* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Exam 1, Problem 6
Nucleic acid analogue wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Genome evolution wikipedia , lookup
Transcriptional regulation wikipedia , lookup
List of types of proteins wikipedia , lookup
Gene expression wikipedia , lookup
Gene desert wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Protein moonlighting wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Gene expression profiling wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Point mutation wikipedia , lookup
Gene regulatory network wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Non-coding DNA wikipedia , lookup
Community fingerprinting wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
Molecular evolution wikipedia , lookup
Welcome to
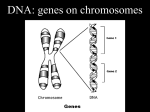
Introduction to Bioinformatics
Wednesday, 13 April 2005
Rehash of Exam 1 (selected)
Rehash of Exam 2 (selected)
Discussion of DGPB, Chapter 6
Exam 1, Problem 6
6c. Find first nucleotides of genes that don’t encode protein.
(LOAD-SHARED-FILE
noncoding-genes-of)
(DEFINE "nc-genes" (NONCODING GENES OF S7942)
(FOR EACH (gene IN nc-genes))
WITH beginning = SEQUENCE OF gene 1 – 3
DO COLLECT beginning
(DEFINE variable AS value)
Exam 1, Problem 6
6c. Find first nucleotides of genes that don’t encode protein.
(LOAD-SHARED-FILE "noncoding-genes-of")
(DEFINE
nc-genes
AS (NONCODING-GENES-OF S7942)
(FOR-EACH gene IN nc-genes
AS
beginning = (SEQUENCE-OF gene FROM 1 TO 3)
COLLECT beginning)
(DEFINE variable AS value)
Exam 1, Problem 6
6c. Find first nucleotides of genes that don’t encode protein.
(LOAD-SHARED-FILE "noncoding-genes-of")
(DEFINE
nc-genes
AS (NONCODING-GENES-OF S7942)
(FOR-EACH gene IN nc-genes
AS
beginning = (SEQUENCE-OF gene FROM 1 TO 3)
COLLECT beginning)
:: ("GCG" "GCG" "GGA" "GCC" "GCC" "GGA" "GCG"
"GGG" "GCC" "GGG" "GCG" "GCC" "GCG" "GGA" "GGG"
"GCG" "GGG" "TCC" "GGT" "GGG" "GGG" "AAA" "GGA"
"CCA" "TCC" "GGC" "GGC" "CGC" "CGG" "GGG" "GGG"
"GCG" "AAA" "GGG" "GGG" "GGT" "TCC" "GGC" "TGG"
"GGG" "GCG" "GGG" "GCC" "GGG" "GCC" "GGG" "CGG"
"CGG" "GGG" "GCG" "GGG" "GGG")
Exam 1, Problem 8
8. Thermophilus extremus G+C% content = 80%.
8a. Frequency of cutting of MseI (TTAA)
G+C=
A+T=
A=
T=
0.8
0.2
0.1
0.1
Expected frequency of TTAA
= 0.1 * 0.1 * 0.1 * 0.1
= 10-4
Exam 1, Problem 8
8. Thermophilus extremus G+C% content = 80%.
8d. Test answer with 1000 random DNA sequences
1000-nucleotides in length (G+C% = 80%)
(FOR-EACH iteration FROM 1 TO 1000
AS seq
= (RANDOM-DNA A 1 T 1 C 4 G 4 LENGTH 1000)
AS counts = (COUNT-OF "TTAA" IN seq)
SUM counts)
:: 103
??? hits per trial?
Exam 1, Problem 8
8. Thermophilus extremus G+C% content = 80%.
8d. Test answer with 1000 random DNA sequences
1000-nucleotides in length (G+C% = 80%)
(FOR-EACH iteration FROM 1 TO 1000
AS seq
= (RANDOM-DNA A 1 T 1 C 4 G 4 LENGTH 1000)
AS counts = (COUNT-OF "TTAA" IN seq)
SUM counts)
:: 103
0.103 hits per trial?
Exam 1, Problem 8
8. Thermophilus extremus G+C% content = 80%.
8e. Test answer with 1000 random DNA sequences
3000-nucleotides in length (G+C% = 80%)
(FOR-EACH iteration FROM 1 TO 1000
AS seq
= (RANDOM-DNA A 1 T 1 C 4 G 4 LENGTH 3000)
AS counts = (COUNT-OF "TTAA" IN seq)
SUM counts)
:: 314
??? hits per trial?
Exam 1, Problem 8
8. Thermophilus extremus G+C% content = 80%.
8e. Test answer with 1000 random DNA sequences
3000-nucleotides in length (G+C% = 80%)
(FOR-EACH iteration FROM 1 TO 1000
AS seq
= (RANDOM-DNA A 1 T 1 C 4 G 4 LENGTH 3000)
AS counts = (COUNT-OF "TTAA" IN seq)
SUM counts)
:: 314
0.314 hits per trial?
Exam 1, Problem 8
8. Thermophilus extremus G+C% content = 80%.
8f. Interpret your results in light of the definition
of E-value (or Expect value).
Your results:
E-value
Expected frequency = 10-4
1000 1000-nucleotides DNA sequences
0.103 per trial
1000 3000-nucleotides DNA sequences
0.314 per trial
E-value = (expected frequency) · (search space)
0.1
0.3
Exam 1, Problem 10
Examine Fig. 4.11 in the text, focusing on the spot labeled
TDH1.
Exam 1, Problem 10
Examine Fig. 4.11 in the text, focusing on the spot labeled
TDH1.
10b. From what you can learn of the function of the gene,
why might this result make sense?
glucose
glycolysis
Glyceraldehyde-3phosphate
dehydrogenase
pyruvate
gluconeogenesis
Exam 2, Problem 4
Consider Chi-Squared.
4a. Define a function that calculates chi-squared
scores, given two input arguments…
(DEFINE-FUNCTION chi-square (observed expected-freqs)
(LET* ((total (SUM-OF observed))
(expected
(FOR-EACH freq IN expected-freqs
COLLECT (* freq total))))
(FOR-EACH O IN observed
FOR E IN expected
AS diff = (- O E)
AS numerator = (* diff diff)
SUM (/ numerator E))))
Exam 2, Problem 4
Consider Chi-Squared.
4b. How do you interpret the 1.44 result from my
example?
The probability of getting a value of 1.44 is likely to occur in the
gene 100-nt population
Exam 2, Problem 4
Consider Chi-Squared.
4b. How do you interpret the 1.44 result from my
example?
This means that there is a > 5% chance that the population given fits
the expected ratios.
Exam 2, Problem 4
Consider Chi-Squared.
4b. How do you interpret the 1.44 result from my
example?
there is a 5% chance that the A:C:G:T ratio of 28:22:28:22 is due to
random chance.
Exam 2, Problem 5
5h. Rerun the program you wrote in 5d but using a single
population: random DNA sequences.
DGPB 6.1 Associating proteins with functions
UPTAG
DOWNTAG
AGTCGT…TGTAACG…CGTGC… AGTCGT…CATGGGA…CGTGC…
gene
DGPB 6.1 Associating proteins with functions
DGPB 6.1 Associating proteins with functions
DGPB 6.1 Associating proteins with functions
DGPB 6.1 Associating proteins with functions
Sampling problem