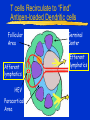
* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Innate Immunity
Gluten immunochemistry wikipedia , lookup
Lymphopoiesis wikipedia , lookup
Hygiene hypothesis wikipedia , lookup
Complement system wikipedia , lookup
Immune system wikipedia , lookup
Molecular mimicry wikipedia , lookup
Cancer immunotherapy wikipedia , lookup
Polyclonal B cell response wikipedia , lookup
Psychoneuroimmunology wikipedia , lookup
Adoptive cell transfer wikipedia , lookup
Adaptive immune system wikipedia , lookup
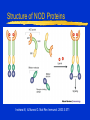
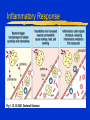
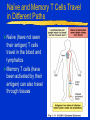
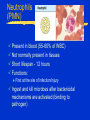
Innate Immunity -M261 Spring 2005 May 6, 2005 Kathleen A. Kelly Reading: Immunobiology (6th Edition) Janeway, Travers, Walpert & Capra Chapter 2 (p. 37-100), and Chapter 6 (p. 209-212) Fundamental Immunology (5th Edition) Lippincott, Williams & Wilkins Chapter 17 (p.497-517) Janeway, CA, et al. Innate Immune Recognition. Annu. Rev. Immunol. 20:197-252, 2002 Innate Immunity -M261 Spring 2004 Kathleen A. Kelly Innate immunity predates development of adaptive immunity Does not produce protective immunity ● No memory response ● Prerequisite for developing adaptive immunity Non-antigen-specific immunity ● Found in plants, invertebrates and vertebrates Innate Immunity 1. Provides a barrier to prevent the spread of infection ● Mechanical (tight junctions, movement) ● Chemical (fatty acids, enzymes, pH, antimicrobial peptides) ● Microbiological (normal flora) ● Mucosal surfaces o Nasopharyngeal, Oral, Respiratory, Intestinal tract Urogenital tract ● Skin (epithelial cells) o Wounds, burns, insect bites Innate Immunity 2. Identifies and eliminates pathogens ● Non-adaptive recognition systems ● Activates molecules that target the microbe and aid in it’s identification. o These factors may be expressed at the surface or within cells, released from immune cells or are secreted and present within circulatory system Innate Immunity 3. Initiates an inflammatory response ● Reaction to injury or infection o Trauma to tissues or cells o Presence of foreign matter (self vs. non-self) o Infectious agents (viruses, bacteria, fungi) ● Delivers effector molecules & immune cells to the site of infection ● Components o Leukocytes & secreted factors o Blood vessels o Plasma proteins Innate Immunity 4. Provides signals to activate and regulate the type of adaptive immune response generated ● Stimulation of co-stimulatory molecules o B7 family (CD80/86, PD-L, ICOSL) o TNFR family (OX40L) ● Induction of a cytokine/chemokine response o Cytokines: IL-12, IL-23, IL-4 o Chemokines: CXCR1, CXCR2, CCL20 • a variety and depends on stimulus The Phases of Immunity Identification of Microbes Recognition ● Receptors – Pattern Recognition Receptors (PRRs) o Fixed in the genome, ie gene rearrangement is not needed ● Distribution o Non-clonal, ie all cells of a class are identical Differentiation ● Pathogen vs. Commensal Identification of Microbes PRR ● Recognize conserved molecular patterns on microbes called microbe associated molecular patterns (MAMPs) which are not present on the host o Not limited to pathogens ● Identify a class of microbes o LPS, LTA, peptidoglycan, lipoarabinomannan, dsRNA, mannans, b-glycans ● MAMPs are often essential for microbe survival Action Time ● Immediate activation of effectors ● Delays need for adaptive immunity Pattern Recognition Receptors (PRRs) Three broad classes of PRRs based on expression profile, localization, function ● 1) PRRs that signal an infection o Include the Toll Receptor Family o Expressed external or internally o Activation of “pro-inflammatory” signaling pathways NFkB and MAP kinase signaling pathways • Antimicrobial peptides (Defensins) / lysozyme, • Inflammatory cytokines (TNFa, IL-8, IL-1) o Regulate activation of adaptive immune response • co-stimulatory molecules Pattern Recognition Receptors (PRRs) ● 2) Phagocytic (endocytic) PRRs o Expressed on the surface of phagocytic cells (MQs, PMNs, DCs) o Mediate uptake of microbe into phagocytes ● 3) Secreted PRRs o Secreted by MQs, epithelial cells, liver o Activate C’, opsonize microbial cells, function as accessory proteins for MAMP recognition Toll-like Receptor Family PPR receptor Toll-like Receptor family ● Found both on the surface and within cells ● First discovered in Drosophila ● Currently 13 receptors o 1-9 mouse & human o 10 human o 11-13 mouse Curr. Opin. Hematology 9:2-10, 2002 Intracellular PRRs: Present in the Cytosol of Host Cells 1. Protein kinase receptor (PKR) ● Activated upon binding to dsRNA (viruses) o Blocks viral & cellular protein synthesis (eIF2a) o Activates NFkB, MAP kinase STATs & IRF signaling pathways o Induces apoptosis & IFNa/b production of infected cells 2. 2’-5’ Oligoadenylate Synthase & RNaseL ● Family of IFN-inducible enzymes o dsRNA activates OAS o RNaseL degrades viral and host RNA o Induces apoptosis Intracellular PRRs: Present in the Cytosol of Host Cells 3. NOD proteins or nucleotide-binding oligomerization domain ● Recognize intracellular peptidoglycan-derived MAMPs and transduce signals ● three distinct functional domains o carboxy-terminal ligand-recognition domain (LRD) o centrally located NOD o amino-terminal effector-binding domain (EBD) CARD domains in mammals Interacts and activates RIP2 inducing NFkB and MAPkinase pathways Structure of NOD Proteins Inohara N, & Nunez G. Nat Rev Immunol. 2003 3:371 NOD Proteins Inohara N, & Nunez G. Nat Rev Immunol. 2003 3:371 Phagocytic (endocytic) PRRs Bind Carbohydrates 1. Macrophage Mannose Receptor (C-type lectin) ● Type 1 transmembrane receptor ● Recognizes patterns of mannose residues in a certain spatial orientation unique to microbes (CRD) ● Only found on macrophages (not monocytes or PMNs) 2. Glucan Receptor (Dectin-1) ● Type 2 transmembrane receptor ● Recognizes b-1,3 & b-1,6 linked glycans ● Present on all phagocytes Phagocytic (endocytic) PRRs: Cont. 3. Scavenger Receptors ● Recognize charged ligands o Polyanionic ligands (ds-RNA, LPS, LTA) o Acetylated low-density lipoproteins (LDL) ● Found on all phagocytes ● MARCO (macrophage receptor with collagenous struction) o binds bacterial cell walls but not yeast ● Phagocytose apoptotic cells o new factor MFG-E8 (released from activated macrophages and binds to apoptotic cells via phosphatidylserine) Secreted PRRs activate the Complement (C’) System Complement system is activated by innate immunity Recognition by Complement receptors (CR) o CR1, CR2, CR3, CR4, C5a, C3a Comprised of plasma proteins that when activated forms a triggered enzyme cascade ● Zymogens – activated by the cleavage of other proteases o Precursor enzymes Function ● Facilitates the uptake & destruction of pathogens by phagocytes ● Induces an inflammatory responses Activation of C’ System b C4b + C2b C3b + Bb Secreted Pattern Recognition Molecules Activation of Complement Opsonization of microbial cells Primarily produced by the liver but can be produced by phagocytes Acute Phase Proteins Secreted Pattern Recognition Molecules 1. Collectins ● Recognizes microbial carbohydrates (CRD domain) ● Effector function mediated by collagenous domain ● Mannan-binding lectin (MBL) o Recognizes patterns of mannose & fucose residues in a certain spatial orientation unique to microbes o Initiates the lectin pathway of C’ cleaving C2 & C4 o Can function as an opsonin Binds a receptor on phagocytes (C1qRp) ● Surfactant proteins (SP-A / SP-D) o lung Collectins • Structure is conserved and similar to other proteins with similar function: o Some Complement proteins & Mannose Binding Protein o Binds to bacteria, fungi & viruses • Function by binding microbes and are important for mediating phagocytosis of alveolar macrophages Collagen helix Microbe a-coiled helix C-type Lectin domain Secreted Pattern Recognition Molecules – Cont. 2. Pentraxin ● Members include o Serum amyloid protein (SAP) o C-reactive protein (CRP) ● Recognize phosphorylcholines on microbes ● Functions as an opsonins ● Binds to C1q & activate classical C’ pathway Secreted Pattern Recognition Molecules – Cont. 3. Lipid Transferases ● LPS binding protein (LBP) o Opsonin ● Bactericidal permeability increasing protein (BPI) o Bactericidal protein 4. Peptidoglycan recognition proteins (PGRS) ● Recognizes peptidoglycans in evolutionarily distant organisms ● 4 human PGRS ● Function is unknown o One has bactericidal effects o Triggers a serine protease cascade in insects ? Complement cascade ? Inflammatory Response Inflammatory Response Leukocyte Adhesion Naïve and Memory T Cells Travel in Different Paths Naïve (have not seen their antigen) T cells travel in the blood and lymphatics Memory T cells (have been activated by their antigen) can also travel through tissues Lymphocyte Trafficking Patterns of Naïve T Cells CD62L:selectin CCR7:chemokine receptor Peripheral Blood HEV GlyCAM-1 Lymphatic system ICAM-1 Peripheral lymph nodes Chemokines CCL21 / SLC CCL19 / ELC (MIP-3b) Lymphocyte Trafficking Patterns of Naïve T Cells CD62L:selectin CCR7:chemokine receptor Peripheral Blood HEV GlyCAM-1 Lymphatic system ICAM-1 Peripheral lymph nodes Chemokines CCL21 / SLC CCL19 / ELC (MIP-3b) Lymphocyte Trafficking Patterns of Effector/Memory T Cells Inflammation CXCR3: chemokine receptor PSGL-1:selectin a4b1:Integrin Peripheral lymph nodes Lymphatic system HEV CD44 ICAM-1 Any Tissue Chemokines CXCL9 / MIG CXCL10 / IP-10 CXCL11 / I-TAC Steps in Lymphocyte Trafficking 1. Tethering 2. Triggering 3. Firm adhesion 4. Diapedesis Blood Vessel Lymphocyte Chemokines Pathogens Stromal cells Endothelial Cell Cytokines Macrophage Phagocytosis Phagocytosis ● Definition: uptake of large particles (>0.5 mm) ● Actin-dependent, clathrin-independent ● High rate & efficiency of internalization Professional phagocytic cells ● Macrophages ● Neutrophils These cells have phagocytic receptors o External receptors FcR, CR3, Mannose receptor o Internal receptors TLRs Macrophages (MQ) Blood - Called monocytes (1-6% WBC) Tissues - Called macrophages ● mature form of monocytes ● normally found in tissues such as gastrointestinal tract, lung, liver and spleen Functions: ● Phagocytose and kills after bactericidal mechanisms are activated (T cells) ● Produce cytokines/chemokines (initiates inflammation) ● Is an antigen presenting cell (co-stim. Molecules) Neutrophils (PMN) Present in blood (55-60% of WBC) Not normally present in tissues Short lifespan - 12 hours Functions: ● First at the site of infection/injury Ingest and kill microbes after bactericidal mechanisms are activated (binding to pathogen) Phagocytosis (MQ & PMN) Active process initiated by binding to pathogen Pathogen is surrounded and then internalized Signaling Interactions during Phagocytosis Ann. Rev. Immunol. 20:825-852, 2002 Killing Mechanisms Phagosome - membrane bounded vesicle that becomes acidified Lysozome - granules that contain products that damage or kill pathogens ● Enzymes o Lysozyme - dissolves cell walls of some bacteria o Acid hydrolases - digests bacteria ● Proteins o Lactoferrin - binds Fe++ needed for bacterial growth o Vitamin B12-binding protein ● Peptides o Defensins and cationic proteins - direct antimicrobials Killing Mechanisms - cont. Respiratory Burst ● ● ● ● Activated following phagocytosis Stimulated by PRR Requires increased oxygen consumption Produces substances that are directly toxic to the bacteria o Oxygen-derived products O2-, H2O2 & Myeloperoxidase o Nitrogen-derived products NO (nitrogen oxide) Produced by inducible NO synthase (iNOS) enzyme Enzyme is induced by cytokines (LT, TNFb) NADPH Oxidase Mitochondrial-independent respiratory burst P47phox & p67phox normally resides in the cytoplasma. P47phox becomes hyperhposphorylated following phagocytosis and binds to p67phox. These components move to the membrane and bind the NADPH complex resulting in an active complex. Enzyme Reactions of Respiratory Burst Respiratory Burst NADPH + 2 O2 NADP+ 2 O- Superoxide dismutase H2O2 Myeloperoxidase ● Enzyme which is stored in primary granules of PMN & MQ and uses the products of the respiratory burst. ● H2O2 + C1Chloramines Professional APC Regulation of Adaptive Response Veterinary Immunology & Immunopathology 91: 1, 2003 T cells Recirculate to “Find” Antigen-loaded Dendritic cells Follicular Area Afferent lymphatics HEV Paracortical Area Germinal Center Efferent lymphatics Mucosal Immunity – Reading Assignment May 10, Spring 2004 Immunobiology (6th Edition) Janeway, Travers, Walpert and Capra Chapter 10 (p. 432-445). Neutra, MR et al Antigen sampling across epithelial barriers and induction of mucosal immune responses. Annu. Rev. Immunol. 14:275-300, 1996 Wright, JR. Immunoregulatory Functions of Surfactant Proteins. Nature Review Immunol. 5:58-68, 2005. Cheroutre, H. Start at the beginning: new perspectives on the biology of mucosal T cells. Annu. Rev. Immunol. 22:217-46, 2004. Weiner, H. Oral tolerance: immune mechanism and the generation of Th3type TGF-beta-secreting regulatory cells. Microbes & Infection 3:947954, 2001.