* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
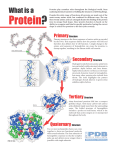
Download SSE – secondary structure element (ex. helices, sheets)
Paracrine signalling wikipedia , lookup
Point mutation wikipedia , lookup
Biochemistry wikipedia , lookup
Gene expression wikipedia , lookup
Expression vector wikipedia , lookup
Magnesium transporter wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Bimolecular fluorescence complementation wikipedia , lookup
Metalloprotein wikipedia , lookup
Interactome wikipedia , lookup
Protein purification wikipedia , lookup
Western blot wikipedia , lookup
Two-hybrid screening wikipedia , lookup
An Efficient Index-based Protein Structure Database Searching Method 陳冠宇 Introduction More than 18,000 protein structures stored in PDB (September 2002) Structural comparison(3D) and database searching – other methods practice exhaustive searching Their design philosophy: Filter-and-refine Using Indexed-based searching method Results: 16 times faster than DALI Filter-and-Refine query ProtDex Actual alignment Top 100 proteins Database 20,000 proteins result Problem Definition Protein Structures 3D Structural Comparison Structural Database Searching A protein is composed of a sequence of amino acid (AA) residues. SSE – secondary structure element (ex. helices, sheets) Loop Regions (no specific shape) Sequence Comparison vs. Structural Comparison One cannot determine the similarity of two remotely homologous proteins by sequence comparison. We try to superimpose one protein structure over another in order to obtain the minimum root mean square deviation (RMSD) between them. -> O(n4m4) The ProtDex Method Step 1: Extracting Information from PDB database Step 2: Building Intra-molecular Distance Matrices Design rationale: two protein structures are similar if their distance matrices are similar Step 3: Cutting Fixed Matrices and Extracting Properties Step 4: Building Inverted File Index Step 1: Extracting Information For each protein chain in PDB file: PDB id - chain id; No. of AA residues; No. of SSEs For each AA Residue: 3D coordinate (x, y, z) of C carbon For each SSE: SSE type (Helix or Sheet); SSE Start position; SSE length Step 2: Representation - Building Distance Matrices Protein 9xxxx with 7 AA residues Step 3-1: Contact Patterns & FixedSize Matrices SSE(H) SSE(E) contact patterns Fixed-size matrix Step 3-2: Extracting Properties For the 2X2 sub-matrix starting at the cell (2, 2), we store the values: 8, HH, (3,3), (1,1), (1,1) For the 2X2 sub-matrix starting at the cell (3,6), we store the values: 49, HE, (3,2), (1,2), (2,1), etc. Step 4: Building Inverted File Index Implemented as sorted list Searching a Protein Structure S(Q,P) = WFMCount(Q,P) X WGSum(I,j) X Sigma(match(I,j)[ (WTerm(i) X max(match(a,b)^PdbIdb=P)( WArea(a,b) X WARatio(a,b) X WOrdinal(a,b) ) ] WFMCount is to compensate the effect that the large proteins being matched and scored more frequently than the small ones. WTerm is to add more weight to the query index terms that rarely occur in the database. Discussion Design: representation of structures scoring schemes comparison algorithms assessment of the results Performance Accuracy – SCOP classification hierarchy is made of 4 levels: class, fold, superfamily and family Pros and Cons of ProtDex Conclusions Advantages: Speed (need not to scan through each structure in the database) Disadvantages: Cannot provide the actual alignment Storage overhead for the index structure (the entire index: 1.2GB) Time requirement to build and update the index (building the entire index: 30min 38 sec)