations occur at identical positions on all five G proteins. (iii) The membrane anchoring sequence of hydrophobic amino acids is located at positions 465–483 of Cocal G protein. Although the transmembrane sequences of glycoproteins of related virus usually do not show homology, the R.S. Bhella et al. / Virus Research 54 (1998) 197–205 sequence of the transmembrane domain of Cocal G protein shows a 53% identity with that of VSV (IND) G protein. (iv) The cytoplasmic domain of Cocal and the other four vesiculovirus G proteins showed very little conservation. However, the cytoplasmic sequence was shown to be essential in virus assembly presumably by recognition of the nucleoprotein complex (Whitt et al., 1989). (v) Three amino acids, Trp, Pro and Cys, are found to occur in non-variant positions in the G proteins of all five vesiculoviruses. Conserved Pro residues could define domains while the conserved Cys residues are possibly involved in disulfide bond formation essential for proper folding. It may be noted that no Cys residues are present after amino acid 301. (vi) Several blocks of amino acids are also conserved in all five sequences. The function of the conserved 17-amino acid stretch spanning residue 83–99 is not yet known. The conserved region spanning residue 124 – 137 has recently been identified as a fusogenic domain of G protein of VSV (IND) (Li et al., 1993; Zhang and Ghosh, 1994; Durrer et al., 1995; Frederickson and Whitt, 1995). This region may thus represent the fusion peptide of vesiculovirus G proteins. A conserved region encompassing residues 395–424 near the membrane anchoring domain of VSV (IND) G protein has also been recently shown to be important for membrane fusogenic activity of G protein and may play a key role in control of low-pH induced conformational change of rhabdovirus glycoproteins (Li et al., 1993; Gaudin et al., 1996; Shokralla et al., 1998). The functional role of the other conserved blocks containing 4 – 6 amino acids is yet to be established. Mutagenesis in the conserved regions of G protein and the effect of the mutations on biological activities of G protein may provide some information on the structure-function relationship of G proteins. (vii) Finally, phylogenetic analysis of the amino acid sequence differences among representative vesiculoviruses (Fig. 5) clearly indicates that Cocal virus is closely related to VSV (IND) but represents a distinct lineage separate from classical VSV (IND) strains. This corre- 203 Fig. 5. Phylogenetic relationship of vesiculoviruses based on analysis of G protein deduced amino acid sequence differences. Phylogenetic analysis was performed by the maximum parsimony method using PAUP 3.1 software (Swofford, 1991). The analysis was done using the Branch and Bound (BANDB) and MULPARS options, with PROTPARS weighting of the data matrix (Felsenstein, 1993), and bootstrap confidence limits obtained from 1000 repetitions of the analysis. The more distantly related vesiculovirus, Chandipura virus, G protein sequence was used to outgroup root the tree output. The horizontal distances represent the number of amino acid step differences (indicated by scale bar) present between branch nodes and taxa (i.e. viruses). Vertical distances are for graphic representation only. Bootstrap confidence limits exceeding 50% are indicated next to each branch node. lates with previous findings showing the serological cross-reactivity of Cocal virus with a VSV (IND) serotype prototypic virus isolate (Travassos da Rosa et al., 1984). Acknowledgements We thank Xiaynan Li for technical assistance, Pamuk Bilsel for preliminary sequences and L. Kush and M.M. Strong for manuscript preparation. This work was supported by Medical Research Council of Canada (H.P.G.). 204 R.S. Bhella et al. / Virus Research 54 (1998) 197–205 References Banerjee, A.K., Barik, S., 1992. Gene expression of vesicular stomatitis virus genome RNA. Virology 188, 417– 428. Bilsel, P.A., Nichol, S.T., 1990. Polymerase errors accumulating during natural evolution of the glycoprotein gene of vesicular stomatitis virus Indiana serotype isolates. J. Virol. 64, 4873 – 4883. Brun, G., Bao, X.K., Prevec, L., 1995. The relationship of Piry virus to other vesiculoviruses: a re-evaluation based on the glycoprotein gene sequence. Intervirology 38, 274 – 282. Durrer, P., Gaudin, Y., Ruigrok, R.W., Graf, R., Brunner, J., 1995. Photolabeling identifies a putative fusion domain in the envelope glycoprotein of rabies and vesicular stomatitis viruses. J. Biol. Chem. 270, 17575–17581. Einfeld, D., Hunter, E., 1991. Transport of membrane proteins to cell surface. Curr. Top. Microbiol. Immunol. 170, 107 – 139. Felsenstein, J., 1993. PHYLIP: Phylogeny Inference Package, version 3.5c. University of Washington, Seattle, WA. Florkiewitz, R.Z., Rose, J.K., 1984. A cell line expressing vesicular stomatitis virus glycoprotein fuses at low pH. Science 225, 721 – 723. Frederickson, B.L., Whitt, M.A., 1995. Vesicular stomatitis virus glycoprotein mutations that affect membrane fusion activity and abolish virus infectivity. J. Virol. 69, 1435 – 1443. Gallione, C.J., Rose, J.K., 1983. Nucleotide sequence of a cDNA clone encoding the entire glycoprotein from the New Jersey serotype of vesicular stomatitis virus. J. Virol. 46, 162 – 169. Garoff, H., 1985. Using recombinant DNA techniques to study protein targeting in the eurkaryotic cell. Ann. Rev. Cell Biol. 1, 403–446. Gaudin, Y., Raux, H., Flamand, A., Ruigrok, R.W.H., 1996. Identification of amino acids controlling the lowpH-induced conformational change of rabies virus glycoprotein. J. Virol. 70, 7371–7378. Ghosh, H.P., 1980. Synthesis and maturation of glycoproteins of enveloped animal viruses. Rev. Infect. Dis 2, 26– 39. Gubler, U., Hoffman, B.J., 1983. A simple and very efficient method for generating cDNA libraries. Gene 25, 263 – 269. Henikoff, S., 1984. Unidirectional digestion with exonuclease III creates targeted breakpoints for DNA sequencing. Gene 28, 351 – 359. Jonkers, A.H., Shope, R.E., Aitken, T.H.G., Spence, L., 1984. Cocal virus, a new agent in Trinidad related to vesicular stomatitus virus, type Indiana. Am. J. Vet. Res. 25, 236 – 242. Karabatsos, N., 1985. International Catalogue of Arboviruses Including Certain Other Viruses of Vertebrates. Am. Soc. of Trop. Med. Hyg., San Antonio, TX, pp. 337 – 338. Kotwal, G.J., Capone, J., Irving, R.A., Rhee, S.H., Bilan, P., Toneguzzo, F., Hofmann, T., Ghosh, H.P., 1983. Viral membrane glycoproteins: comparison of amino terminal amino acid sequences of the precursor and mature glycoproteins of three serotypes of vesicular stomatitis virus. Virology 129, 1 – 11. Kotwal, G.J., Ghosh, H.P., 1984. Role of fatty acid acylation of membrane glycoproteins. Absence of palmitic acid in glycoproteins of two serotypes of vesicular stomatitis virus. J. Biol. Chem. 259, 4699 – 4701. Kornfeld, R., Kornfeld, S., 1985. Assembly of asparaginelinked oligosaccharides. Ann. Rev. Biochem. 54, 631 – 664. Kozak, M., 1989. The scanning model for translation: an update. J. Cell. Biol. 108, 229 – 241. Li, Y., Drone, C., Sat, E., Ghosh, H.P., 1993. Mutational analysis of the vesicular stomatitis virus glycoprotein G for membrane fusion domains. J. Virol. 67, 4070 – 4077. Masters, P.S., Bhella, R.S., Butcher, M., Patel, B., Ghosh, H.P., Banerjee, A.K., 1989. Structure and expression of the glycoprotein gene of Chandipura virus. Virology 171, 285 – 290. Masters, P.S., Bhella, R.S., Butcher, M., Patel, B., Ghosh, H.P., Banerjee, A.K., 1990. Structure and expression of the glycoprotein gene of Chandipura virus. Virology 174, 630. Pettersson, R.F., 1991. Protein localization and virus assembly at intracellular membranes. Curr. Topics Microbiol. Immun. 170, 67 – 106. Reidel, H., Kondor-Koch, C., Garoff, H., 1984. Cell surface expression of fusogenic vesicular stomatitis virus G protein from cloned cDNA. EMBO J. 3, 1477 – 1483. Rose, J.K., 1980. Complete intergenic and flanking gene sequences from the genome of vesicular stomatitis virus. Cell 19, 415 – 421. Rose, J.K., Gallione, C.J., 1981. Nucleotide sequences of the mRNA’s encoding the vesicular stomatitis virus G and M proteins determined from cDNA clones containing the complete coding regions. J. Virol. 39, 519 – 528. Sanger, F., Nicklen, S., Coulson, A.R., 1977. DNA sequencing with chain-terminating inhibitors. Proc. Natl. Acad. Sci. USA 74, 5463 – 5467. Shokralla, S., He, Y., Wanas, E., Ghosh, H.P., 1998. Mutations in a carboxy-terminal region of vesicular stomatitis virus glycoprotein G that affect membrane fusion activity. Virology 242, 39 – 50. Swofford, D.L., 1991. PAUP: Phylogenetic Analysis Using Parsimony, version 3.0s. Computer program. Illinois Natural History Survey, Champaign, IL. Travassos da Rosa, A.P.A., Tesh, R.B., Travassos da Rosa, J.F., Herve, J.P., Main, A.J., 1984. Carajas and Maraba viruses, two new vesiculoviruses isolated from phlebotomine sand flies in Brazil. Am. J. Trop. Med. Hyg. 33, 999 – 1006. Wagner, R.R., 1990. Rhabdoviridge and their replication. In: Fields, B. (Ed.), Field’s Virology. Raven Press, New York, pp. 867 – 882. R.S. Bhella et al. / Virus Research 54 (1998) 197–205 Whitt, M.A., Chong, L., Rose, J.K., 1989. Glycoprotein cytoplasmic domain sequences required for rescue of a vesicular stomatitis virus glycoprotein mutant. J. Virol. 63, 3569 – 3578. Yang, Y.-C., Ciarletta, A.B., Temple, P.A., Chung, M.P., Kovacic, S., Witek-Giannotti, J.S., Leary, A.C., Kriz, R., Donahue, R.E., Wong, G.G., Clark, S.C., 1986. Human . . 205 IL-3 (multi-CSF): identification by expression cloning of a novel hematopoietic growth factor related to murine IL-3. Cell 47, 3 – 10. Zhang, L., Ghosh, H.P., 1994. Characterization of the putative fusogenic domain in vesicular stomatitis virus glycoprotein G. J. Virol. 68, 2186 – 2193.