* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Recovery and analysis of old/ancient DNA: molecular archaeology
Deoxyribozyme wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Gene expression profiling wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Designer baby wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Microevolution wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
RNA silencing wikipedia , lookup
Genetic engineering wikipedia , lookup
Mir-92 microRNA precursor family wikipedia , lookup
Genetically modified crops wikipedia , lookup
Primary transcript wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
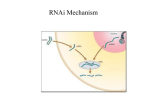
Double-stranded RNA-induced RNA interference causes destruction of a specific mRNA in C. elegans uninjected, no probe uninjected, mex-3 probe antisense mex-3 RNA, mex-3 probe double-stranded mex-3 RNA injected, mex-3 probe Guo, S. and Kemphues, K. J. Cell 81, 611-620 (1995) Fire, A. et al. Nature 391, 809 (1998) Key points of C. elegans experiment •substoichiometric amounts of dsRNA relative to the targeted mRNA are required to completely eliminate the mRNA (i.e. the dsRNA is catalytic) •dsRNA is 10-100X better than antisense or sense RNA •doesn’t work if introns or promoters are targeted by the dsRNA •doesn’t interfere with transcription initiation or elongation (it is possible to target a single gene in an operon) (i.e. RNAi is a post-transcriptional phenomena) •the targeted mRNA is degraded (i.e. it can’t be detected by probes) •dsRNA can cross cellular boundaries (i.e. there is a transport mechanism) Mechanism of RNAi post-transcriptional gene silencing dsRNA 5’ small interfering RNAs 5’ (inactive, 250-500kDa complex) (a critical step in the activation of RISC) RNA-induced silencing complex (active, 100kDa complex) (endonucleolytic cleavage in the region of homology) Zamore, P. D. Science 296, 1265-1269 (2002) Dicer Dicer contains 5 domains • 2 catalytic RNase domains • a dsRNA-binding domain • a helicase domain • a PAZ domain Dicer is thought to work as a dimer One of the catalytic sites in Dicer is defective Thus, instead of cleaving from ~9-11 nt, like bacterial RNase III Dicer cleaves ~22 nt (see panel B) Hannon, G. J. Nature 418, 244-251 (2002) RISC RISC contains at least 4 subunits •Argonaute (5 homologs in Dros.) •dFXR (the Dros. homologue of human fragile X mental retardation protein) •Vasa intronic gene (VIG) •nuclease Activated RISC uses the unwound siRNA as a guide to substrate selection Hannon, G. J. Nature 418, 244-251 (2002) Unanswered question •How does RNAi spread throughout and organism, even when triggered by minute quantities of dsRNA? -Requires a system to pass a signal from one cell to another -Requires a strategy for amplifying the signal RNAi can silence gene expression through other mechanisms: Histone methylation occurs at loci homologous to the siRNA target histone methyltransferase RNA-dependent RNA polymerase Matzke, M. and Matzke, A. J. M. Science 301, 1060-1061 (2003) Schramke, V. and Allshire, R. Science 301, 1069-1074 (2003) Questions about RNAi •What is the endogenous biological function of the RNAi machinery? Gene Regulation Protection from viruses. It could to silence transposons •Is RNAi under negative regulation? how about RNases that digest siRNAs •What are neurons refractory to RNAi? mammals Synthesis of double-stranded RNA (dsRNA) in vitro cut A cut B cut A, SP6 polymerase, NTPs cut A SP6 T7 plasmid 3’ 5’ cut B, T7 polymerase, NTPs SP6 T7 5’ complementary 21-mer RNA oligonucleotides 5’-GGCAAGGUCCAGCUGAACU-3’ ||||||||||||||||| 3’-UACCGUUCCAGGUCGACUU-5’ anneal 5’ 3’ 3’ 5’ 800-1200 bp dsRNA cut B 3’ diploid and polyploid species of Chrysanthemum Gene transfer to plants • Plant callus and cell culture • Agrobacterium (and Rhizobium) mediated transformation • Direct DNA transformation • Plant virus vectors Somatic plant cells have high developmental plasticity (in contrast to most animal cells)--it’s as if they are all embryonic stem cells – Cells from most parts of a plant can regenerate an entire new plant (totipotent) – Plant cells can be cultured in suspension, genetically manipulated, then used to generate transgenic plants Plant transgenics have been used to engineer: – Insect resistance (Bacillus thuringiensis [Bt] toxin) – Microbial pathogen resistance (e.g. overexpression of naturally-occurring plant defense genes, anti-fungal peptides “defensins” – Herbicide tolerance (glyphosate [Roundup] resistance gene) – Improved nutritional value (“golden rice” containing 1 bacterial and 2 daffodil genes for Vitamin A production). Plant tissue culture • “explant” (tissue sample) taken, disinfected • With correct balance of plant hormones, a “callus” will form • Callus transferred to liquid medium and agitated to yield a cell culture • Single cells can be grown on plates to yield new calluses Regeneration of plants from cell culture • Change phytohormone levels to get differentiation – High auxin, roots develop – High cytokinin, shoots develop – In both cases plant formation can be induced OR • Induce somatic embryogenesis: under specific conditions cells produce embryo-like structures that can develop into fertile plants Plant cell transformation • Infection by Agrobacterium tumefaciens and its relatives • Chemical transformation of protoplasts (cells lacking cell walls) • Particle bombardment • Viral infection Agrobacterium transformation • Plant tumors (crown gall disease) induced by bacterium Agrobacterium tumefaciens • Tumors induced by Ti plasmids (140-235 kb) transferred to the plant cells by bacterium (Ti, Tumor inducing plasmids) • Part of plasmid (T-DNA, 23 kb) integrates into plant genome (randomly): confers unregulated growth (hence a tumor) and directs synthesis of “opines” • Opines provide food (carbon and nitrogen source) for bacterium, tumor provides a home, what a deal A typical Ti plasmid • Virulence genes are responsible for T-DNA transfer--induced following bacterial attachment to plant wound • T-DNA is flanked by “border sequences” (25 bp imperfect direct repeats) involved in the transfer process: the right-hand border sequence is sufficient for transfer • T-DNA is excised by Vir gene products and transferred to plant cell via a conjugative pilus T-DNA genes • Encode phytohormones to promote unregulated growth (oncogenes) • These oncogenes can be deleted to “disarm” the T-DNA • Disarmed T-DNA sequences are used for transformation An early ‘disarmed’ Ti plasmid Transfer could be screened for by opine synthesis (product of the nos gene) Manipulating the Ti plasmid • Very large--difficult to manipulate in vitro • Transgene-containing T-DNA can be created by recombination in Agrobacterium: T-DNA Manipulating the Ti vector • Alternative approach: use two plasmids – One with genes for virulence (“helper” plasmid) – The other with T-DNA sequences--smaller, can use classical cloning techniques Selection for T-DNA transfer • • Drug resistance (e.g. aminoglycoside antibiotics) Herbicide resistance (e.g. glyphosate [Roundup]) • Concern over potential harm (to health and/or environment) from these markers has driven development of other methods – manA gene: confers growth on the sugar mannose as a sole carbon source – Use of cre-lox mediated deletions to remove markers from transgenic plants Agrobacterium-mediated transformation Kanamycin: selection for T-DNA transfer Carbenicillin: kills Agrobacteriu New developments: not just Agrobacterium can transfer genes via a T-DNA vector Closely related bacteria (Rhizobium species) can do the same This circumvents many patents, and researchers behind this work have made the technology freely available www.bioforge.net, see the “Transbacter” section Fig. 10.25. Genetically engineered Direct DNA transfer to plants • Protoplast transformation – – – – – Make protoplasts (plant cells lacking cell walls) Addition of DNA in the presence of polyethylene glycol Electroporation Transformants selected on the basis of marker genes Regenerate whole plants Direct DNA transfer to plants • Particle bombardment – Particles coated with transforming DNA, fired through plant cells (and nuclei) – Valuable method for transforming plants that cannot be transformed by previous methods – Works with cell cultures, embryos, leaves, etc. Chloroplast transformation • Many chloroplasts in each cell (high copy number and expression of transgene) • Chloroplasts are not transmitted by pollen (easier to contain the transgene) • Chloroplast transgenes are not subject to position effects on transgene expression • Target integration into chloroplasts using chloroplast homology regions Plant viruses as vectors • Naturally transforming • High-level transgene expression • No integration into host chromosome • Mostly used for expression of foreign proteins • DNA viruses • RNA viruses – Most plant viruses are RNA viruses – cDNA copies of the viral genome are used for engineering Terminator technology Protection of technology and market share: “technology protection” (Monsanto) “terminator technology” (critics) Farmers historically save a small proportion of seeds from this year’s crop for next year’s crop Transgenic seeds: buy them once, never buy them again because of saved seeds Unless those transgenic plants produce sterile seeds How to produce sterile seeds on demand: Ribosome inactivating protein (RIP) under the control of a late embryonic development induced promoter, remove transcription terminator by Cre/loxP system Plants produce viable seeds x stop RIP loxP loxP RIP No viable seeds (the seeds are soaked in Tet before sale) Syngenta and the release of Bt-10 Bt-11: approved by govt. for sale to farmers Bt-10: not approved… differs from Bt-11 in that it has a marker gene (conferring ampicillin resistance) $375,000 fine for Syngenta Embarrassment (damage?) to the US biotech industry Issues with recombinant plants How will the transgene affect: 1. Ecosystem health a. Will the plant be more invasive? b. Will the plant transgenes transmit to the native plant population c. Will the plant harm beneficial animals and insects? d. How quickly will resistant strains of pathogens arise? 2. Human health is recombinant DNA-containing plant tissue safe to eat over the long term? 3. Economics: will the choice to not grow transgenics harm farmers? • US govt. website on agricultural biotechnology regulations: http://www.aphis.usda.gov/brs/index.html • The Onion and recombinant broccoli