* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download 3.1.molecular_evolution - T
Genome (book) wikipedia , lookup
Viral phylodynamics wikipedia , lookup
Non-coding DNA wikipedia , lookup
Human genome wikipedia , lookup
Dual inheritance theory wikipedia , lookup
Genetic drift wikipedia , lookup
Group selection wikipedia , lookup
History of genetic engineering wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Genetic code wikipedia , lookup
Human genetic variation wikipedia , lookup
Frameshift mutation wikipedia , lookup
Adaptive evolution in the human genome wikipedia , lookup
Genome evolution wikipedia , lookup
Koinophilia wikipedia , lookup
Point mutation wikipedia , lookup
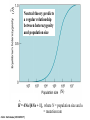
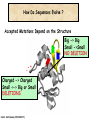
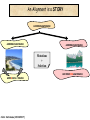
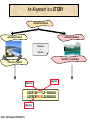
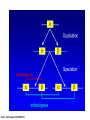
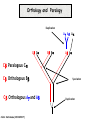
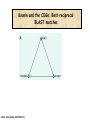
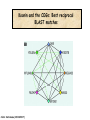
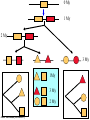
Molecular Evolution Cédric Notredame Cédric Notredame (29/04/2017) Molecular Evolution is a rapidly developing field……….. Cédric Notredame (29/04/2017) Phenotypic diversity in populations suggests underlying genetic diversity Cédric Notredame (29/04/2017) Other molecular level studies reveal even more variation Nucleotide sequence variation Cédric Notredame (29/04/2017) Is Modern Genetic Data Compatible with Darwinism Cédric Notredame (29/04/2017) If each variable locus is influenced by selection, then how can populations be so genetically variable? Can each variable “locus” influence population fitness? Genotype: Fitness: A1A1 1 A1A2 1 A2A2 1-s What about the total “cost of selection”when ALL these variable loci are taken into account? Cédric Notredame (29/04/2017) J.B.S. Haldane Neutral Theory of Molecular Evolution Genotype: Fitness: A1A1 1 A1A2 1 A2A2 1 M. Kimura Cédric Notredame (29/04/2017) How Do Sequences Evolve Each Portion of a Genome has its own Agenda. Cédric Notredame (29/04/2017) The Genetic Code is Redundant: Third-position codon changes are often “silent” Cédric Notredame (29/04/2017) Neutral Theory postulates that much of the genetic variation present in populations is not influenced by selection--rather it is a reflection of two interacting processes: (1) Mutation (2) Drift Cédric Notredame (29/04/2017) ^ (H) Neutral theory predicts a regular relationship between heterozygosity and population size (N) ^ H = 4Nu/[4Nu + 1], where N = population size and u = mutation rate Cédric Notredame (29/04/2017) The Molecular Clock Cédric Notredame (29/04/2017) THE MOLECULAR CLOCK HYPOTHESIS Another prediction of Neutral Theory--Because mutation is regular (or “clocklike”) and because selection does not influence the rate of divergence, divergence of DNA and protein molecules in two separate lineages should occur in a REGULAR, clocklike manner i.e., the longer the time separating two species from their common ancestor, the more divergent will be a given protein or DNA molecule (and this relationship should be LINEAR) Cédric Notredame (29/04/2017) Amino acid substitutions /site Cédric Notredame (29/04/2017) Amino Acid Substituions/Site x 10 If most mutations are selectively neutral, we might expect to find that molecular divergence is “clocklike”. Do we, in fact, see this? 3.2 3 2.8 2.5 2.2 2 1.8 1.5 1.2 1 150 250 350 450 550 Independent evolution (MYA) 650 Different molecular clocks for different proteins--another prediction Cédric Notredame (29/04/2017) How Do Sequences Evolve ? CONSTRAINED Genome Positions Evolve SLOWLY EVERY Protein Family Has its Own Level Of Constraint Family KS KA Histone3 Insulin Interleukin I a-Globin Apolipoprot. AI Interferon G 6.4 4.0 4.6 5.1 4.5 8.6 0 0.1 1.4 0.6 1.6 2.8 Rates in Substitutions/site/Billion Years as measured on Mouse Vs Human (80 Million years) Ks Synonymous Mutations, Ka Non-Neutral. Cédric Notredame (29/04/2017) Cédric Notredame (29/04/2017) How Can We Compare Sequences ? Using Knowledge Could Work C Aliphatic L V I A G T Aromatic F Y W H Small P G CC D K E R S N Q Hydrophobic Polar But we do not know enough about Evolution and Structure. Using Data works better. Cédric Notredame (29/04/2017) How Do Sequences Evolve ? In a structure, each Amino Acid plays a Special Role + On the surface, CHARGE MATTERS OmpR, Cter Domain Cédric Notredame (29/04/2017) In the core, SIZE MATTERS How Do Sequences Evolve ? Accepted Mutations Depend on the Structure Big -> Big Small ->Small NO DELETION + - Charged -> Charged Small <-> Big or Small DELETIONS Cédric Notredame (29/04/2017) Alignments and Evolution Cédric Notredame (29/04/2017) An Alignment is a STORY ADKPKRPLSAYMLWLN ADKPKRPLSAYMLWLN ADKPKRPLSAYMLWLN Mutations + Selection ADKPKRPKPRLSAYMLWLN ADKPRRPLS-YMLWLN Cédric Notredame (29/04/2017) An Alignment is a STORY ADKPKRPLSAYMLWLN ADKPKRPLSAYMLWLN ADKPKRPLSAYMLWLN Mutations + Selection ADKPKRPKPRLSAYMLWLN ADKPRRPLS-YMLWLN Insertion Deletion ADKPRRP---LS-YMLWLN ADKPKRPKPRLSAYMLWLN Mutation Cédric Notredame (29/04/2017) Comparing Is Reconstructing Evolution Cédric Notredame (29/04/2017) Divergeant or Convergeant Evolution ? Cédric Notredame (29/04/2017) Evolution is NOT Always Divergent… Chen et al, 97, PNAS, 94, 3811-16 AFGP with (ThrAlaAla)n Similar To Trypsynogen N S AFGP with (ThrAlaAla)n NOT Similar to Trypsinogen Cédric Notredame (29/04/2017) Evolution is NOT Always Divergent AFGP with (ThrAlaAla)n Similar To Trypsynogen N S AFGP with (ThrAlaAla)n NOT Similar to Trypsinogen SIMILAR Sequences BUT DIFFERENT origin Cédric Notredame (29/04/2017) Orthologous And Paralogous Genes Cédric Notredame (29/04/2017) Cédric Notredame (29/04/2017) Orthology and Paralogy Duplication Ag Ad Aa Cb Ca Bb Ba Ab Aa Cb Paralogous Ca Cb Orthologous Bb Cb Orthologous Ag and Ad Cédric Notredame (29/04/2017) Speciation Duplication Koonin and the COGs: Best reciprocal BLAST matches Cédric Notredame (29/04/2017) Koonin and the COGs: Best reciprocal BLAST matches Cédric Notredame (29/04/2017) Cédric Notredame (29/04/2017) Cédric Notredame (29/04/2017) Cédric Notredame (29/04/2017) Mixing Orthologs and Paralogs leads to inexact Phylogenetic trees Cédric Notredame (29/04/2017) 0 My 1 My 2 My 3 My 1My 3 My 2 My Cédric Notredame (29/04/2017) Positions are not enough to infer Orthology Cédric Notredame (29/04/2017) Comparative Map of the Human and Mouse Genomes Human chromosomes Mouse chromosomes Similar colored blocks = blocks of conserved synteny (blocks of similar genes) Cédric Notredame (29/04/2017)