* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Bioinformatics - Department of Computer Science
Multi-state modeling of biomolecules wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Molecular cloning wikipedia , lookup
Genetic code wikipedia , lookup
Whole genome sequencing wikipedia , lookup
Community fingerprinting wikipedia , lookup
List of types of proteins wikipedia , lookup
Genome evolution wikipedia , lookup
Gene expression wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Homology modeling wikipedia , lookup
Non-coding DNA wikipedia , lookup
Protein structure prediction wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Biochemistry wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
History of molecular evolution wikipedia , lookup
CS369 Computational Biology
University of Auckland
Course coordinator: Alexei Drummond
CS369 Computational Biology
• This course provides an overview of algorithms
and scientific computing techniques in
computational biology and bioinformatics.
• A hands-on introduction to topics including
–
–
–
–
–
–
CS369 2007
Dynamic programming and string algorithms,
Markov models, Hidden Markov models,
Heuristic search algorithms,
Pairwise and multiple sequence alignment,
Phylogenetic Reconstruction
Gene finding and secondary structure prediction
2
Your lecturers
Alexei Drummond
Michael Dinneen
Patricia Riddle
Coordinator
Algorithmics
Learning and
pattern discovery
CS369 2007
3
Recommended Texts
• Bioinformatics: Sequence and Genome
analysis, 1st & 2nd Editions
– by David W Mount (2001; 2004)
– 1st Edition is available as an E-book in the library
– 2nd edition is available on short term loan
• Biological sequence analysis
– by Durbin, Eddy, Krogh and Mitchinson (1998)
– Available on short term loan
• Algorithm Design
– by Kleinberg J. and Tardos E (2006)
CS369 2007
4
Assessment
• 2 x 2 hour labs (3rd April and 29th May)
– 5% each
• 1 assignment (1st May handout)
– 20%
• 1 Mid-semester test
– 10%
• 1 Exam
– 60%
CS369 2007
5
Lecture schedule - first half
CS369 2007
6
Lecture schedule - second half
CS369 2007
7
What is computational biology?
• Computers and
biology?
• the organization and
the analysis of
biological data with
computers?
• The new biology of the
21st century?
CS369 2007
8
What is bioinformatics?
• Rapidly growing biological
databases contain information
about
– Cellular and molecular biology
– Ecology and Evolutionary
biology
– Microbiology
– Genomics
– Proteomics
• The application of
computational biology to
understand this data.
CS369 2007
9
What is computational biology?
• Computational biology combines the tools and techniques
of
– mathematics,
– statistics,
– computer science,
– biology
• in order to understand the masses of biological data
CS369 2007
10
Computational biology
CS369 2007
and the tree of life
11
CS369 2007
12
The human 18S gene
CS369 2007
QuickTime™ and a
TIFF (Uncompressed) decompressor
are needed to see this picture.
13
The frog 18S gene
CS369 2007
QuickTime™ and a
TIFF (Uncompressed) decompressor
are needed to see this picture.
14
Pairwise alignment of human and frog
CS369 2007
Humans and Frogs are 94% similar!
15
Pairwise sequence alignment
Sequences
x=acggts
y=awgcctt
Alignment
x = a – c g g – t s
y = a w – g c c t t
CS369 2007
16
Dynamic programming algorithm
• computation is carried out bottom-up
• store solutions to subproblems in a table
• all possible subproblems solved once each, beginning
with smallest subproblems
• work up to original problem instance
• only optimal solutions to subproblems are used to
compute solution to problem at next level
• DO NOT carry out computation in recursive, top-down
manner
– same subproblems would be solved many times
CS369 2007
17
Principle of Optimality
Auckland
Te Kuiti
Wellington
CS369 2007
18
Scoring
• Numeric score associated with each column
• Total score = sum of column scores
• Column types:
(1) Identical (+ve)
(2) Conservative (+ve)
x = a – c g g – t s
y = a w – g c c t t
(3) Non-conservative (-ve)
CS369 2007
(4) Gap (-ve)
19
Comparing many species
CS369 2007
This is called a “multiple sequence alignment”
20
QuickTime™ and a
TIFF (Uncompressed) decompressor
are needed to see this picture.
QuickTime™ and a
TIFF (Uncompressed) decompressor
are needed to see this picture.
QuickTime™ and a
TIFF (Uncompressed) decompressor
are needed to see this picture.
QuickTime™ and a
TIFF (Uncompressed) decompressor
are needed to see this picture.
QuickTime™ and a
TIFF (Uncompressed) decompressor
are needed to see this picture.
QuickTime™ and a
TIFF (Uncompressed) decompressor
are needed to see this picture.
QuickTime™ and a
TIFF (Uncompressed) decompressor
are needed to see this picture.
QuickTime™ and a
TIFF (Uncompressed) decompressor
are needed to see this picture.
CS369 2007
21
Tree of 3000 species
CS369 2007
22
Molecular tracking of the
Southern Saltmarsh Mosquito
extract
multiply
decode
A C G T T A C C G T A A G T G
build tree
CS369 2007
align
23
Bioinformatics and genome
evolution
QuickTime™ and a
TIFF (Uncompressed) decompressor
are needed to see this picture.
CS369 2007
24
Computational biology and
disease
• Bacterial infection can
causes serious
illnesses, e.g.
– Necrotising Fasciitis
(“Flesh eating
disease”)
– Toxic Shock Syndrome
(TSS)
• (e.g. the illness Lana Cockroft
got from the cut in her foot
during “Celebrity Treasure
Island”).
CS369 2007
T: Staphylococcus
aureus
B: Streptococcus
pyogenes,
25
Computational biology and
Disease
Necrotizing fasciitis
CS369 2007
26
Computational biology and
Disease
CS369 2007
Identify protein structure of superantigens
- so that potential drug targets can be uncovered.
27
Computational biology and sequencing
technology
• “Old” Technology
–
–
–
–
CS369 2007
1977
Small Scale
Slow
Length: long ≈ 700bp
• New Technology: 454
–
–
–
–
–
2003
Large scale: Whole genome
Fast
Length: short ≈ 100bp
http://www.454.com
28
Bring the mammoth back
Indian
Mammoth
African
Using this new sequencing technology and
bioinformatics to compare the sequences to
the African elephant genome,
scientists sequenced 28 million nucleotides
of the mammoth genome from frozen bones!
CS369 2007
29
Where does the DNA come from?
• Scientists Sequence DNA
of Woolly Mammoth
(extinct 10,000 years ago)
– http://www.science.psu.edu/
alert/Schuster12-2005.htm
QuickTime™ and a
TIFF (Uncompressed) decompressor
are needed to see this picture.
CS369 2007
30
Conclusion
• Computational biology and bioinformatics are fast
becoming central to the biological sciences
–
–
–
–
–
Ecology
Evolutionary biology (tree of life)
Health and disease (flesh-eating bacteria)
Molecular biology and cell biology
Personalized medicine
• Computational biology is heavily algorithmic and often
statistical, drawing from the ‘hard-as-in-rock’ sciences like
mathematics, computer science and statistics - this type of
knowledge is becoming increasingly important in biology.
CS369 2007
31
Reading
• Chapter 1 “Biological Sequence Analysis”,
Durbin et al
• Chapter 1 of Mount
– Don’t worry about any biological terms that you
don’t immediately understand! This chapter is
more to give you a sense of where we are
going to go. The pertinent details will be
explained along the way.
CS369 2007
32
Biomolecules and the Central
Dogma of Molecular Biology
The central dogma of molecular
biology
Replication
Transcription
DNA
CS369 2007
Translation
RNA
Protein
34
What is a macromolecule?
• A chain molecule made up of a large number of
repeating units.
– Homo-polymers are made up of the same units
– Hetero-polymers are made up of a number of different
units
• Nylon, Polyester, DNA (poly-nucleotide) and
Protein (poly-amino acid) are examples
• Macromolecular 3D structure is a question
conformational change rather than breaking and
forming strong bonds.
CS369 2007
35
Biomolecular sequences
DNA
5’-ACGATCGACTGGTATATCGATGCT-3’
Xi {A,C,G,T}
RNA
Protein
CS369 2007
5’-ACGAUCGACUGGUAUAUCGAUGCU-3’
Xi {A,C,G,U}
MFINRWLFSTNHKDIGTLYLLFGAW
Xi {A,R,N,D,C,E,Q,G,H,I,L,K, M,F,P,S,T,W,Y,V}
36
DNA
5’-ACGATCGACTGGTATATCGATGCT-3’
3’-TGCTAGCTGACCATATAGCTACGA-5’
A-T
Watson-Crick
Base pairing
In nature DNA exists in a
double helix of two
complementary
sequences.
G-C
CS369 2007
37
RNA
RNA
5’-ACGAUCGACUGGUAUAUCGAUGCU-3’
G-C
A-U
3 H bonds
G-U
1 H bonds
3D structure
CS369 2007
2 H bonds
Secondary structure
38
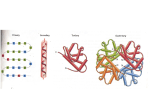
Proteins
• Proteins are hetero-polymers made
up of 20 standard amino acids and
are “coded for” by genes.
• They form 3D molecular structures
that do work in the cell.
• An open problem is to determine
structure computationally from the
primary sequence of amino acids.
A cartoon of the 3D structure of
the myoglobin protein
CS369 2007
39
The genetic code
Amino acids
CS369 2007
40
The genetic code
CS369 2007
41
CS369 2007
42
CS369 2007
43