* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download No Slide Title
Marcus theory wikipedia , lookup
Physical organic chemistry wikipedia , lookup
George S. Hammond wikipedia , lookup
Enantioselective synthesis wikipedia , lookup
Elias James Corey wikipedia , lookup
Asymmetric induction wikipedia , lookup
Ring-closing metathesis wikipedia , lookup
Hydroformylation wikipedia , lookup
Vinylcyclopropane rearrangement wikipedia , lookup
Tiffeneau–Demjanov rearrangement wikipedia , lookup
Hofmann–Löffler reaction wikipedia , lookup
Discodermolide wikipedia , lookup
Bottromycin wikipedia , lookup
Petasis reaction wikipedia , lookup
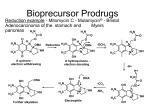
Synthesis and Retrosynthesis of Peptidomimetic Inhibitors Thrombin Presented by: Kevin Condel Overview / Terminology • Goal: To design and synthesize a peptidomimetic that competes to inhibit the enzyme thrombin. • Thrombin is part of a cascade leading to the formation of insoluble fibrin, a material found in blood clots. Unregulated clotting may lead to cardiac arrest or stroke. • A Peptidomimetic is any small organic molecule that mimics the transition state of a natural substrate. • Peptidomimetics competitively inhibit the enzyme process, preventing the natural reaction from occurring. – i.e. - The peptidomimetic binds more readily to thrombin than the substrate. • Hydroxy-aldehydes are important components of peptidomimetic inhibitors of the thrombin system. This work involves the development of a simple, yet effective protocol for the generation of hydroxy-aldehydes. • Blood clotting must be regulated. Thrombin • Errors in blood clotting lead to: – cardiac arrest (in the heart) – stroke (in the brain) • Thrombin begins inactive and is shown on the bottom-left. • Inactive thrombin has extra domains, colored blue, which are clipped off during activation. • The purple atoms are Ca2+ ions, bound to modified glutamate amino acids. – The strong (+) charge on these ions tether the protein to the surfaces of blood vessels, holding thrombin and localizing it to one spot. • Since inactive thrombin is held, blood clots will generally not spread to other areas. – Only the thrombin adjacent to the damage will be activated. • Activated thrombin (the upper structure shown here) lasts only seconds, serving also to limit the clot to the area of damage. • Thrombin is simply part of a cascade which serves to synthesize the cross-linked fibrin polymers found in blood clots. Thrombin Click to enlarge Thrombin as an Enzyme • Thrombin has an active site consisting of the catalytic triad: Ser 195, His 57, and Asp 102 Enzyme Binding Site • In addition to the active site, thrombin has three binding sites, labeled as S1, S2 and S3, that determine the strength and specificity of binding • The lipophilicity of S3 has been well determined • Lipophilicity represents the affinity of a molecule or moiety for a lipophilic environment (i.e. hydrophobicity) Inhibition of the Active Site • Leeches synthesize proteins that block thrombin (and other enzymes), stopping the formation of the clot. • One example, a protein called hirudin, is shown here on the left in blue. Notice how it fits the active site of thrombin perfectly. Peptidomimetics • Small peptide-like molecules that mimic transition state of substrate and work by competitively inhibiting the binding of the natural substrate (ex. Hirudin from the leech) • Peptide analog must be stable. • Drug must be a reversible inhibitor of the enzyme but can be irreversible if the enzyme is unique to the disease. Saquinavir Project Design • Goals: – Design a polypeptide isotere based on a natural thrombin substrate (natural Phe-Pro-Arg tripeptide shown below) – Optimize a generalized scheme for isotere synthesis – Model the S2 and S3 steric and hydrophobic requirements Project Setup • All reactions required an anhydrous environment. • Nitrogen steadily flushed through the system to exclude water vapor. • Various temperatures were achieved as follows: – -78C (Dry Ice + Acetone) – 0C (Ice Bath) Overall Reaction S S Br 1. BuLi 2. TMSCl SiMe 3 N N OR OR OR CHO unmasking N 2-TST R CHO R S OH CHO R OH S SiMe 3 2-TST N 1. 4.22ml n-Butyl-Lithium was injected into a round bottom flask containing a swirling solution of 50 ml ether through the septum using the syringe 2. 2.33ml 2-Bromothiazole in 50 ml of ether slowly added over 30 min via a separatory funnel into a controlled -78ºC nitrogenous environment. 3. Mixture allowed to stir for 30 min 4. 4.33ml (CH3)SiCl in 50 ml ether added drop wise for 30 min via the sep funnel Bu S Br Bu- + Li S + N N Br Li S Me 3Si SiMe 3 Cl + N Li Cl 2-TST NMR Identification S SiMe 3 N Benzaldehyde 1. 7.33g Benzoic Acid was added under inert nitrogenous conditions to a flask containing 15 mL Tetrahydrofuran (THF). 2. Under 0ºC conditions, 45 mL of pre-cooled 1.0 M Lithium Aluminum Hydride (LiAlH4) in THF was added dropwise with vigorous stirring. C OH LiAlH4 0ºC O RT PCC C H O RT, 6-12 h CH2 OH (No isolation) CH2 OH Benzaldehyde 3. After the hydrogen had evolved, the solution was cooled to room temp. and stirred for 30 min. 4. In a separate flask under the same nitrogenous conditions, 14.3 g Pyridinium Chlorochromate (PCC) was added to 100mL Methylene Chloride and stirred into solution. C OH LiAlH4 0ºC O RT PCC C H O RT, 6-12 h CH2 OH (No isolation) CH2 OH Benzaldehyde 5. The alkoxyaluminum salt in THF created by mixing LiAlH4 with benzoic acid was next added dropwise at room temperature via a separatory funnel 6. The reaction mixture was stirred for 12 hours at room temperature, diluted with diethyl ether, and filtered and washed to remove the supernatant liquid. The ether was then distilled from the filtrate to obtain benzaldehyde. C OH LiAlH4 0ºC O RT PCC C H O RT, 6-12 h CH2 OH (No isolation) CH2 OH Benzaldehyde NMR Identification C H O Amino Acid Reduction Methodology (Theoretical) Click to enlarge • Use benzaldehyde synthesis schematic to reduce the amino acid argenine • When reduced argenine is combined with 2-TST it creates the Arg side chain of the Phe-Pro-Arg tripeptide peptidomimetic • Unmasking protocol removes thiazole ring and replaces it with CHO group, creating the active site inhibitor. • Phe-Pro addition will be conducted in future experimentation. Reduction of Argenine (Novel Approach) 1. 12.64g Argenine was added under inert nitrogenous conditions to a flask containing 20 mL Methyl Sulfoxide as a solvent. 2. Under 0ºC conditions, 45 mL of pre-cooled 1.0 M Lithium Aluminum Hydride (LiAlH4) in THF was added dropwise with vigorous stirring. + + H3N C O C + NH H2N H3N OH LiAlH4 0ºC RT + CH2 NH H2N NH2 OH PCC C + H3N NH2 (No isolation of alkoxy-salt) 0ºC 2 h RT 6-12 h C + C H NH H2N NH2 O Reduction of Argenine (Novel Approach) 3. After the hydrogen had evolved (2 h), the solution was cooled to room temp. and stirred for 30 min. The intermediate was not isolated. 4. In a separate flask under the same nitrogenous conditions, 14.3 g Pyridinium Chlorochromate (PCC) was added to 100mL Methylene Chloride and stirred into solution. + + H3N C O C + NH H2N H3N OH LiAlH4 0ºC RT + CH2 NH H2N NH2 OH PCC C + H3N NH2 (No isolation of alkoxy-salt) 0ºC 2 h RT 6-12 h C + C H NH H2N NH2 O Reduction of Argenine (Novel Approach) 5. The intermediate in Methyl Sulfoxide created by mixing LiAlH4 with argenine was next added dropwise at room temperature via a separatory funnel. 6. Mixture stirred for 12 hours at room temperature, diluted with diethyl ether, filtered and washed. The ether was then distilled from the filtrate to obtain the aldehyde of argenine. + + H3N C O C + NH H2N H3N OH LiAlH4 0ºC RT + CH2 NH H2N NH2 OH PCC C + H3N NH2 (No isolation of alkoxy-salt) 0ºC 2 h RT 6-12 h C + C H NH H2N NH2 O Methodological Problems with Argenine Reduction • Need a nonpolar, nonreactive solvent to dissolve argenine without interfering with the reaction. • Methyl Sulfoxide NOT efficient as a solvent for this reaction due to its exothermicity. – i.e (Broken Manifold and intense sulfur scent) • Length of complete reaction and temperature requirements are dependent on the solvent used to dissolve argenine. – With Methyl Sulfoxide, it is proposed that 0ºC conditions must exist for at least two hours prior to addition of PCC. Retrosynthesis and Unmasking Protocol Mechanism (Theory) + H3N C + + C H3N H O N RT, 4 h NH H2N Me3Si + S C OH S NH H2N NH2 Aldehyde of Argenine C N NH2 Intermediate 2-TST RT, 4 h Retrosynthesis and Unmasking Protocol Mechanism (Theory) Me + H3N C + + N H3N C OH S NH H2N CF3SO3Me4 NaBH4 C + N C OH S NH H2N NH2 NH2 Previous research has determined that CF3SO3Me4 is added to N-methylate the thiazole ring. NaBH4 is added to reduce the mixture and break the pi bonds. Retrosynthesis and Unmasking Protocol Mechanism (Theory) Me + H3N C + N H3N C OH NH O + S Cu++ / Water C + C C H OH NH H2N H2N NH2 NH2 Cu++ and water are added to hydrolyze the system, removing the thiazole ring and adding yet another aldehyde. Retrosynthesis and Unmasking Protocol Mechanism (Theory) + H3N C + C CH2 O + NR2 H3N C C H OH NaBH3CN NH HNR2 C + H2N NH2 OH NH H2N NH2 HNR2 is added as part of a dehydration reaction to remove water and add NR2 to the molecule. This NR2 represents the part of the molecule that will be interacting with the active site of thrombin. References • Alessandro Dondoni, et al.; Synthesis of TSTs and Reactions with Carbonyl Compounds; J. Org. Chem. 1988, 53, 1748-1761 • Benoit Bachand , et al.; Synthesis and Structure-Reactivity of Potent Bicyclic Lactam Thrombin Inhibitors; Bioinorg. & Med. Chem. 1999, 9, 913-918 • Jin Soon Cha, et al.; Preparation of Aldehydes from Carboxylic Acids by Reductive Oxidation with Lithium Aluminum Hydride and Pyridinium Chlorochromate or Pyridinium Dichromate; Bull. Korean Chem Soc. 1999, Vol. 20, No. 4 Acknowledgements • We gratefully acknowledge the support of the Welch Foundation in the form of a Departmental Research Grant Questions? Thrombin Theory Structure Peptidomimetics Inhibition Project Goals Method Argenine Reduction Catalytic Triad Apparatus Problems Binding Sites Overall Reaction Retrosynthesis 2-TST Reaction Benzaldehyde Reaction NMR Identification NMR Identification Proposed Peptidomimetic Click to view natural substrate Natural Substrate Click to view proposed peptidomimetic PCC Pyridinium Chlorochromate NH+ CrO3Cl-