* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Dr Una Fairbrother
Survey
Document related concepts
Western blot wikipedia , lookup
Structural alignment wikipedia , lookup
List of types of proteins wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Homology modeling wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Protein mass spectrometry wikipedia , lookup
Protein folding wikipedia , lookup
Protein domain wikipedia , lookup
Circular dichroism wikipedia , lookup
Intrinsically disordered proteins wikipedia , lookup
Transcript
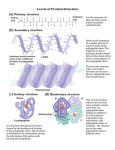
Proteins Dr Una Fairbrother Dipeptides Two amino acids are combined as in the diagram, to form a dipeptide. Water is the other product Peptides Peptides are normally written with the terminal amino group (N-terminal) to the left and the carboxyl group (C-terminal) to the right. Polypeptides Continued formation of peptide bonds extends the molecule to many amino acids linked by peptide bonds. Polymers of amino acids called POLYPEPTIDES Individual units of the polypeptide are called amino acid RESIDUES Can estimate the no. of amino acid residues in a polypeptide or protein by its molecular weight (Mr). Assume the mean Mr of an amino acid residue is 110 dalton Protein Structure or Hierarchy Protein structure is considered at different levels. Primary Secondary Tertiary Quaternary Primary structure Describes the unique sequence of amino acids which make up the polypeptide(s). i.e a bead necklace where each different coloured bead represents an amino acid. The beads can be arranged in any order or have any frequency Secondary structure is content of regular or repeating structures i.e. a helix and b pleated sheets. For the a helix consider the bead necklace twisted into a coil. The nature and structure of the a helix was elucidated by Linus Pauling and Robert Corey using Xray diffraction analysis and some simple chemical rules. polypeptide chain follows a coiled path X ray diffraction a-keratin - wool b- keratin - silk a b a helix Thermodyamically favoured structure - the preferred and thus most stable structure of a polypeptide in the absence of interactions Stabilised by H-bonding between the carbonyl oxygen and amino hydrogen of the peptide linkages. Each linkage is H-bonded to 2 other linkages one three units ahead and one three units behind. The H-bonds are approximately parallel to the long axis of the helix. Alpha helix 10Å = 1nm Each turn has 3.6 amino acid residues Each turn extends 5.4Å along the long axis Hydrogen bonds are between every fourth amino acid residue lie parallel to the long axis occur between carbonyl oxygens and amino hydrogens within different peptide linkages Helices and other structures If a protein contains long stretches of a helix it will be semi-rigid and fibrous. E.g a keratin found in hair and horn Silk or b-Keratin,excreted by the caterpillar of the silk moth. A polypeptide of glycine, alanine, and smaller amounts of other amino acids called fibroin b-Keratin molecules do not form a helix they lie on top of each other to give ridged sheets of linked amino acids, with glycine appearing on only one side of the sheets. The sheets then stack one on top of the other. This planar structure is felt when you touch the smooth surface of silk. b pleated sheets Polypeptide extended, not coiled polypeptide regions may come to lie alongside each other. These regions stabilised by H-bonds between the polypeptide regions. Here, H-bonds are roughly at right-angles to the long axis of the polypeptide chain in contrast to the a helix. b pleated sheet types Globular proteins Contain only short regions of a helix No systematic structures. single chains, two or more chains which interact in the usual ways portions of the chains with: helical structures, pleated structures, or completely random structures. Relatively spherical in shape Common globular proteins include egg albumin, hemoglobin, myoglobin, insulin, serum globulins in blood, and many enzymes. Globular proteins and Proline (a) (a)Regions can be lined up (b) as parallel (N C to N C) or antiparallel (N C to C N) Proline forces the chain to kink and does not allow the a helix to continue it is a helix breaker residue. often found in globular proteins at the end of regular sequences where the polypeptide chain bends back on itself. (b) proline in green and glycine in yellow. the side chain of proline forms a ring attached to the amino N atom (in blue). The N atom has no hydrogen so can't act as an H bond donor. This "breaks the chain" of H-bonds in helix Tertiary Structure Describes the superfolding of the polypeptide. The resultant structure contains regular regions of secondary structure It is stabilised by a range of different interactions or bonds. Bonds in tertiary structure Hydrogen bonding is between side chains of the amino acid residues (compare with H-bonding in secondary structure which is between peptide linkages) Ionic bonds between oppositely charged side chains (eg positively charged lysine residues and negatively charged glutamic acid residues). Hydrophobic interactions between the hydrocarbon side chains in phenylalanine, leucine, isoleucine and valine. Disulphide bridges between cysteine residues, these are covalent and more difficult to break. Hexokinase An example of a protein showing helices, -structure and connecting loops Hexokinase phosphorylates glucose Bonds and denaturing agents Bonds o r l inkage s Dena tur ing agen ts Ion ic bond ( sal t br idge) -COOH +H 3 N Di sulph ide br idge -S-S - Aci d (- COOH - t o ŠCOO H) Alk ali ( + H 3 N t o H 2 N-) Redu cing age nt s eg mer cap toe thano l (- S-S- to ŠSH H S-) Organ ic s ol ven ts (non po lar) and de tergen ts Urea ( N 2 N-C=O | N H 2) Hea t, ion si ng rad iati on Hy drophob ic b onds Hy drogen bond s ~C=O ..... ...... H-N~ Fo rma tion of unna tural Š S-Sbr idge s Str uc tural d ist or ti on Sal ts of me tals su ch a s c opper , lead , mer cur y and cadm ium Quaternary structure (a) the association of individual polypeptide subunits into a multisubunit or multimeric protein. Polypeptides with surface regions of hydrophobic amino acids will tend to associate in order to bring those patches together and reduce interactions with water. (b) Hexokinase, domain 1 and 2 What stabilises quaternary structure? Hydrophobic bonding Ionic bonding Hydrogen bonding unlike tertiary structure there is no covalent bonding such as would be obtained with -S-Sbridges Summary