* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Slide 1
Pharmacogenomics wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Gene expression profiling wikipedia , lookup
Nutriepigenomics wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Genetic testing wikipedia , lookup
Genetic drift wikipedia , lookup
Koinophilia wikipedia , lookup
Gene expression programming wikipedia , lookup
Behavioural genetics wikipedia , lookup
Genealogical DNA test wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Human genetic variation wikipedia , lookup
Population genetics wikipedia , lookup
Genetic engineering wikipedia , lookup
History of genetic engineering wikipedia , lookup
Public health genomics wikipedia , lookup
Genome (book) wikipedia , lookup
Heritability of IQ wikipedia , lookup
Designer baby wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
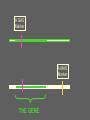
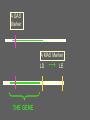
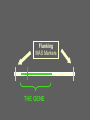
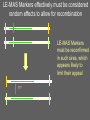
Multiple-Trait Selection in a Single-Gene World David Notter Department of Animal and Poultry Sciences Virginia Tech Genetic Markers and NCE • Genetic Markers have the potential to improve the effectiveness of NCE • However, for most traits, genetic markers will not account for enough of the genetic variation to allow them to be used as the only selection criterion • Instead, methods must be developed to combine information on genetic markers with performance data Types of Marker-Assisted Selection • Gene-Assisted Selection (GAS) – A DNA sequence variant exists within the gene – May be the actual causal mutation or just associated with it • Linkage-Disequilibrium MAS (LD-MAS) – Marker is not a part of the gene, but is very tightly linked with the favorable form of the QTL • Linkage-Equilibrium MAS (LE-MAS) – Marker is loosely linked to the QTL. The association can differ among families (sires) Garrick and Johnson, 2003 A GAS Marker A MAS Marker THE GENE A GAS Marker A MAS Marker LD THE GENE → LE Flanking MAS Markers THE GENE Judging the Importance of a Marker • The Size of Marker Effect – How different are the different marker genotypes? • Degree of Dominance of the Marker – Are heterozygotes intermediate or do they resemble one of the homozygotes? • The Marker Frequency – Is the marker common or rare within a breed? • The Proportion of the Genetic Variance in the Trait Accounted for by the Marker – How much variation (opportunity for improvement) exists independent of the marker? The Marker Effect Genotype MM Effect +a Marker mm -a = 2a Effect For the GeneStar marbling (thyroglobulin) marker, the difference in marbling score between homozygotes is 3.5 to 11% (Hetzel, 2003). In Angus cattle (AAA, 2004), with a mean marbling score of about 6.0, this would give 2a = 0.48, or about one half of a marbling score. Degree of Dominance of the Marker Genotype MM Effect +a Mm Mm Mm +a → Dominant marker 0 → Co-dominant marker -a → Recessive marker mm -a The GeneStar marbling marker is approximately codominant in its effect on marbling Genestar Genetic Marker Percentage Choice Percent Choice 80 70 60 57 60 64 50 40 30 No Stars One Star No. of "Stars" Two Stars Genetic Variance Accounted for by a Codominant Marker 2 σ 2 h M A-M = = 2 2p(1-p)a 2 [2p(1-p)a ] / 2 σ P • h2M is the marker heritability • a is the marker effect • p is the frequency of the marker • σ2P is the phenotypic variance Frequency of the Marker p 0.50 h2 0.50 h2M 0.125 h2M = .25 h2 at p = .50 Frequency of the Marker p 0.50 h2 0.50 h2M 0.125 0.75 0.48 0.100 h2M = .25 h2 at p = .50 Frequency of the Marker p 0.50 h2 0.50 h2M 0.125 0.75 0.48 0.100 0.90 0.46 0.050 For GeneStar Marbling with p = 0.50, h2M ~ 0.04, which accounts for 11% of the additive variance in marbling score. Thus 89% is associated with other, currently unidentified, genes. Frequency of the GeneStar Marbling Marker in Various Breeds Breed B. Angus Frequency* 0.30 R. Angus 0.45 Simmental 0.29 Wagyu 0.65 * From Hetzel, 2003--Approximate Overview of Issues Involved in Marker Assisted Selection 1. The size of the effect: what is the difference (2a) between individuals homozygous for alternative marker alleles? • Must be estimated and validated 2. The importance of the effect: what is the economic effect of a change in marker genotype? 3. The mechanism of gene action: is the marker dominant, recessive, or co-dominant? Overview of Issues Involved in Marker Assisted Selection 4. The importance of other genes: compare the marker heritability (h2M) to the overall h2 to determine the need for continued data recording and gene discovery. 5. The frequency of the favorable marker: • • • Frequencies near 0.5 support the most rapid and immediate improvement High frequencies imply limited impact Low frequency markers result in a lag period, and have lots of potential, but raise concern about loss of genetic diversity and impact on other traits Integrating Marker Information into National Genetic Evaluations • Genes and markers will continue to be discovered • Many will not be of general utility, but some will be useful • Comprehensive genotyping of many animals may be possible but is not yet a reality • Partial genotyping of subsamples of animals is more realistic for the immediate future. How Might Breed Associations Respond? • We are effectively being told that there is something outside NCE that makes an animals better or worse than his EPDs might indicate • Yet for proven sires, the EPD is a more definitive predictor of progeny performance and genetic worth – Markers are valuable mainly for young (unproven) animals, for traits not included in the EPDs, or for traits that take a long time to evaluate accurately How Might Breed Associations Respond? • Explicitly identify the genes and markers of interest to the breed • Develop a DNA collection strategy • Develop a genotyping strategy • Develop validation strategies • Incorporate marker information into NCE How Might Breed Associations Respond? Explicitly identify the genes and markers of interest to the breed • Identify the known genes and LD markers of interest to the breed • Might also identify a set of informative microsatellite markers for use in gene discovery • This will be an evolving array, but provides guidance for the genes and markers that will be supported in NCE How Might Breed Associations Respond? Develop a DNA collection strategy • Evaluate simple techniques for DNA acquisition and physical storage: fluoroacetate papers, hair, etc. • Don’t extract DNA until you need it. • Capacity for repeated extractions. • Identify high-priority animals, but don’t necessarily rule out storage of (for example) blood on all registered or performance-recorded animals Blood samples on Perforated FTATM Cards How Might Breed Associations Respond? Develop a genotyping strategy • Breed associations need to ensure access to genotypes on their animals and become repositories for those genotypes • “Multiplex” genotyping capacity is needed to allow efficient genotyping of individual animals for many genes/markers • Develop a genotyping plan for high-use (“legacy”) sires, and perhaps samples of their calves (i.e., to screen for segregating markers) How Might Breed Associations Respond? Develop validation strategies • New markers must be validated to determine if initial results are repeatable • New markers must be validated in different breeds • Markers must be validated for both the primary trait and for correlated traits • Genotyping strategies can be designed to support validation strategies How Might Breed Associations Respond? Incorporate marker information into NCE • We know very little about how this will happen! • We do know that marker information will continue to evolve—we will always be behind! • Must be able to continuously incorporate new markers into NCE • Marker information will enhance, but certainly not replace, performance data and EPDs How Might Breed Associations Respond? Take Control of the Use of Genetic Markers in NCE • Knowledge and resources to allow breeders and their organizations to impact marker detection and development • Rapid evaluation of frequencies of new genetic variants and markers • Rapid and efficient validation of newly proposed markers How Might Breed Associations Respond? Issues in the Incorporation of Marker Information into NCE • Are marker effects “fixed” or “random”? • What is the genetic base for a marker effect? • What are the effects of a marker on other traits? How do we estimate these accurately? • How do we validate marker effects in different environments and management systems? • How do we check if a marker “stops working”? • How to handle animals that are not genotyped or genotyped for only a few markers? Using Markers in NCE Are the marker effects “fixed” or “random”? • If fixed, then we make a constant adjustment to the EPD based on marker genotype • If random, then even if we know the genotype exactly, we still hedge our bets to allow for recombination, interactions of the marker with the environment or the background genotype, or other unknown variations in the gene • But HOW do we hedge? Estimate sire x marker or marker x environment interaction variances? A GAS Marker can, at least hypothetically, be considered a fixed effect (but somehow it seems too simple!) A GAS Marker THE GENE Genotype MM Mm Mm EPD +α d -α A GAS Marker THE GENE Other, unknown sequence variants could be present in some animals and invalidate the effects of the known marker. We need to prepare ourselves for things like this! LE-MAS Markers effectively must be considered random effects to allow for recombination LE-MAS Markers must be reconfirmed in such sires, which appears likely to limit their appeal ??? Using Markers in NCE What is the genetic base for a marker effect? • Depends on, and changes with, the frequency of the marker • As marker approaches fixation, the favorable form becomes less and less useful. Using Markers in NCE What are the effects of a marker on other traits? How do we estimate these accurately? • Major validation issue • We will immediately credit an animal for the known, favorable marker effect, but only slowly identify that animal as possibly inferior for correlated effects • Linkage with performance records is mandatory, as is adequate genotyping of offspring of both sexes Using Markers in NCE What about animals that are not genotyped or are genotyped for only a few markers? • Many animals will likely not be genotyped • We will therefore need to infer the possible genotypes of such animals using the genotypes of their relatives • Thallman has developed one methodology to accomplish this—there may be others Conclusions • The search for markers will continue • The bovine gene map will accelerate the search for and the rate of discovery of genetic markers • BIF is facing a developmental effort to use these DNA technologies that may rival the implementation of BLUP EPDs • The BIF Guidelines are going to get thicker again!