* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download ImageSurfer: Visualization of Dendritic Spines
Feature detection (nervous system) wikipedia , lookup
Biological neuron model wikipedia , lookup
Convolutional neural network wikipedia , lookup
Neuroanatomy wikipedia , lookup
Stimulus (physiology) wikipedia , lookup
Synaptic gating wikipedia , lookup
Nervous system network models wikipedia , lookup
Synaptogenesis wikipedia , lookup
Visual servoing wikipedia , lookup
Holonomic brain theory wikipedia , lookup
Apical dendrite wikipedia , lookup
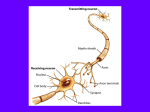
ImageSurfer: Visualization of Dendritic Spines CISMM: Computer Integrated Systems for Microscopy and Manipulation The Problem: Finding Relationships Scientific Domain Neurons transmit information using electrochemical reactions. When a neuron fires, it sends an electrical impulse down a long arm-like structure called an axon. At the end of the axon the impulse sets off a chemical transfer. The chemicals diffuse across a gap to dendrites, on neighboring receiver neurons. These neurons then fire their own electrical impulses thus propagating the signal. Dendritic spines are tiny structures (0.5-2 m length, 0.1 m diameter) that protrude along the length the dendrites. These small bodies are shaped like miniature light bulbs with distinct head and neck regions. In addition, these spines may play a role in insulating specific regions of the dendrite to signal molecules from neighboring neurons at the cellular level and might even be the basis of memory. Scientific Questions 1. How is the concentration of PMCA distributed within dendritic spines? 2. In a given spine, where is the greatest concentration of PMCA? Where is the least concentration? Data Collection Images of the sample are taken using a confocal microscope. A single slice of the sample is illuminated with a wavelength that causes the a phosphorescing tag named DiO (dioctadecyloxacarbocyanine perchlorate) to glow and an image is captured. The phosphorescing DiO reveals the lipid layor of the dendrites. The sample is then illuminated with another wavelength that causes the PMCA (plasma membrane calcium ATPase) antibody to phosphoresce and another image is captured. This process yields two images for every slice. The resulting image pairs show the locations and concentration of PMCA proteins as well as the general structure of the nerve cells. Figure 1: Former Method, images of a single slice of DiO (left image) and PMCA (right image). The collaborators would flip between these two datasets to find relationships between the data. In addition to being time consuming, only the spines that fell into the image plane could be analyzed in this technique. The Solution: Visualizing Both Datasets Colored Isosurface The isosurface is a 3D reconstruction of the DiO dataset. The surface structure exhibits the shape of the dendritic spine and color exhibits the concentration of the protein, PMCA, at the surface. Magic Cut Plane Spline Tool This tool can be thought of as a cross between a magic lens and a cut plane. The magic cut plane enables capturing spines at arbitrary angles. This increases the number of spines that can be analyzed beyond those that lie in the image plane. The spline tool enables users to capture concentrations of PMCA and DiO along a user-defined curve. Height Field The magic cut plane captures a slice from the DiO and PMCA datasets and maps the PMCA concentration to a height field. The color indicates DiO, thus providing an outline of the dendrite. Graph After the DiO and PMCA are captured, the graph window displays the concentrations as a function of distance from the start of the spline. The red line indicates the DiO and the green line indicates PMCA. http://www.cs.unc.edu/Research/nano/cismm/ Collaborators: Alain Burette and Richard Weinberg, UNC Cell Development and Biology. Project Lead: Russell Taylor . Investigator: Dennis Jen. December 2003