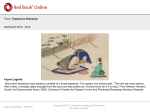
* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Introduction
Point mutation wikipedia , lookup
Genetic code wikipedia , lookup
Gene expression wikipedia , lookup
Expression vector wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Magnesium transporter wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Structural alignment wikipedia , lookup
Protein purification wikipedia , lookup
Interactome wikipedia , lookup
Western blot wikipedia , lookup
Homology modeling wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Biochemistry wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Introduction Andrew Torda, wintersemester 2006 / 2007, 00.904 Angewandte … • who am I ? • language .. English .. verhandelbar • Zettel • www.bioinformatics.uni-hamburg.de/research/BM/torda/lehre.html • + stine • Übungen also on web • where should information go ? stine or our pages ? 23/05/2017 [ 1 ] Administration People • Andrew Torda 42838 7331 1. Stock / 105 [email protected] (but use phone or stairs) • sekr (Annette Schade) 7330 • • • • Gundolf Schenk Paul Reuter (maths as well) Nasir Mahmood Stefan Bienert (more in DNA) Vorlesungen Mittwoch 12:30 Übungen (flexibel) jede zweite Woche Freitag 12:15 23/05/2017 [ 2 ] Homework / Übungen Not too much • enough from other courses Übungen • very short report (schriftlich) Textbooks • any biochemistry book (Stryer, Biochemistry as per chem dept) • expensive, not used too much • statistics, Ewens, W.J. and Grant, G.R., Statistical Methods in Bioinformatics, Springer, 2001 • Leach, Andrew, “Molecular Modelling” very good for future semesters • Folien should be sufficient 23/05/2017 [ 3 ] Exams • any facts that are mentioned in these lectures and Übungen • schriftliche Klausur 16 Feb 23/05/2017 [ 4 ] Protein Structure - the problem - sociological • Easy ? boring ? • Essential • How many people have done biology ? chemistry ? • Mein Vorschläge • Freitag 12:30 protein structure lecture or • Schnell hier • The chemists can correct me + • Freitag 12:30 optional protein structure with details • For everybody in normal lecture slot 23/05/2017 [ 5 ] Broad themes Theme of Semester • given some information about a macromolecule (protein) • what can be calculated ? predicted ? • how much would you trust predictions ? • limitation, applicability, reliability • typical information • a protein sequence (lots known) • a protein structure (less known) • a DNA sequence (think of genomes) 23/05/2017 [ 6 ] Specific and general models Dream • Feed data to box and have it interpreted • given my protein, what is the structure ? • given my spectrum where is the centre of the peak ? Model types • Specific • you know the structure of your data, fit points to the observations • General • look for some patterns in data – little understanding of the underlying theory • examples 23/05/2017 [ 7 ] Interpreting spectroscopic data • just an example (no spectroscopy in this course) • many kinds of peaks in spectroscopy look like 0.09 0.07 theoretical peak 0.05 amplitude 0.03 0.01 -0.01 0 20 40 60 80 frequency (Hz) • my mission • find centre (≈24) and height (≈0.08) • but they have noise 23/05/2017 [ 8 ] noisy data 0.12 • real world has noise • we still want centre, height 0.1 real peak with noise 0.08 amplitude 0.06 0.04 0.02 • try simple smoothing • no assumptions about data 0 0 50 100 frequency (Hz) 0.12 0.1 • claim • centre around 23 • looks believable smoothed data 0.08 amplitude 0.06 0.04 0.02 0 0 50 100 frequency (Hz) 23/05/2017 [ 9 ] Using prior knowledge • I expect peaks like a2 (a 2 x 2 ) 0.15 • A fit of a calculated peak… • something is clearly wrong amplitude 0.1 • if peak has a certain width it 0.05 must have an appropriate height 0 a2 (a 2 x 2 ) best fit of a theoretical peak 0 50 100 frequency (Hz) • What looked good is not the correct form 0.15 amplitude 0.1 0.05 0 0 50 100 frequency (Hz) 23/05/2017 [ 10 ] More appropriate fitting • what if we used two peaks ? shape of two peaks added 0.15 0.1 amplitude 0.05 • peaks centred at 20 and 26 • very different explanation of data 0 0 50 100 frequency (Hz) 0.15 amplitude two peaks explain data almost perfectly 0.1 0.05 0 0 50 100 frequency (Hz) 23/05/2017 [ 11 ] General vs appropriate modelling 0.12 • general smoothing method suggested one peak • looks good • appears to explain observations • generally applicable 0.1 smoothed data 0.08 amplitude 0.06 0.04 0.02 0 0 50 100 frequency (Hz) 0.15 amplitude best fit of a theoretical peak 0.1 0.05 0 0 • testing with correct model suggested this is a trick 50 100 frequency (Hz) 0.15 • fitting with best model (two peaks) • near perfect • summary • if you know the underlying model, use it • always applicable ? • back to biological questions amplitude two peaks explain data almost perfectly 0.1 0.05 0 0 50 100 frequency (Hz) 23/05/2017 [ 12 ] General purpose modelling • Proteins have "secondary structure • It appears to reflect the sequence of amino acids • what is the rule ? • 20 amino acids, N positions, • 20N sequences, patterns not clear • what to do ? • correct model – think of all atomic interactions • see where atoms should be placed • not practical • or • forget physics • use dumb statistics / machine learning approaches http://www.cgl.ucsf.edu/chimera/ImageGallery/, bluetongue virus capsid protein 23/05/2017 [ 13 ] Mixtures of specific and general Will a ligand (Wirkstoff) bind to a protein ? • with physics • model all atomic interactions, best physical model • calculate free energy (∆G) • difference in solution / bound • more generally • gather idea of important terms (H-bonds, overlap, ..) • try to find some function which often works • do not stick to real physics Will my drug dissolve in water or oil (lipid) ? (important) • sounds like chemistry • usually approached by machine learning • number of atoms, types of atoms, … http://www.cgl.ucsf.edu/chimera/ImageGallery/, bluetongue virus capsid protein 23/05/2017 [ 14 ] Similarity 6 4 • Important in all bioinformatics 2 • I have a protein of unknown • structure / function / cell localisation 0 0 2 4 set 1 (units) • is it similar to one of known structure, function … • Similarity seems obvious • two sets of numbers (above) • two protein sequences ACDEACDE rather similar - but quantified ? ADDEAQDE • how many positions differ ? how long are proteins ? • could the similarity be by chance ? synteny plot: http://home.cc.umanitoba.ca/~umlawda/39.769/presentation/presentation.html, Dr. Brian Fristensky set 2 (units) 6 23/05/2017 [ 15 ] Similarity Two genomes similarity • what are the descriptors ? • how many genes are common ? • is the order preserved ? Potential drugs • drug 1 binds, will drug 2 ? • how similar ? synteny plot: http://home.cc.umanitoba.ca/~umlawda/39.769/presentation/presentation.html, Fristensky, B. ligands from, Wang, N., DeLisle, R. K. and Diller, D.J. (2005), J. Med. Chem., 48, 6980-6990 23/05/2017 [ 16 ] Detection and Quantification • Models for prediction and interpretation • often not well justified • Similarity in these applications • detection (finding / recognising) • quantification • Each in the context of applications • first protein structure … 23/05/2017 [ 17 ] Summary so far A model can explain observations, make predictions or both A model may be based • on a belief of the underlying chemistry / physics • purely mathematical, probabilistic Similarity • we have objects with some information (proteins, ligands, genomes, sequences, …) • we want to find similar objects and hope they have the same properties • similarity has a different meaning in different areas 23/05/2017 [ 18 ] Proteins - who cares ? • Most important molecules in life ? Ask the DNA / RNA people • • • • • • • • structural (keratin / hair) enzymes (catalysts) messengers (hormones) regulation (bind to other proteins, DNA, ..) industrial – biosensors to washing powder receptors transporters (O2, sugars, fats) anti-freeze … 23/05/2017 [ 19 ] Proteins are easy • data (protein data bank, www.rcsb.org) • nearly 40 000 structures • literature on function, interactions, structure • software • viewers, molecular dynamics simulators, docking, .. • nomenclature and rules Proteins are not friendly • one cannot take a sequence and predict structure /function • data formats are full of surprises, mostly old formats • data contains error and mistakes 23/05/2017 [ 20 ] Protein Rules • Physics /chemistry versus rules / dogma / beliefs / folklore • Physics / Chemistry • protein + water = set of interacting atoms • can be calculated (not really) • Rules (not quantified) • proteins unfold if you heat them (exceptions ?) • if they contain lots of charged amino acids, they are soluble • if they are more than 300 residues, they have more than one domain, • proteins fold to a unique structure (could you prove this ?) • lowest free energy structure 23/05/2017 [ 21 ] Protein chemistry • Chemists / biochemists may sleep (quietly) • Short version • proteins are sets of building blocks (amino acids, residues, Reste) • 20 types of residue • chains of length few to 103 ( 100 or 200 typical) • small ones (< ≈50) are peptides • Longer version 23/05/2017 [ 22 ] Sizes • 1 Å = 10-10m or 0.1 nm structure size bond CH CC protein radius 1Å 1.5 Å 102 - 103 Å α-helix spacing 5½Å Cαi to Cαi+1 3.8 Å myoglobin picture 1gjn from www.rcsb.org 23/05/2017 [ 23 ] proteins are polymers • simple polymers A X B many times gives A X X X X X X X B example what kind of polymer would this give ? Is it obvious what R is ? 23/05/2017 [ 24 ] Why are proteins interesting polymers ? boring polymer gives uninteresting structures OK for plastic bags, haushaltsfolie. Not nice regular structures.. What can we do to make things more protein like ? 23/05/2017 [ 25 ] Giving proteins character 1 • more complicated backbone with H-bond • donor R R • acceptor R • basis of standard regular structures in proteins (secondary structure) R R • repeating polymer unit: R • if this was all there was • all proteins would be the same 23/05/2017 [ 26 ] protein chemistry amino acids (monomers) all look like: H NH2 C C OH O R maybe H NH3 + C C O } O sidechain R α carbon or Cα • how can we construct specific structures ? • different kinds of "R" groups 23/05/2017 [ 27 ] Putting monomers together NH2 H C R1 OH C O NH2 + H C R2 NH2 H C R1 O H C N H C R2 OH C O + NH2 H C R3 O H C N H C C R3 C OH O OH O • protein synthesis story (biochemistry lectures) ? • peptides and proteins • < 30 or 40 residues = peptide • > 30 or 40 residues = protein 23/05/2017 [ 28 ] side chain possibilities • • • • big / small charged +, charged -, polar hydrophobic (not water soluble), polar interactions between sites… A C C R W S T G B 23/05/2017 [ 29 ] Backbone and consequences • peptide bond is planar • partial double bond character • shorter than other C-N • nearly always trans NH2 H C O C R1 N H H C R2 O C N H H C R3 OH C O • two bonds can rotate NH2 H C O C R1 N H phi φ H C R2 O C N H H C R3 C OH O psi ψ 23/05/2017 [ 30 ] ramachandran plot • can we rotate freely ? • no… steric hindrance 180 β 120 • Ramachandran plot 60 ψ psi -180 0 α -120 -60 diagrams from http://www.cgl.ucsf.edu/home/glasfeld/tutorial/AAA/AAA.html -60 0 60 120 180 -120 -180 φ phi 23/05/2017 [ 31 ] Backbone H bonds • oxygen is slightly negative • NH bond is polar δ+ C H N δ- O δ+ δ- H N C O • H-bonds • can be near or far in sequence • fairly stable at room temperature 23/05/2017 [ 32 ] Secondary structure • regular structures using information so far • rotate phi, psi angles so as to • form H-bonds where possible • do not force side chains to hit each other (steric clash) • two common structures • α-helix • β-strand / sheet 23/05/2017 [ 33 ] α helix • • • • each CO H-bonded to NH 3 or 4 away 3.6 residues per turn 2 H-bonds per residue side chains well separated 23/05/2017 [ 34 ] β-sheet • β-strand • stretch out backbone and make NH and CO groups point out • β-sheet • join these strands together with H-bonds (2 H-bonds/residue) • anti-parallel • or parallel 23/05/2017 [ 35 ] After α-helix and β-sheet • do helices and sheets explain everything ? • no • there is flexibility in the angles (look at plot) • geometry is not perfectly defined • there are local deviations and exceptions 180 • other common structures 120 • tighter helices 60 • some turns ψ psi 0 • other structure -180 -120 -60 0 -60 • coil, random, not named 60 120 180 -120 -180 φ phi 23/05/2017 [ 36 ] What determines secondary structure ? So far • secondary structure pattern of H-bonding Almost all residues have H-bond acceptor and donor • all could form α-helix or β-sheet ? No Difference ? • sequence of side-chains – overall folding Why else are sidechains important • chemistry of proteins (interactions, catalysis) Fundamental dogma • the sequence of sidechains determines the protein shape • why is dogma a good word ? 23/05/2017 [ 37 ] Side chain properties • properties • big / small • neutral / polar / charged • special (…) • example • phenylalanine side chain looks like benzene (benzin) • very insoluble • benzene would rather interact with benzene than water • what if you have phe-phe-phe… poly-phe ? • does not happen in nature (can be made) • would be insoluble • not like a real peptide • phe is a constituent of real proteins – has a role 23/05/2017 [ 38 ] Properties are not clear cut • You can be big / small, hydrophic / polar • combinations are possible • Do not memorise this figure Taylor, W.R. (1986) J. Theor. Biol., 215-218 23/05/2017 [ 39 ] Sidechain interactions • • • • ionic (if the sidechains have charge) hydrophobic (insoluble sidechains) H-bonds (some donors and acceptors) repulsive 23/05/2017 [ 40 ] summary of amino acids N O O N O O N N O N Alanyl N Arginyl [Ala] O N N Asparaginyl Aspartyl [Asn] [Asp] [Arg] O O O O O O O N O S N N Cysteinyl [Cys] O N N Glutamyl [Glu] Glutaminyl [Gln] O Glycyl [Gly] O N O N N N N Isoleucyl [Ile] Histidyl [His] O O S N N Leucyl [Leu] O Lysyl [Lys] O O N Methionyl [Met] N N Phenylalanyl [Phe] N Prolyl [Pro] Seryl [Ser] O O O O N Threonyl [Thr] N N Tryptophanyl [Trp] O N O Tyrosyl [Tyr] N Valyl [Val] Diagram from MDL isis draw. Better pictures at http://www.cryst.bbk.ac.uk/pps97/course/section2/aa_diags.html 23/05/2017 [ 41 ] Amino Acids by property aromatic O tryptophan N N O phenylalanine N O tyrosine O N 23/05/2017 [ 42 ] rather hydrophobic O leucine O isoleucine N N O cysteine methionine S O S N N O alanine O proline N N O glycine O valine N N 23/05/2017 [ 43 ] Polar O O threonine N serine O O N glutamine O O N N asparagine O O N N 23/05/2017 [ 44 ] charged N O histidine N N O N N N lysine arginine N N O aspartate O O O N N glutamate O O O N 23/05/2017 [ 45 ] Hydrophobicity – how serious ? • very serious, but simplified • the lists above are • pH dependent • difficult to measure experimentally (some aspects) • is hydrophobicity really defined ? Other properties - size big … trp gly small ala 23/05/2017 [ 46 ] Other properties – chemistry / geometry O • proline • only one rotatable angle ! • peptide bond sometimes cis N 180 • pro ramachandran plot • 4000 points 120 60 ψ psi -180 -120 0 -60 0 -60 60 120 180 -120 -180 φ phi 23/05/2017 [ 47 ] gly and cys • glycine • no side chain • can visit forbidden parts of phi-psi map (4000 points here) 180 120 60 ψ psi -180 -120 0 -60 0 -60 60 120 180 -120 -180 φ phi • cysteine • forms covalent links with other cys picture from Stryer, L, Biochemistry, WH Freeman, 1981 23/05/2017 [ 48 ] Summary so far • proteins are heteropolymers • from the backbone alone form α-helices and β-strands (and more) • side-chains determine the • pattern of secondary structure • overall protein shape • special amino acids • cys (forms disulfide bridges) • gly (can visit "forbidden" regions of ramachandran plot • pro (no H-bond donor) • last bits of nomenclature… 23/05/2017 [ 49 ] Nomenclature • some rules are unavoidable Alanine Cysteine Aspartic acid Glutamic acid Phenylalanine Glycine Histidine Isoleucine Lysine Leucine Methionine Asparagine Proline Glutamine Arginine Serine Threonine Valine Tryptophan Tyrosine Ala Cys Asp Glu Phe Gly His Ile Lys Leu Met Asn Pro Gln Arg Ser Thr Val Trp Tyr A C D E F G H I K L M N P Q R S T V W Y • always write from N to C terminal • important convention 23/05/2017 [ 50 ] Definitions, primary, secondary … • first, some more definitions • primary structure • sequence of amino acids • ACDF (ala cys asp phe…) • secondary structure • α-helix, β-sheet (+ few more) • structure defined by local backbone • tertiary structure • how these units fold together • coordinates of a protein • quaternary structure • how proteins interact 23/05/2017 [ 51 ] Protein structure general comments • primary, secondary, tertiary structure … how real ? • primary/secondary well defined • edges can blur • supersecondary struct / tertiary 23/05/2017 [ 52 ] Representation • Ultimately, our representation of a structure… ATOM ATOM ATOM ATOM ATOM ATOM ATOM ATOM ATOM ATOM ATOM ATOM ATOM ATOM ATOM ATOM ATOM ATOM ATOM 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 N CA C O CB CG CD NE CZ NH1 NH2 N CA C O CB CG CD N ARG ARG ARG ARG ARG ARG ARG ARG ARG ARG ARG PRO PRO PRO PRO PRO PRO PRO ASP 1 1 1 1 1 1 1 1 1 1 1 2 2 2 2 2 2 2 3 31.758 31.718 33.154 33.996 30.886 29.594 28.700 27.267 26.661 27.370 25.367 33.800 34.976 34.960 33.962 34.922 34.058 33.371 36.192 13.358 13.292 13.224 12.441 12.103 11.968 13.182 12.895 13.087 13.558 12.797 13.936 13.367 11.922 11.306 14.145 15.391 15.273 11.317 -13.673 -12.188 -11.664 -12.225 -11.724 -12.534 -12.299 -12.546 -13.727 -14.735 -13.838 -10.586 -9.840 -9.660 -9.391 -8.523 -8.737 -10.096 -9.707 x, y, z coordinates 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 18.79 14.26 18.25 20.10 16.74 15.96 15.45 12.82 17.38 18.38 25.73 17.07 14.99 13.11 10.57 15.81 18.91 19.41 8.73 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 • drawing the structure ? 23/05/2017 [ 53 ] Drawings 3 ways of looking at “ras” protein bare Cα trace ribbons ribbons and cylinders are these just cosmetic differences ? diagrams made with molscript http://www.avatar.se/molscript/ 23/05/2017 [ 54 ] Different levels of abstraction pictures from "Structural Bioinformatics", ed Bourne, PE and Weissig, H., Wiley New York (2003) 23/05/2017 [ 55 ] Atomistic For details • where does a ligand bind ? • which interactions is a residue involved in ? Ribbons Overview • shape • number secondary struct elements • symmetry pictures from "Structural Bioinformatics", ed Bourne, PE and Weissig, H., Wiley New York (2003) strands helices 23/05/2017 [ 56 ] More abstract • no idea of real shape • very quickly classify a protein – example • lots of serine proteases • lots of different sequences • all very similar at this level of abstraction pictures from "Structural Bioinformatics", ed Bourne, PE and Weissig, H., Wiley New York (2003) 23/05/2017 [ 57 ] Why does structure matter ? • • • • what residues can I change and preserve function ? what is the reaction mechanism of an enzyme ? what small molecules would bind and block the enzyme ? is this protein the same shape as some other of known function ? Where do structures come from ? • • • • topic of other course (lots of detail) X-ray crystallography NMR + a bit of small angle X-ray scattering, electron diffraction, … 23/05/2017 [ 58 ] Atomic coordinates - warnings • remember the coordinate file ? • lots of problems • atoms and residues missing • numbering can be peculiar • history • suits fortran 66 (think columns) • non-standard amino acids • nucleotides, ligands • accuracy ATOM ATOM ATOM ATOM ATOM ATOM ATOM ATOM ATOM ATOM ATOM ATOM ATOM ATOM ATOM ATOM ATOM ATOM ATOM 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 N CA C O CB CG CD NE CZ NH1 NH2 N CA C O CB CG CD N ARG ARG ARG ARG ARG ARG ARG ARG ARG ARG ARG PRO PRO PRO PRO PRO PRO PRO ASP 1 1 1 1 1 1 1 1 1 1 1 2 2 2 2 2 2 2 3 31.758 31.718 33.154 33.996 30.886 29.594 28.700 27.267 26.661 27.370 25.367 33.800 34.976 34.960 33.962 34.922 34.058 33.371 36.192 13.358 13.292 13.224 12.441 12.103 11.968 13.182 12.895 13.087 13.558 12.797 13.936 13.367 11.922 11.306 14.145 15.391 15.273 11.317 -13.673 -12.188 -11.664 -12.225 -11.724 -12.534 -12.299 -12.546 -13.727 -14.735 -13.838 -10.586 -9.840 -9.660 -9.391 -8.523 -8.737 -10.096 -9.707 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 1.00 18.79 14.26 18.25 20.10 16.74 15.96 15.45 12.82 17.38 18.38 25.73 17.07 14.99 13.11 10.57 15.81 18.91 19.41 8.73 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 1BPI 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 23/05/2017 [ 59 ] resolution, precision, accuracy • coordinates 1.1 1.0 8.5 • what do they mean ? • random errors • non-systematic / noise / uncertainty • should be scattered around correct point • from any measurement there are errors ±x.y • x-ray crystallography has model for data • uncertainty (probability) • resolution (experimental) • < 1 Å (good) • > 5 Å (bad, but excusable – monster structures) 23/05/2017 [ 60 ] X-ray crystallography • non-systematic errors • small problems: (O and N look the same) • few huge problems • newer structures are better • proteins are not static • overall motion • local motion N C Cα Cβ N N O Cα C Cβ O N 23/05/2017 [ 61 ] NMR structures • different philosophy to X-ray • lots of little internal distances • do not quite define structure • generate 50 or 102 solutions • look at scatter of solutions • as with X-ray • some parts are well defined • some not structure 1sm7 from www.rcsb.org 23/05/2017 [ 62 ]