* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Document
Bimolecular fluorescence complementation wikipedia , lookup
List of types of proteins wikipedia , lookup
Protein design wikipedia , lookup
Protein purification wikipedia , lookup
Circular dichroism wikipedia , lookup
Protein folding wikipedia , lookup
Protein domain wikipedia , lookup
Western blot wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Intrinsically disordered proteins wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Protein mass spectrometry wikipedia , lookup
Alpha helix wikipedia , lookup
Structural alignment wikipedia , lookup
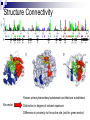
Protein Sectors: Evolutionary Units of Three-Dimensional Structure Najeeb Halabi, Olivier Rivoire, Stanislas Leibler, and Rama Ranganthan Cell 138, 774-786, August 21, 2009 Journal Club Yizhou Yin Sep 23, 2009 Sequence Conservation “…sequence conservation – the degree to which the frequency of amino acids at a given position deviates from random expectation in a well sampled multiple sequence alignment of the protein family...” evolution sequence structure Evolutionary relationship sequence conservation Structural/functional importance property/function Hypothesis -However, in the 3-dimensional structure of protein, the large amount of interactions between amino acid residues are also fundamental “structural elements”. -Amino acid distributions at individual position should not be taken as independent of one another. -Investigation of correlations between sequence positions in protein family leads to decomposition of the protein into groups of coevolving amino acids – “sectors”. Hypothesis: the sectors are features of proteins structures and reflect the evolutionary histories of their conserved biological properties. S1A Family Serine protease Clan Family Sub-family SA SB … S1 S2 … S1A … Catalytic triad – active site rat trypsin (3TGI) Member trypsin chymotrypsin tryptase kallikrein granzyme … Broad distribution and functions Prokaryotes Digestion Invertebrates Blood clotting Vertebrates Inflammation … Binding site - specificity Method Outline Identification of sectors Statistical Coupling Analysis Statistical Independence Correlated entropy Physical connectivity Distinct biochemical properties Alanine mutagenesis Catalytic power & thermal stability assays Independent divergence Sequence similarity analysis From Sequence to Sectors Multiple sequence alignment of 1470 members of the S1A family (single domain) NCBI nonredundant database through iterative PSI-BLAST Alignment: Cn3D, ClustalX Standard manual adjustment methods Position Conservation Di(a): Divergence (or relative entropy) fi(a): Observed frequency of amino acid a at position i q(a): Background frequency of a in all proteins Statistical Coupling Analysis (SCA) SCA matrix (conservation-weighted covariance matrix) Cijab: frequency-based correlation between position i and j ~Cijab is a measure of the significance of observed correlations as judged by the conservation of the amino acids under consideration After binary approximation: Binary approximation Di(ai): the conservation of ai, which is the most prevalent amino acid at that position Spectral cleaning to separate functional correlation from statistical and historical noise Principal Component Analysis Spectral decomposition of ~Cij matrix to partially sort out the different contributions to the correlations 223 eigenvalues Lowest 218 – Statistical noise Randomized alignments retaining the same size and amino acid propensities at sites show eigenvalues of similar magnitude First mode makes the dominant contribution to ~Cij – historical noise The first eiganvelue is well approximated by a first order approximation, proves that the first eigenvector should just report the net contribution of each position to the total correlation Sector Identification using modes 2 to 5 Overview of Sectors Statistical Independence Compute correlation entropy to quantitatively measure the independence of sectors Minimum discriminatory information method i.e. S is small set of position, specifically, the top five positions contributing to each sector Structure Connectivity Known primary/secondary/subdomain-architecture subdivision No sector Distinction in degree of solvent exposure Difference in proximity to the active site (not for green sector) Red: focus on S1 pocket catalytic specificity Blue: more distributed property Green: focus around catalytic triad catalytic activity Without information about tertiary structure and only ~10% of total sequence positions contributes strongly to each sector, each sector reveals obvious intrasector physical connectivity and only a few inter-sector contacts. Biochemical Independence Mutations of red and blue sectors showed very different effects focused either on catalytic power or thermal stability Additive effects from combination of mutations between two groups (magenta: observed | white: predicted) Independent Sequence Divergence Sequence similarity analysis of each sector classifies members in the family effectively only by the related property, while the analysis on all positions failed to do the classification (442 members with functional annotation) Evidence of “Sector” theory in Other Protein Families PDZ PAS SH2 SH3 Different regulatory mechnisms Discussion Novel Structural Organization Implication for Physical Properties of Proteins Alternative View to Calculate Residue Covariance Technical Challenges Protein Modulization Adaptive Advantage