* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download document
Deoxyribozyme wikipedia , lookup
Adenosine triphosphate wikipedia , lookup
Fatty acid metabolism wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Catalytic triad wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Fatty acid synthesis wikipedia , lookup
Butyric acid wikipedia , lookup
Citric acid cycle wikipedia , lookup
Proteolysis wikipedia , lookup
Metalloprotein wikipedia , lookup
Point mutation wikipedia , lookup
Peptide synthesis wikipedia , lookup
Protein structure prediction wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Epitranscriptome wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Biochemistry wikipedia , lookup
Genetic code wikipedia , lookup
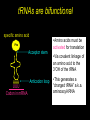
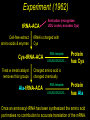
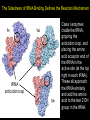
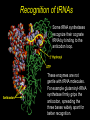
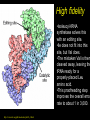
Chapter 32 The “2nd Genetic Code” Pages 1075 to 1086 Learning objectives: Understand the following • The Raney nickel experiment • The class differences of aminoacyl tRNA synthetases? • How aminoacyl tRNA synthetases recognize their cognate tRNA • Crick’s Wobble Hypothesis All rights reserved. Requests for permission to make copies of any part of the work should be mailed to: Permissions Department, Harcourt Brace & Company, Sea Harbor Drive, Orlando, Florida 32887-6777 6277 tRNAs are bifunctional specific amino acid Phe Acceptor stem Anticodon loop AAA UUU Codon in mRNA •Amino acids must be activated for translation •Via covalent linkage of an amino acid to the 3’OH of the tRNA •This generates a “charged tRNA” a.k.a. aminoacyl-tRNA tRNA activation must be specific •When a ribosome pairs a "CGC" tRNA with "GCG"codon, it expects to find an alanine carried by the tRNA. It has no way of checking. •This is because codons in the mRNA are recognized by the anti-codon loop in the aminoacyl-tRNAs •The delivery of the amino acid is specified by this codon-anticodon interaction (regardless of which amino acid is attached to the tRNA) HOW DO WE KNOW THIS??? From:http://www.rcsb.org/pdb/molecules/pdb16_1.html Experiment (1962) tRNA-ACA Cell-free extract amino acids & enymes tRNA is charged with Cys Cys-tRNA-ACA Treat w metal catalyst removes thiol groups (Raney nickel) Anticodon (recognizes UGU codon, encodes Cys) RNA template UGUGUGUGUG... Charged amino acid is changed chemically Ala-tRNA-ACA Protein has Cys Experiment (1962) tRNA-ACA Cell-free extract amino acids & enymes tRNA is charged with Cys Cys-tRNA-ACA Treat w metal catalyst removes thiol groups Anticodon (recognizes UGU codon, encodes Cys) RNA template UGUGUGUGUG... Protein has Cys Charged amino acid is changed chemically Ala-tRNA-ACA RNA template UGUGUGUGUG... Protein has Ala Once an aminoacyl-tRNA has been synthesized the amino acid part makes no contribution to accurate translation of the mRNA. tRNA activation must be specific •The delivery of the amino acid is specified by this codon-anticodon interaction (regardless of which amino acid is attached to the tRNA) •Each tRNA is matched with its amino acid long before it reaches the ribosome. •The match is made by a collection of remarkable enzymes, the aminoacyl-tRNA synthetases. •These enzymes charge each tRNA with the proper amino acid, thus allowing each tRNA to make the proper translation from the genetic code of DNA into the amino acid code of proteins. From:http://www.rcsb.org/pdb/molecules/pdb16_1.html Aminoacyl-tRNA Synthetases Have two roles 1) Aminoacyl-tRNA synthetases do the critical job linking the right amino acid with "cognate" tRNA • They act as a “scaffold” to match up the tRNA with its correct (“cognate”) amino acid • They catalyze a two-step reaction • This generates an ester linkage between -the 3’OH of the tRNA (on the acceptor stem) -and the COO- group of the amino acid 2) This reaction activates the amino acid for protein synthesis The Aminoacyl-tRNA Synthetase Reaction • The goal of this reaction is to activate an amino acid by forming an ester linkage with the correct tRNA O Adenine O-P-OCH2 O - O O OH H CC 3’-most ntd from the CCA acceptor stem OH C - C - R group NH3+ Amino acid The Aminoacyl-tRNA Synthetase Reaction is two steps 1) Activate the amino acid first, by reacting with ATP Amino acid + ATP Aminoacyl AMP + PPi 2Pi An enzyme-bound intermediate 2) Transfer the activated amino acid to its cognate tRNA Aminoacyl AMP +tRNA Aminoacyl-tRNA + AMP The Aminoacyl-tRNA Synthetase Reaction • The goal of this reaction is to activate an amino acid by forming an ester linkage with the correct tRNA The Aminoacyl-tRNA Synthetase Reaction - Step 1 All aminoacyl tRNA synthetase enzymes have this step in common Step 2 - differs depending on the enzyme Transfer of the amino acid to the 2’OH of the tRNA first, then the 3’OH Transfer of the amino acid to the 3’OH of the tRNA Aminoacyl-tRNA Synthetases • Despite their common function, the synthetases are a very diverse collection of enzymes • Four different quaternary structures: , 2, 4 and 22 • The subunits vary in size from 334 to more than 1000 amino acids • Two different reaction mechanisms (as seen on previous slide) There are at least 20 different AminoacyltRNA Synthetases How do we name them? tRNAAla + Ala Ala-tRNAAla Enzyme = alanyl tRNA synthetase • Are grouped into Class I or Class II based on: 1) monomers or oligomers 2) type of reaction mechanism 3) general features of their amino acid substrate 4) short stretches of amino acid similarity The two Classes of Aminoacyl-tRNA Synthetases • Larger, more hydrophobic amino acid substrates • Class I enzymes are monomers Class I Class II Arg Cys Gln Glu Ile Leu Met Trp Tyr Val Ala Asn Asp Gly His Lys Phe Pro Ser Thr • Smaller, more hydrophilic amino acid substrates • Class II enzymes are multimers Different Structures of Aminoacyl-tRNA Synthetases •The one shown in the next slide,which charges aspartic acid onto the proper tRNA, is a dimer of two identical subunits (colored blue and green, the two tRNA molecules are colored red). •Others are small monomers or large monomers, or dimers, or even tetramers of one or more different types of subunits. •Some have wildly exotic shapes, such as the serine enzyme. Aspartyl-tRNA Synthetase (dimer of identical subunits in green and blue - tRNAs in red) Representative Class I (left) & Class II (right) structures Different classes bind to different “sides” of the tRNA tRNA Class I enzyme Class II enzyme The Sidedness of tRNA Binding Defines the Reaction Mechanism tRNA anticodon loop Class I enzymes cradle the tRNA, gripping the anticodon loop, and placing the amino acid acceptor end of the tRNA in the active site (at the top right in each tRNA). These all approach the tRNA similarly and add the amino acid to the last 2’OH group in the tRNA. The Sidedness of tRNA Binding Defines the Reaction Mechanism The Class II enzymes such as the phenlyalaninyl tRNA synthetase approach the tRNA from the other side, and add the amino acid to the 3’ hydroxyl on the last tRNA base. http://www.rcsb.org/pdb/molecules/pdb16_2.html Aminoacyl-tRNA Synthetases • All have a common 2-domain structure A catalytic domain A variable domain Interacts with the tRNA 3’OH Interacts with the specific bases on the tRNA that identify that tRNA Recognizes and binds the cognate amino acid Aminoacyl-tRNA Synthetases • Must exhibit high specificity (fidelity) • This is a two different levels: 1) They must be able to recognize and bind to the correct tRNA 2) They must be able to recognize and bind to the correct amino acid (have an editing function for this one) Recognition of tRNAs Some tRNA synthetases recognize their cognate tRNA by binding to the anticodon loop. These enzymes are not gentle with tRNA molecules. For example glutaminyl-tRNA synthetase firmly grips the anticodon, spreading the three bases widely apart for better recognition. Recognition of tRNAs • Recognition of tRNA molecules using the anticodon is not always possible. • For example 6 different codons specify Ser, so seryltRNA synthetase must recognize six tRNA molecules with six different anticodons (isoacceptor tRNAs). • Therefore, tRNA molecules are also recognized using bases elsewhere in the molecule. • Base number 73 in the sequence, seems to play a major role in many cases, but in other cases it is completely ignored. tRNA structure and recognition elements Recognition of tRNAs No common set of rules for tRNA recognition !!! • Anticodon region is not the only recognition site • The "inside of the L" and other regions of the tRNA molecule are also important • Specificity of several aminoacyl-tRNA synthetases: 1) one or more bases in anticodon, 2) one or more bases in the acceptor stem, 3) discriminator base 73 Identity elements in tRNAs tRNAMet tRNAVal Identity elements reside in the anticodon Alterating the anticodon of tRNAVal to the Met anticodon results in recognition of the mutant tRNA by methyonyl-tRNA synthetase Identity elements in tRNAs tRNAPhe The 5 bases that are identity elements reside in the anticodon (3), G20 in the D loop and A73 near the 3’end G20 may be especially important in recognition of tRNAPhe since it is not found in any other tRNA Identity elements in tRNAs tRNASer Six codons for Ser, which are quite different from one another. Six “isoacceptor” tRNAs It makes sense that the anti-codon loop is not used to recognize tRNASer Identity elements in tRNAs tRNAAla Single G3:U70 pair defines specificity G:C, A:U or U:G do not work Including this base pair in other tRNAs allows them to be recognized by the alanyl tRNA synthetase A completely synthetic “microhelix” can be aminoacylated provided that G3:U70 is present tRNAAla Single G3:U70 pair defines specificity G:C, A:U or U:G do not work Including this base pair in other tRNAs allows them to be recognized by the alanyl tRNA synthetase High fidelity in amino acid selection •Aminoacyl-tRNA synthetases must perform their tasks with high accuracy, since every mistake will result in a misplaced amino acid when new proteins are constructed. •These enzymes make about one mistake in 10,000. For most amino acids, this level of accuracy is not too difficult to achieve. •Most of the amino acids are quite different from one another. •But in a few cases, it is difficult to choose just the right amino acids and these enzymes must resort to special techniques. http://www.rcsb.org/pdb/molecules/pdb16_3.html High fidelity in amino acid selection •Isoleucine is a particularly difficult example. •It is recognized by an Ile-shaped hole in the enzyme, too small to fit larger amino acids like Met and Phe, and too hydrophobic to bind anything with polar side chains. •But, the slightly smaller amino acid Val, different by only a single methyl group, also fits nicely into this pocket, binding instead of Ile in about 1 in 150 times. •This is far too many errors, so corrective steps must be taken. http://www.rcsb.org/pdb/molecules/pdb16_3.html High fidelity •Isoleucyl-tRNA synthetase solves this with an editing site. •Ile does not fit into this site, but Val does. •The mistaken Val is then cleaved away, leaving the tRNA ready for a properly-placed Leu amino acid. •This proofreading step improves the overall error rate to about 1 in 3,000. http://www.rcsb.org/pdb/molecules/pdb16_3.html 3’ 5’ CGG Codon-Anticodon Interaction Anticodon loop 3 2 1 1 2 3 5’ GCC Codon in mRNA 3’ • Predict that every codon should have a corresponding tRNA (and anticodon) • BUT IT IS NOT THAT SIMPLE • There are 61 different codons specifying amino acids • BUT there are far fewer than 61 different tRNAs • we can conclude that some tRNAs must bind to > 1 codon Codon-Anticodon Interaction For Example: Yeast tRNAAla GCU GCC GCA • Recognition of the 3rd codon base is not as “precise” as the first 2 codon bases • The genetic code is consistent with this (3rd base degeneracy) This led Crick to propose a “Wobble Hypothesis” -there are canonical base pairs for the 1st and 2nd bases of the codon, and non-canonical base pairs for the 3rd base Codon-anticodon interactions Anticodon (base #1) Codon (base #3) C A G U I G U C,U A,G U,C,A • with these rules a minimum of 31 different tRNAs is required to recognize all 61 codons that encode amino acids Third-Base Degeneracy and the Wobble Hypothesis • Codon-anticodon pairing is the crucial feature of the "reading of the code" • But what accounts for "degeneracy": are there 61 different anticodons, or can you get by with fewer than 61, due to lack of specificity at the third position? • Crick's Wobble Hypothesis argues for the second possibility - the first base of the anticodon (which matches the 3rd base of the codon) is referred to as the "wobble position" The Wobble Hypothesis How? • Canonical base pairing with the 1st two codon bases • Loose, weak base pair interactions with the 3rd codon base • #1 nucleotide of the anticodon is in a flexible domain of the tRNA Why? • Kinetic advantage • tRNA can dissociate more readily from the RNA template • Allows faster protein synthesis Codon Usage • More than one codon exist for most amino acids (except Met and Trp) • Organism may have a preferred codon for a particular amino acid • Preferred codons depend on A:T/G:C content • Codon usage correlates with abundance of tRNAs (preferred codons are represented by abundant tRNAs) • Rare tRNAs correspond to rarely used codons • mRNAs containing rare codons experience slow translation Nonsense suppression • Mutations that produce in-frame TAA, TAG and TGA result in premature termination of protein synthesis • Second mutations may appear that suppress the effect of nonsense mutations: these suppressors are tRNAs!!! • In these tRNA mutations, anticodons are altered so that “stop” signals could insert amino acids • Suppressor tRNAs originate from minor isoacceptor tRNAs so that they do not interfere with translation of highly utilized codons.