* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download The Proton Motive Force
Biochemistry wikipedia , lookup
Citric acid cycle wikipedia , lookup
Adenosine triphosphate wikipedia , lookup
Photosynthesis wikipedia , lookup
NADH:ubiquinone oxidoreductase (H+-translocating) wikipedia , lookup
Metalloprotein wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Photosynthetic reaction centre wikipedia , lookup
Microbial metabolism wikipedia , lookup
Electron transport chain wikipedia , lookup
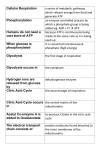
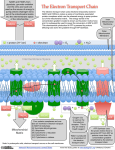
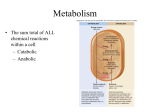
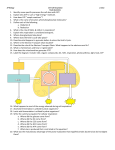
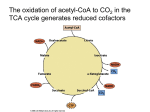
BSC 260 2/1/12 Metabolism Metabolism-The sum total of all chemical reactions that occur in a cell catabolism Energy-releasing metabolic reactions anabolism Energy-requiring metabolic reactions Nutrients Supply of monomers (or precursors of) required by cells for growth Macronutrients Nutrients required in large amounts Carbon Required by all cells Typical bacterial cell ~50% carbon (by dry weight) Major element in all classes of macromolecules Nitrogen Typical bacterial cell ~12% nitrogen Key element in proteins, nucleic acids, and many more cell constituents Other Macronutrients Phosphorus, Sulfur, Potassium, Magnesium, Calcium, and Sodium Micronutrients Nutrients required in trace amount Iron Key component of cytochromes and FeS proteins involved in electron transport Cells produce siderophores (iron-binding agents) to obtain iron from insoluble mineral form Culture Media Nutrient solutions used to grow microbes in the laboratory Two broad classes Defined media: precise chemical composition is known Complex media: composed of digests of chemically undefined substances (e.g., yeast and meat extracts) Selective Media Contains compounds that selectively inhibit growth of some microbes but not others Differential Media Contains an indicator, usually a dye, that detects particular chemical reactions occurring during growth Enzymes Typically proteins (some RNAs) Highly specific Typically rely on weak bonds Examples: hydrogen bonds, van der Waals forces, hydrophobic interactions Active site: region of enzyme that binds substrate Increase the rate of chemical reactions by 108 to 1020 times the spontaneous rate Oxidation–Reduction and Energy-Rich Compounds Energy from oxidation–reduction (redox) reactions is used in synthesis of energy-rich compounds (e.g., ATP) Redox reactions occur in pairs (two half reactions; Figure 4.8) Electron donor: the substance oxidized in a redox reaction Electron acceptor: the substance reduced in a redox reaction Essentials of Catabolism Two reaction series are linked to energy conservation in chemoorganotrophs: fermentation and respiration (Figure 4.13) Differ in mechanism of ATP synthesis Fermentation: substrate-level phosphorylation; ATP directly synthesized from an energy-rich intermediate Respiration: oxidative phosphorylation; ATP produced from proton motive force formed by transport of electrons Glycolysis Glucose consumed Two ATPs produced Fermentation products generated Some harnessed by humans for consumption Respiration and Electron Carriers Aerobic Respiration Oxidation using O2 as the terminal electron acceptor Higher ATP yield than fermentations ATP produced at the expense of the proton motive force, which is generated by electron transport Electron Transport Systems Membrane associated Mediate transfer of electrons Conserve some of the energy released during transfer and use it to synthesize ATP Many oxidation–reduction enzymes are involved in electron transport (e.g., NADH dehydrogenases, flavoproteins, iron–sulfur proteins, cytochromes) NADH dehydrogenases: proteins bound to inside surface of cytoplasmic membrane; active site binds NADH and accepts 2 electrons and 2 protons that are passed to flavoproteins Flavoproteins: contains flavin prosthetic group (e.g., FMN, FAD) that accepts 2 electrons and 2 protons but only donates the electrons to the next protein in the chain Cytochromes: Proteins that contain heme prosthetic groups Accept and donate a single electron via the iron atom in heme Iron–Sulfur Proteins: Contain clusters of iron and sulfur Example: ferredoxin Quinones: Hydrophobic non-protein-containing molecules that participate in electron transport Accept electrons and protons but pass along electrons only The Proton Motive Force Electron transport system oriented in cytoplasmic membrane so that electrons are separated from protons The final carrier in the chain donates the electrons and protons to the terminal electron acceptor During electron transfer, several protons are released on outside of the membrane Results in generation of pH gradient and an electrochemical potential across the membrane (the proton motive force) Terminal oxidase; reduces O2 to H2O ATP synthase (ATPase): complex that converts proton motive force into ATP; two components F1: multiprotein extramembrane complex, faces cytoplasm Fo: proton-conducting intramembrane channel Reversible; dissipates proton motive force The Citric Acid Cycle Citric acid cycle (CAC): pathway through which pyruvate is completely oxidized to CO2 Per glucose molecule, 6 CO2 molecules released and NADH and FADH generated Several high energy electrons stored in carrier molecules Energetics advantage to aerobic respiration Catabolic Diversity Microorganisms demonstrate a wide range of mechanisms for generating energy (Figure 4.22) Fermentation Aerobic respiration Anaerobic respiration Chemolithotrophy Phototrophy Anaerobic Respiration The use of electron acceptors other than oxygen Examples include nitrate (NO3), ferric iron (Fe3+), sulfate (SO42), carbonate (CO32), certain organic compounds Less energy released compared to aerobic respiration Dependent on electron transport, generation of a proton motive force, and ATPase activity Chemolithotrophy Uses inorganic chemicals as electron donors Examples include hydrogen sulfide (H2S), hydrogen gas (H2), ferrous iron (Fe2+), ammonia (NH3) Typically aerobic Begins with oxidation of inorganic electron donor Uses electron transport chain and proton motive force Autotrophic; uses CO2 as carbon source Phototrophy: uses light as energy source Photophosphorylation: light-mediated ATP synthesis Photoautotrophs: use ATP for assimilation of CO2 for biosynthesis Photoheterotrophs: use ATP for assimilation of organic carbon for biosynthesis