* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Homology Modeling via Protein Threading - lmm
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Genetic code wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Multi-state modeling of biomolecules wikipedia , lookup
Gene expression wikipedia , lookup
Magnesium transporter wikipedia , lookup
Expression vector wikipedia , lookup
Biochemistry wikipedia , lookup
Point mutation wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Bimolecular fluorescence complementation wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Metalloprotein wikipedia , lookup
Western blot wikipedia , lookup
Protein purification wikipedia , lookup
Interactome wikipedia , lookup
Proteolysis wikipedia , lookup
Homology Modeling
via
Protein Threading
Kristen Huber
ECE 697S
Topics in Computational Biology
April 19, 2006
Fundamentals of Protein Threading
Protein Modeling
Homology Modeling
Protein Threading
Generalized Overview of a Threading Score
Score Methodology based on Multiple
Protein Structure Alignment
Protein Modeling
20,000 entries of proteins in the PDB
1000 - 2000 distinct protein folds in nature
Thought to be only several thousand
unique folds in all
Protein Structure Prediction
aim
of determining the three-dimensional
structure of proteins from their amino acid
sequences
Types of Structure Prediction
De novo protein
methods
seek to build three-dimensional
protein models "from scratch"
Example: Rosetta
Comparative protein
modeling
uses previously solved structures as
starting points, or templates.
Example: protein threading
Factors that Make Protein Structure
Prediction a Difficult Task
The number of possible structures that proteins
may possess is extremely large, as highlighted
by the Levinthal paradox
The physical basis of protein structural stability
is not fully understood.
The primary sequence may not fully specify the
tertiary structure.
chaperones
Direct simulation of protein folding is not
generally tractable for both practical and
theoretical reasons.
Homology Modeling
Homolog a protein related to it by
divergent evolution from a
common ancestor
40 % amino-acid identity with its
homolog
NO large insertions or deletions
Produces a predicted structure
equivalent to that of a medium
resolution experimentally solved
structure
25 % of known protein sequences
fall in a safe area implying they
can be modeled reliably
Homology Modeling Defined
Homology modeling
Based
on the reasonable assumption that two
homologous proteins will share very similar
structures.
Given the amino acid sequence of an unknown
structure and the solved structure of a homologous
protein, each amino acid in the solved structure is
mutated computationally, into the corresponding
amino acid from the unknown structure.
Homology Modeling Limitations
Cannot study conformational changes
Cannot find new catalytic/binding sites
Brainstorm lack of activity vs activity
Chymotrypsionogen,
trypsinogen and plasminogen
40%
homologous
2 active, 1 no activity, cannot explain why
Large Bias towards structure of template
Models cannot be docked together
Why Homology Modeling?
Value in structure based drug design
Find common catalytic sites/molecular
recognition sites
Use as a guide to planning and interpreting
experiments
70-80 % chance a protein has a similar fold to
the target protein due to X-ray crystallography or
NMR spectroscopy
Sometimes it’s the only option or best guess
Protein Threading
A target sequence is threaded through the
backbone structure of a collection of template
proteins (fold library)
Quantitative measure of how well the sequence
fits the fold
Based on assumptions
3-D
structures of proteins have characteristics that
are semi-quantitatively predictable
reflect the physical-chemical properties of amino
acids
Limited types of interactions allowed within folding
Fold Recognition Methods
Bowie, Lüthy and Eisenberg (1991)
2 approaches to recognition methods
Derive a 1-D profile for each structure in the fold
library and align the target sequence to these
profiles
Identify
amino acids based on core or external
positions
Part of secondary structure
Consider the full 3-D structure of the protein
template
Modeled as a set of inter-atomic distances
NP-Hard (if include interactions of multiple residues)
Protein Threading
The word threading implies that one drags the
sequence (ACDEFG...) step by step through
each location on each template
Protein Threading
Generalized Threading Score
Want to correctly recognize arrangements of residues
Building a score function
G(rAB) = kTln (ρAB/ ρAB°)
potentials of mean force
from an optimization calculation.
G, free energy
k and T Boltzmanns constant and temperature respectively
ρ is the observed frequency of AB pairs at distance r.
ρ° the frequency of AB pairs at distance r you would expect to
see by chance.
Z-score = (ENat - <Ealt>)/σ Ealt
Natural energies and mean energies of all the wrong structures/
standard deviation
Scoring Different Folds
Goodness of fit score
Based
on empirical energy
function
Modify to take into account
pairwise interactions and
solvation terms
High score means good fit
Low score means nothing
learned
Some Threading Programs
3D-pssm (ICNET). Based on sequence profiles, solvatation potentials and
secondary structure.
TOPITS (PredictProtein server) (EMBL). Based on coincidence of
secondary structure and accesibility.
UCLA-DOE Structure Prediction Server (UCLA). Executes various threading
programs and report a consensus.
123D+ Combines substitution matrix, secondary structure prediction, and
contact capacity potentials.
SAM/HMM (UCSC). Basen on Markov models of alignments of crystalized
proteins.
FAS (Burnham Institute). Based on profile-profile matching algorithms of the
query sequence with sequences from clustered PDB database.
PSIPRED-GenThreader (Brunel)
THREADER2 (Warwick). Based on solvatation potentials and contacts
obtained from crystalized proteins.
ProFIT CAME (Salzburg)
Process of 3D Structure
Prediction by Threading
Has this protein sequence similarity to other with a
known structure?
Structure related information in the databases
Results from threading programs
Predicted folding comparison
Threading on the structure and mapping of the
known data
A comparison between the threading predicted
structure and the actual one
Protein Threading Based on Multiple Protein
Structure Alignment
Tatsuya Akutsu and Kim Lan Sim
Human Genome Center, Institute of Medical Science,
University of Tokyo
NP-Hard if include interactions between 2 or
more AA
Determine multiple structural alignments based
on pair wise structure alignments
Center
Star Method
Center Star Method
Let I0 be the maximum number of gap symbols placed before the
first residue of S0 in any of the alignments A(S0; S1); : : : ;A(S0; SN).
Let IS0j be the maximum number of gaps placed after the last
character of S0 in any of the alignments, and let Ii be the maximum
number of gaps placed between character S0;i and S0;i+1, where Sj:i
denotes the i-th letter of string Si
Create a string S0 by inserting I0 gaps before S0, IjSo gaps after S0,
and Ij gaps between S0;I and S0;i+1.
For each Sj (j > 0), create a pairwise alignment A(S0; Sj) between S0
and Sj by inserting gaps into Sj so that deletion of the columns
consisting of gaps from A(S0; Sj) results in the same alignment as
A(S0; Sj).
Simply arrange A(S0; Sj )'s into a single matrix A (note that all A(S0;
Sj )'s have the same length).
Simple Threading Algorithm
Apply simple score function based on structure alignment algorithm
Let X = x1……xN (input amino acid sequence)
Ci ( i-th column in A)
Test and analyze results and/or apply constraints
Protein Threading with Constraints
Assume part of the input sequence xi…xi+k must
correspond to part of the structure alignment
cj…cj+k
Apply constraints
Prediction Power
Entered in CASP3 competition
17 predictions made
3 targets evaluated as similar to correct folds
Only team to create a nearly correct model for
structure T0043
Best in competition
8
evaluated as similar to correct
Next time….
In depth detail of
Multiple
structural alignment program
Multiprospector
Global
Optimum Protein Threading with
Gapped Alignment
Quality measures for protein threading
models
Improvements on threading-based models
Gapped Alignment
Review
Homology Modeling
Based
on the reasonable assumption that two
homologous proteins will share very similar
structures.
Threading
Modeled
as a set of inter-atomic distances
NP-Hard (if include interactions of multiple residues)
Build a score function based on energies in order to
correctly recognize arrangements of residues
Threading via multiple structural alignment
Score
function based upon alignment matrix
Specifics of Protein Threading
Different Threading Types
Multiprospector:
Predictions of Protein-Protein
Interaction by Multimeric Threading
Global Optimum Protein Threading with
Gapped Alignment
Quality measures for protein threading
models
Improvements on threading-based models
MULTIPROSPECTOR
An algorithm for the prediction of protein-protein
interactions by multimeric threading
Protein–protein
interactions are fundamental to
cellular function and are associated with processes
such as enzymatic activity, immunological
recognition, DNA repair and replication, and cell
signaling.
Function can be inferred from the nature of the
protein with its interactants
Use properties related to the topology of the
interface, solvent-accessible surface area and
hydrophobicity
Addressed limitations of existing approaches
Method Basis
Thread the sequences through a representative structure template
library that, in addition to monomers, also includes each of the
chains in representative protein dimer structures.
Compute the interaction energy between a pair of protein chains for
those protein structures involved in dimeric complexes.
Stable complex formation determined by the magnitude of the
interfacial potentials and the Z-scores of the complex structures
relative to that of the monomers.
Interfacial Statistical Potentials
Interfacial pair potentials
Calculated by examining each interface of the selected
dimers
P(i, j), (i=1, …, 20; j =1, …,20),
Nobs(i, j) is the observed number of interacting pairs of i, j
between two chains.
Nexp(i, j) is the expected number of interacting pairs of i, j {Nexp (i,
j) = Xi * Xj * Ntotal}
Apply Boltzman Principal to the ratio to obtain potential
of mean force between 2 residues
Multimeric Threading Strategy
and Z-Score
Z-score of the score for each probetemplate alignment is used to decide
if a correct fold is found:
is the standard deviation of
energies; Ei is the energy of the ith sequence of M alternative folds
(i 1, …, M).
Multimeric Threading
Results
Global Optimum Protein Threading
with Gapped Alignment and
Empirical Pair Score Functions
The structural model corresponds to an annotated
backbone trace of the secondary structure segments in
the conserved core fold.
Loops are not considered part of the conserved fold, and
are modeled by an arbitrary sequence-specific loop
score function.
Alignment gaps are confined to the connecting non-core
loop regions
Each distinct threading is assigned a score by an
assumed score function
Exponentially large search space of possible threadings
NP-hard search spaces as large as 9.6x1031 at rates
ranging as high as 6.8 x1028 equivalent threadings per
second
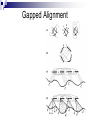
Gapped Protein
Threading Methodology
Common core of four secondary
structure segments
Spatial interactions. Small circles
represent amino acid residue positions
(core elements), and thin lines connect
neighbors in the folded core.
Thread through model by placing
successive sequence amino acid
residues into adjacent core elements. Tax
indexes the sequence residue placed
into the first element of segment X.
Sequence regions between core
segments become connecting turns or
loops.
Sets used in the branch-and-bound
search are defined by lower and upper
limits (dark arrows, labeled bax and dax
for segment X)
General Pairwise Score Function
For any threading t, let fv(v, t)
be the score assigned to core
element or vertex v
fe({u, v}, t) the score assigned
to interaction or edge {u, v}
f1(λi , t) the score assigned to
loop region λi
Then the total score of the
threading is:
Rewrite function of threading
pairs of core segments
Branch-and-Bound Search Algorithm
branch-and-bound search requires the ability to
represent the entire search space as a set of possibilities
split any set into subsets
compute a lower bound on the best score achievable within any
subset
After some finite number of steps, the chosen set will
contain only one threading (equals its lower bound)
Splitting the Search Space
The set of all legal threadings is
represented by the hyper-rectangle
lower bound on the score f(t)
attainable by any threading t in the
set T
summing lower bounds on each
term separately
The enclosing mintЄT ensures that the lower bound will be instantiated on a specific legal
threading tlbЄT. This will be used in splitting T, below. The equation further ensures that the
singleton term, in g1(i, ti ), remains consistent both with the terms that reflect loop scores, in
g2(i - 1, i, ti-1, ti ), and with the other (non-loop) pairwise terms, in g2(i, j, ti , uj ). The inner
minuЄT allows a different vector u for each i, but requires u to be a legal threading.
Search
Space
Results
Threading Results
Quality Measures for Protein
Threading Models
Evaluation of different prediction methods for protein
threading
Purpose:
determine if one method to build a model is better than another
optimize the performance of existing methods.
Threading Assessment:
ability to predict the correct fold
the similarity of the model to the correct structure
Methods of Comparison Defined
Global
Alignment Dependent
based on an exact match between the residues in the model and
the correct structure
Alignment Independent
consider all residues in both the model and the correct structure
in an "alignment dependent“ fashion
based on a structural superposition between the model and the
correct structure
Template Based
available for models that are created from the sequence being
aligned onto a single structural template.
Methods of Comparison
Comparison Results
Most methods correlate to
each other
0.51 model-normalized
0.41 template-normalized
High quality homology-models
correlate less with the rest of
the data
Measures of same type
correlate well and tend to
cluster
A Need for Improvement
Resulting models obtained from threading approaches
are usually of very low quality, with gaps and insertions
in threading alignments that somehow have to be
connected or closed
Various threading methods and their associated scoring
functions only focus on aspects of protein structure and
a subset of their possible interactions.
Method of Improvement
Employs a lattice model
SICHO (Side Chain Only)
The model has been refined by incorporating
evolutionary information into the interaction scheme.
a Monte Carlo annealing procedure attempts to find a
conformation that maintains some (but not all) features of the
original template
optimizes packing and intra-protein interactions
Lattice Model
The model chain consists of a string
of virtual bonds connecting the
interaction centers that correspond to
the center of mass of the side chains
and the backbone alpha carbons.
These interaction centers are
projected onto an underlying cubic
lattice with a lattice spacing of 1.45 A°
A cluster of excluded volume points is
associated with each bead of the
model chain.
Each cluster consists of 19 lattice points
Closest approach distance from
another cluster labels smallest interresidue distance
Interaction Scheme
Starting Model takes on a tube form
Energy potentials.
generic, sequence-independent, biases that penalize against non
protein-like conformations
two-body and multibody potentials extracted from a statistical
analysis of known protein structures.
Evolutionary information extracted from multiple sequence
alignments.
The stiffness/secondary structure bias term has the
following form:
Estiff= - Єgen [Σ min{0.5, max (0, wi ● wi+2)}]
- Єgen [Σ min{0.5, max (0, wi ● wi+4)}]
Interaction Scheme
A weak bias being introduced towards helix-type and
beta-type expanded states
Estruct= Σ{δH1(i) + δ H2(i) + δ E1(i) + δ E2(i)}
Generic packing interactions
Short range interactions
Pairwise Interactions
Multi-body Interactions
δ H1 and δ H2 contributions defined as a broad range of helical/turn
conformations
δ E1 and δ E2 as expanded conformations
statistical potential for residue type A having np parallel and na
anti-parallel contacts.
Emulti= ΣEm(A,np,na)
Total energy
Etotal = Estiff + Emap + 0.875EH-bond + 0.75Eshort + 1.25Epair
+ 0.5Esurface + 0.5Emulti
Threading Model Refinement
a) Generate the threading alignment
between the unknown sequence and the
template structure.
b) Derive the sequence similarity-based
short and long range pairwise potentials.
c) Build the starting continuous model chain
onto the lattice-projected template
structure.
d) Build the tube around the aligned
fragments of the template structure. Then,
perform the first stage of Monte Carlo
refinement.
e) Refinement of the structure
multiple alignments with homologous
sequences of unknown structures were used
in the potential derivation procedures.)
assume to be the new template
Narrow restraints
Select lowest energy structures
All atom models using MODELLER.24
RESULTS
12 targets/template
proteins of low sequence
similarity
3 models used for tuning
6 of 9 yield lower rmsd
than original
Effective parameters
Neglecting part of
threading alignment