* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Classification and Phylogenetic Analysis of the cAMP
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Multilocus sequence typing wikipedia , lookup
NADH:ubiquinone oxidoreductase (H+-translocating) wikipedia , lookup
Gene expression wikipedia , lookup
Biochemistry wikipedia , lookup
Metalloprotein wikipedia , lookup
Community fingerprinting wikipedia , lookup
Magnesium transporter wikipedia , lookup
Western blot wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Genetic code wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Proteolysis wikipedia , lookup
Protein–protein interaction wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Point mutation wikipedia , lookup
Structural alignment wikipedia , lookup
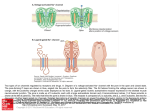
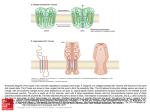
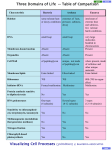
J Mol Evol (2002) 54:17–29 DOI: 10.1007/s00239-001-0013-1 © Springer-Verlag New York Inc. 2002 Classification and Phylogenetic Analysis of the cAMP-Dependent Protein Kinase Regulatory Subunit Family Jaume M. Canaves, Susan S. Taylor Department of Chemistry and Biochemistry, 0654, University of California, San Diego, 9500 Gilman Drive, La Jolla, CA 92093-0654, USA Received: 2 November 2000/Accepted: 14 June 2001 Abstract. The members of the PKA regulatory subunit family (PKA-R family) were analyzed by multiple sequence alignment and clustering based on phylogenetic tree construction. According to the phylogenetic trees generated from multiple sequence alignment of the complete sequences, the PKA-R family was divided into four subfamilies (types I to IV). Members of each subfamily were exclusively from animals (types I and II), fungi (type III), and alveolates (type IV). Application of the same methodology to the cAMP-binding domains, and subsequently to the region delimited by -strands 6 and 7 of the crystal structures of bovine RI␣ and rat RII (the phosphate-binding cassette; PBC), proved that this highly conserved region was enough to classify unequivocally the members of the PKA-R family. A single signature sequence, F–G–E–[LIV]–A–L–[LIMV]–x(3)– [PV]–R–[ANQV]–A, corresponding to the PBC was identified which is characteristic of the PKA-R family and is sufficient to distinguish it from other members of the cyclic nucleotide-binding protein superfamily. Specific determinants for the A and B domains of each Rsubunit type were also identified. Conserved residues defining the signature motif are important for interaction with cAMP or for positioning the residues that directly interact with cAMP. Conversely, residues that define subfamilies or domain types are not conserved and are mostly located on the loop that connects ␣-helix B⬘ and  strand 7. Key words: PKA — cAMP — cAPK — Regulatory Correspondence to: Susan S. Taylor; email: [email protected] subunit — Motif — Classification — Phylogeny — Signature Introduction The cyclic AMP (cAMP)-dependent protein kinase (PKA) regulatory subunit family (PKA-R family) is part of the cyclic nucleotide-binding protein (cNMP) superfamily. Among others, the cNMP superfamily includes a variety of bacterial regulators, such as the catabolite gene activator protein (CAP) (Eron et al. 1971; Weber and Steitz 1987), cyclic nucleotide-gated ion channels (Nakamura and Gold 1987; Ludwig et al. 1990), guanine nucleotide exchange factors (Kawasaki et al. 1998), and the regulators of PKA and PKG. The distinctive feature of the members of the superfamily is the presence of a common structural domain of about 120 residues (Shabb and Corbin 1992) capable of binding cyclic nucleotides. In eukaryotes, PKA mediates a wide range of cellular responses to external stimuli (Taylor et al. 1990). Of the protein kinases, PKA is also one of the simplest and best characterized (Taylor et al. 1990), primarily because the catalytic (C) and regulatory (R) components are the products of distinct genes and the proteins are separated upon activation. In the absence of cAMP, PKA exists as an equimolar tetramer of R and C subunits. In addition to its role as an inhibitor of the C subunit, the R subunit anchors the holoenzyme to specific intracellular locations (DellÁcqua and Scott 1997) and prevents the C subunit from entering the nucleus (Fantozzi et al. 1994). All R-subunit isoforms have a conserved and welldefined domain structure (Taylor et al. 1990) (Fig. 1). 18 Fig. 1. Summary of the general domain structure of a representative dimeric PKA regulatory subunit. A Schematic linear domain structure of bovine RI␣. The N-terminal dimerization domain, inhibitory region, cAMP-binding domain A, and cAMP-binding domain B are shown. The expanded view of the inhibitory sequence shows the different motifs identified for this region. B and C Detailed views of PBCs A and B in the presence of cAMP. A ribbon representation showing the structural elements of the PBCs has been overlaid to a stick representation of the sequence. Each region has its own function, and it also communicates with other regions as part of the conformational changes that are induced by the binding of cAMP. R subunits interact primarily with C subunits through the inhibitory site. PKA regulatory subunits contain two tandem cAMP-binding domains at the C terminus, designated A and B (Takio et al. 1982) (Fig. 1). These cAMPbinding domains, presumably resulting from gene duplication, show extensive sequence similarity and bind cAMP cooperatively (Døskeland and Øgreid 1984; Robinson-Steiner and Corbin 1983). PKA regulatory subunits have typically been classified according to their physicochemical properties, namely, overall charge and molecular mass, and the presence or absence of a serine residue susceptible to phosphorylation in the inhibitory site. In the present work, we propose a new classification based on analysis of the most conserved region of the R subunits: the phosphatebinding cassette (PBC) of each cAMP-binding domain. Also, we have generated a highly specific signature pattern capable of unequivocally identifying the PBCs of PKA regulatory subunits and identified type- and subtype-specific residues. This provides a means for fast assignment of new sequences to specific R subfamilies and could be used as a valuable tool for R-subunit identification and automatic sequence annotation of short protein fragments. Experimental Procedures Sequences and Database Searches. Sources of sequences analyzed in this study are shown in Table 1. Sequence retrieval was performed using BLAST. Initial searches were carried out with PSI-BLAST (po- sition-specific iterated BLAST) (Altschul et al. 1997) set to search the nonredundant database using human RI␣ as the query sequence and a BLOSUM62 weight matrix (Henikoff and Henikoff 1992). Sequences identified in this first stage were used as query sequences for subsequent searches. Sequences unequivocally identified as corresponding to cAMP-dependent kinase regulatory subunits were downloaded from SwissProt and TrEMBL Release 38. In a later stage of the project, a consensus sequence pattern common to all the detected R subunits was used with human RI␣ as template sequence to try to detect additional R subunits using PHI-BLAST (Pattern Hit Initiated BLAST) (Zheng et al. 1998). Search for unidentified R subunits was performed using BLAST on several major genomic databases. The sequences matching the signature pattern were compared to the signature subpatterns for PBC A and B, as well as the different type-specific subpatterns, and classified accordingly. DNA sequences were translated with the Translate utility at Expasy (http://www.expasy.ch). Multiple Sequence Alignment. Multiple sequence alignments were performed using ClustalW 1.6 (Thompson et al. 1994). Priority was set to placing gaps within loops connecting secondary structure elements. The cAMP-binding regions were manually curated with the program GeneDoc (Nicholas et al. 1997), taking into consideration the crystal structures of bovine RI␣ (Su et al. 1995) and rat RII (Diller et al. 2001). The Gonnet weight matrix (Gonnet et al. 1994) was used in the sequence alignments. The complete sequence alignments are available from the authors upon request. Phylogenetic Tree Construction. Phylogenetic trees based on the amino acid alignments of the complete sequences, the cAMP-binding domains, and PBCs were reconstructed using the neighbor-joining method (Saitou and Nei 1987). The statistical significance of the phylogenetic trees obtained was tested by bootstrap analysis with 100 replicates of random additions (Felsenstein 1985). The resulting trees were visualized with the program Treeview (Page 1996). Amino acid sequence distances were corrected for multiple amino acid substitutions according to Kimura (Kimura 1983). Motif Identification and Signature Sequence Generation. Amino acids that were conserved within the entire family of PKA regulatory 19 Table 1. cAMP-dependent protein kinase regulatory subunit family members analyzed in this study: (A) full-length sequences; (B) fragmentsa A Abbreviation b hRI␣ bRI␣ pRI␣ rRI␣ hRI mRI rRI hRII␣ bRII␣ mRII␣ hRII bRII rRII AplyR StroR HemiR CaenR DrosR DictR Sch1R Sch2R BlasR UstiR SaccR EmerR MagnR NeurR CollR EuplR ParaR Source species Accession No. No. AAc MWd pIe Homo sapiens Bos taurus Sus scrofa Rattus norvegicus Homo sapiens Mus musculus Rattus norvegicus Homo sapiens Bos taurus Mus musculus Homo sapiens Bos taurus Rattus norvegicus Aplysia californica Strongylocentrotus purpuratus Hemicentrotus pulcherrimus Caenorhabditis elegans Drosophila melanogaster Dictyostelium discoideum Schizosaccharomyces pombe Schizosaccharomyces pombe Blastocladiella emersonii Ustilago maydis Saccharomyces cerevisiae Emericella nidulans Magnaporthe grisea Neurospora crassa Colletotrichum trifolii Euplotes octocarinatus Paramecium tetraurelia P10644 P00514 P07082 P09456 P31321 P12849 P81377 P13861 P00515 P12367 P31323 P31322 P12369 P31319 Q26619 Q25114 P30625, A21820 P16905 P05987 P36600 O14272 P31320 P49605 P07278 O59922 O14448 Q01386 O42794 Q9XTM6 Q94725 381 379 379 380 380 380 380 403 400 400 417 417 415 377 369 368 376 377 327 411 347 403 522 415 412 390 385 404 338 325 42,981.6 42,761.5 42,790.4 42,963.7 43,026.8 43,092.9 43,150.9 45,387.2 44,962.6 45,257.9 46,214.9 46,204.8 45,991.7 42,606.2 41,788.8 41,679.8 42,649.3 42,367.2 36,835.7 46,363.3 38,917.1 44,467.6 55,957.9 47,087.9 44,903.7 42,270.5 42,156.2 44,778.0 38,786.3 37,275.5 5.27 5.27 5.27 5.27 5.64 5.71 5.60 4.96 4.80 4.78 4.82 4.85 4.90 5.36 4.61 4.66 4.99 5.07 5.65 5.06 5.85 4.82 8.88 7.82 5.09 5.05 5.45 9.17 8.11 7.61 B Abbreviationb Source species Accession no. Comments pRI RbRI RET/TyrK pRII␣ rRII␣ mRII hRII MucoR Sus scrofa Oryctolagus cuniculus Homo sapiens Sus scrofa Rattus norvegicus Mus musculus Homo sapiens Mucor rouxii Q29083 O77795 Q15300 P05207 P12368 P31324 O60380 AAF44694 (GenBank) DD domain A domain A domain A domain A and B domains DD domain B domain A and B domains a The Comments column indicates the structural modules that are contained in the fragment: dimerization-docking domain (DD domain), cAMP-binding domain A (A domain), or cAMP-binding domain B (B domain). Unless specified, all the accession numbers correspond to Swissprot entries. b subunits or only within subfamilies were identified based on the final alignment of all the regulatory subunits. Signature sequences were created manually from the sequence logos corresponding to the cAMPbinding cassettes. Amino acid patterns are abbreviated in one-letter code, where “X” indicates any of the 20 essential amino acids. The signature sequence was evaluated for specificity by carrying out motif searches with the ScanProsite tool (Appel et al. 1994) for searching the Swissprot and TrEMBL protein databases. were generated via the WebLogo server maintained by Steven E. Brenner (http://www.bio.cam.ac.uk/cgi-bin/seqlogo/logo.cgi). In sequence logos, the horizontal axis represents the position of the residue, whereas the vertical axis represents the information content of that position in bits. The height of the one-letter residue symbol at each position is proportional to the information content of the residue at that position. Entropy plots (Schneider and Stephens 1990) were created with the BioEdit sequence alignment editor (Hall 1999). Entropy plots are a measure of the lack of predictability for each alignment position. The entropy or uncertainty (H) was calculated according to the expression Sequence Logos and Entropy Plots. Sequence logos (Schneider and Stephens 1990) corresponding to R-subunit cAMP-binding domains Abbreviation used in this study. Number of amino acids in the protein. d Molecular mass (Da). e Theoretical isoelectric point (pH units). c Hx ⳱ −⌺f(r,x) ⭈ ln(f(r,x)) 20 Fig. 2. Clustering of the members of the PKA-R family listed in Table I according to physicochemical parameters. The estimated molecular mass calculated from the amino acid sequence was plotted against the theoretical isoelectric point for each subunit. Inset: An expanded view of the region with the highest clustering density. where x represents each position along the alignment, r represents each choice of amino acid for the position in question, and f(r,x) is the frequency at which residue r is found at position x. Results Sequence Identification. Abbreviations in the first column in Table 1 are used in all subsequent figures and tables. Thirty full-length and eight fragments of eukaryotic R-subunit sequences were identified by an extensive search of all major databases (Table 1) using known regulatory subunits as query sequences. Only 30 complete and 8 incomplete eukaryotic sequences previously identified and annotated as PKA regulatory subunits were considered. Of the 30 complete sequences, 13 belonged to Chordata and 9 to Fungi. The rest of the sequences were from Echinodermata (2), Alveolata (2), Arthopoda (1), Mollusca (1), Nematoda (1), and Dictyosteliida/slime molds (1). Two entries from Fungi corresponded to R-subunit isoforms from Schizosaccharomyces pombe, one monomeric and one dimeric, both available from Swissprot. Although there are four isoforms of the type I R subunit from Drosophila, three monomeric and one dimeric (Kalderon and Rubin 1988), only the representative sequence (dimeric) incorporated into Swissprot was used for our analysis. Physicochemical Properties of Full-Sequence R Subunits. The size of the subunits used in the present study ranged between 325 amino acids, with a theoretical molecular mass (MW) of 37 kDa (Paramecium tetraurelia; Q94725), and 522 amino acids with a MW of 56 kDa (Ustilago maydis; P49605) (Table 1). The theoretical isoelectric points covered a range from 4.82 (Blastocladiella emersonii; P31320) to 9.17 (Colletotrichum trifolii; O42794) Most of the sequence variation occurred in the N-terminal region preceding the inhibitory sequence, which contains the variable region and, in most cases, contains a dimerization/docking domain. Plotting the predicted molecular mass versus the theoretical isoelectric point for each subunit showed a considerable degree of diversity within the PKA-R family (Fig. 2). The vertebrate type I and II subunits, and their ␣ and  subtypes formed tight and well-defined clusters (Fig. 2, inset). The nonvertebrate type I subunits from Drosophila, Aplysia, and C. elegans clustered closer to type RI␣ than to RI. The type I subunit from Dictyostelium was practically equidistant from the RI␣ and RI clusters. Vertebrate type II subunits also formed two well-defined clusters. In contrast, subunits from Echinodermata, typically considered type II subunits, formed a separate cluster that was equidistant from those for vertebrate types I and II. Finally, the subunits from Fungi, in contrast with the well-defined clustering of vertebrate type I and type II regulatory subunits, were completely dispersed. Based on these observations, it is clear that a classification of the PKA-R family based on their physicochemical properties is appropriate only for the vertebrate subunits; the approach cannot be generalized. Inhibitory Sequence Motif Identification. An inhibitory sequence conforming to the RRx[AG]⌿ motif, where ⌿ is a hydrophobic residue, was identified in each type I subunit (Fig. 1A). Conversely, an RRxS⌿ motif 21 Fig. 3. Radial phylogenetic tree of the PKA-R family. Protein abbreviations are provided in Table 1. Phylogenetic distance is approximately proportional to branch length. Analysis was performed using the neighbor-joining algorithm. The tree is based on a complete sequence alignment of the proteins listed in Table 1A. Clustering patterns are shaded in gray. ␣ and  subtypes are also indicated for type I and type II subunits. A bar for calibration of phylogenetic distances is provided at the bottom. was identified in vertebrate RII type subunits as well as R subunits from Fungi. In all the subunits from Fungi and type I subunits, the ⌿ residue was followed by a serine, whereas in all type II subunits that position was occupied by a cysteine. The R subunit from the ciliate Paramecium tetraurelia contains an atypical inhibitory sequence conforming to the pattern TRxS⌿ (Carlson and Nelson 1996). When the R subunit from Euplotes octocarinatus was scanned for suitable PKA phosphorylation sites that could function as inhibitory sequences, a novel sequence corresponding to the motif KxS⌿ was identified (Fig. 1A). Sequence Alignment and Phylogenetic Analysis. A multiple alignment of all the full-sequence subunits was performed (not shown) and a radial phylogenetic tree was built from it as described under Experimental Procedures (Fig. 3). The general clustering of proteins derived from organisms of the same kingdoms is noteworthy. For that reason, the PKA-R family was subdivided 22 Fig. 4. Phylogeny of the PKA-R family. The phylogenetic tree shown in Fig. 3 was rooted using the sequence of Euplotes octocarinatus (Eup1R) as the outgroup. Protein abbreviations are provided in Table 1. Bootstrapping values are provided. into the “type I,” “type II,” “fungi,” and “alveolate” subfamilies. In the type I and II clusters, two subclusters can be identified corresponding to the subtypes ␣ and . Tentatively, we designated the subunits from fungi as type III and those from alveolates as type IV. When an unrooted tree was constructed based on a sequence alignment of the complete cNMP superfamily (not shown), the deepest-branching clade in the PKA R subunit group was the subunit from Euplotes (Eupl R). Consequently, it was used as outgroup to root the previously described radial tree. The rooted and bootstrapped tree (Fig. 4) showed the clustering of the subunits in separate groups which were consistent with accepted phylogenetic relationships. Although bootstrapping values were relatively low (49%) for the branching between type I and type II subunits, the use of different methodologies and parameters always yielded trees with the same topology in which Fungi were always the first divergent group. Additionally, bootstrapping values were low for the branching point of the sequences of Blastocladiella (BlasR) and Ustilago (UstiR). Upon observation of the complete sequence alignment of all the R subunits (not shown), there was a remarkable lack of conservation of the dimerization/docking domains, the linker regions, and the inhibitory sequences in the most primitive organisms. Accordingly, we focused our study on the most conserved region, which corresponded to cAMP-binding domains A and B (Fig. 5). The alignment of the cAMP-binding domains was curated based on structural evidence from the crystal structures of bovine RI␣ (Su et al. 1995) and rat RII (Diller et al. 2001). Most of the variability in the cAMP-binding domains corresponded to the loop between -strand 4 and -strand 5 of both domain A and domain B, and the C-terminal region. Two distinct blocks, which corresponded to the phosphate-binding cassettes (PBCs), contained the majority of invariant residues (asterisks in Fig. 5). The PBCs are defined as the segments of each cAMPbinding domain that contain most of the key residues for cAMP-mediated activation of PKA (Figs. 1B and C and 5) (Su et al. 1995; Diller et al. 2001). Signature Sequences and Profiles. Sequence logos for PBC A and B and a logo defining the consensus PBC 23 were generated based on the sequence alignment of the cAMP-binding domains. Residue conservation was lower in positions not critical for the interaction with cAMP. Residue conservation (dots in Fig. 6A) was higher for PBC A. The occurrence of an invariant tyrosine residue in position 9 of each PBC A was especially striking, since position 9 is the most variable in PBC B, as shown by the entropy plots in Fig. 6B. The entropy plots also show that conservation is higher in the Nterminal region than in the C-terminal part of the PBC. Signature sequences for the R subunits were generated from the consensus sequence logo shown in Fig. 6. Swissprot and TreMBL were scanned with signature patterns of different length. Shortening the N terminus of the original 19-residue-long signature produced a dramatic loss of specificity. In contrast, shortening of the C terminus of the signature sequence still kept the original specificity. The minimal length of a signature capable to retain enough stringency to detect all the R subunits without any false positives was 14 residues. The signature F–G–E–[LIV]–A–L–[LIMV]–X(3)– [PV]–R–[ANVQ]–A (Table 2 and Fig. 7) can specifically identify a protein belonging to the R-subunit family (X ⳱ any residue; alternative residues at any one position are presented in brackets). When Swissprot and TrEMBL were scanned for proteins matching the signature, all the sequences and fragments of R subunits that contained a cAMP-binding domain were identified. There were no false positives. Thus, it is a genuine signature sequence and can be used to identify new members of the family as they become sequenced. Futhermore, by integrating this pattern with unique residues identified from the sequence logos that define the different PBCs (Fig. 6), it was possible to define subpatterns (Table 2). Those derived signatures are based on the three central residues in the PBC, which are the residues most distant from the cAMP molecule (Figs. 1B and C and 7). Identification of Potential R Subunits. The signature pattern generated for the PKA-R family was used to scan different databases. When the genome of Drosophila melanogaster was scanned, a new putative PKA regulatory subunit was identified (Swissprot/TreMBL AC: Q9V5E8).1 Scanning the complete genome of Drosophila revealed the presence of only one type I and one type II subunit. Additional searches were performed on the nonredundant database, dbEST, and also diverse genomic databases (Caenorhabditis elegans database at the Sanger Center, Dictyostelium discoideum database at the 1 The RII subunit from Drosophila melanogaster was deposited and annotated as a type II PKA regulatory subunit in the Swissprot and the Celera Genomics Flybase databases after the completion of these studies. Therefore, the sequence was not included in our analysis. Sanger Center, TIGR, and the collection of finished and unfinished microbial genomes at the NCBI). As a result, we identified several fragments previously not annotated as belonging to PKA regulatory subunits (data not shown), as well as two new possibly complete sequences. The putative regulatory subunit from Plasmodium falciparum (clone 3D7) corresponds to Contig 04.000625 in chromosome 12. The potential subunit from Candida albicans corresponds to Contig 5-2380. No additional subunits were identified in the partial genomes of C. elegans and Dictyostelium. No recognizable homologues were identified in Bacteria or Archaea. Discussion Two types of R subunits are generally accepted: type I and type II. While all R subunits share the same general domain organization, type I and type II subunits differ in molecular weight, isoelectric point, amino acid sequence, antigenicity, autophosphorylation capacity, cellular location, and tissue distribution. Also, they differ in their affinities for cAMP and C-subunit isoforms (Corbin et al. 1975; Øgreid et al. 1989). The original criterion to assign R subunits to types I and II was the order of elution following ion-exchange chromatography. Subsequently, types I and II were subdivided into ␣ and  subtypes based on their SDS-PAGE apparent mobilities. This classification was formulated from a very limited number of mammalian sequences available at the time. After that, nonmammalian subunits were included into one of those groups based on sequence similarity. While this methodology is adequate in some cases, in the PKA-R family it leads to significant inconsistencies. Our analysis indicates that traditional classifications based on physicochemical properties such as isoelectric point/charge and molecular weight are not adequate. Inhibitory sequence motifs are another criterion used to classify the PKA-R family. The inhibitory site for the mammalian type II isoform contains the “RRxS⌿” consensus sequence for phosphorylation by the PKA catalytic subunit (Rosen and Erlichman 1975). In contrast, the inhibitory site for type I isoforms contains a nonphosphorylatable pseudo-substrate site in which the serine found in type II motif is replaced by a small hydrophobic amino acid residue. One important characteristic of the inhibitory site of animal type II isoforms is a conserved cysteine residue that can form intermolecular disulfide bonds with cysteine residues at the active sites of C subunits (First et al. 1988). To fit the R subunit from Fungi in the classic type I/type II schema, the length of the inhibitory sequence motif is conventionally set to five residues. If the motif is expanded to include the residue following the ⌿ residue, it is apparent that in all the subunits from Fungi, the cysteine residue charac- 24 Fig. 5. Partial multiple sequence alignment of the sequences shown in Table 1A. For method of alignment see text. Alignments are restricted to the C-terminal region of the R subunits that contains the cAMP-binding domains. Dashes indicate gaps introduced during the alignment process. Straight arrows indicate -strands and curvy arrows indicate ␣-helices, according to the bovine RI␣ crystal structure (Su et al. 1995). The boundaries of the phosphate-binding cassettes (PBC) A an B are indicated. All sequences are listed under the abbreviated names indicated in Table 1. Dots under the PBC bars indicate residues that are critical for the activation of the RI␣ subunits. Residues indicated by arrows in domains A and B may be important for interdomain communication. Sequence conservation is shown under the alignment according to the following key: (*) invariant; (:) conserved; (.) partially conserved residue. 25 26 Fig. 6. Sequence logos and entropy plots. A Sequence logos defining PBC A and B, and PBC consensus describing the general residue distribution in any PBC belonging to a protein from the PKA-R family. The bar over the PBC consensus logo indicates the region that was used for the generation of the PKA-R family signature pattern. Dots in the PBC A and B logos indicate residues that are invariant in each PBC. The horizontal axis represents the position of the residue within the PBC motif. The vertical axis represents the amount of information (in bits) that this position holds. The height of the one-letter residue symbol at each position is proportional to the information content of the residue at that position. B Entropy plots corresponding to the sequences depicted in column A. The horizontal axis represents the position of the residue within the PBC motif, whereas the vertical axis represents the entropy or lack of predictability for each alignment position. teristics of all animal type II subunits is replaced by an invariant serine, as in type I subunits. Thus, based on the extended six-residue motif, the subunits from Fungi could be classified as a separate type. However, if differences in the inhibitory were applied as a classification criterion was to the R subunits from Alveolates, each one should be included in a separate group (Paramecium TRxS⌿, Euplotes IKxS⌿, Plasmodium KKxS⌿), or they should all be included in a separate group for “nonconforming” R subunits. In conclusion, the inhibitory sequence cannot be consistently used as a satisfactory criterion to classify the PKA-R family. Phylogenetic trees are more revealing than raw homology data, because they integrate the data of all pairs of protein sequences. Therefore, in the present study, we have used clustering analysis based on phylogenetic trees to classify the PKA-R family. Clustering based on phy- logenetic analysis revealed the existence of four groups/ types. Two of them corresponded to classical type I and II subunits, and the other two to Fungi (designated type III) and Alveolates (designated type IV). The proposed designations are a compromise between the most commonly used nomenclature and the fact that the Fungi and Alveolate clusters have characteristics that clearly set them apart from the type I/type II classification. Our phylogenetic analysis indicates that it is likely that an ancestral R subunit gave rise to type IV subunits before the Metazoa and Fungi lineages separated. All alveolate subunits contain atypical inhibitory sequence motifs and lack dimerization domains. These characteristics suggest that these subunits descended early from an ancestral R subunit. Phylogenetic trees indicate that the emergence of multiple paralogous R subunits (types I and II and subtypes ␣ and ) occurred late in the evo- 27 Table 2. Signature sequence for the cAMP-dependent protein kinase regulatory subunit family (PKA-R) and variable residues in the phosphatebinding cassette defining PKA-R subfamilies Family Signature sequence PKA-R 1 2 3 4 5 6 7 8 9 10 11 12 13 14 F–G–E–[LIV]–A–L–[LIMV]–X1–X2–X3–[PV]–R–[ANVQ]–A PBC A Type & subfamily X1 X2 PBC B X3 Type I␣ mammal X1 X2 X3 M N Type I mammal T G R L Type I nonmammal D Y Type I Dictyostelium S T Type II (␣/) T N K [NDEH] [RKTLAE] N Types III & IV [ASV] Fig. 7. Spatial distribution of the residues that define the signature sequence for the PKA-R family. The diagram schematically depicts the residue distribution in the structural elements that define the PBCs and their relative positions with respect to the cAMP molecule. Lines indicate hydrogen-bonding between the selected residues and cAMP. lutionary process, after the divergence of Metazoa and Fungi. The number of paralogous mammalian R subunits may be explained by multiple gene duplication events. This phenomenon may have occurred in response to the need to maintain a stricter homeostasis and elaborate intercellular communication networks in metazoans. The rooted phylogenetic tree created using the sequence of Euplotes octocarinatus as outgroup showed a [NDHKS] branching pattern that was consistent with known functional differences between R subunits, as well as accepted phylogenetic relationships (Fig. 4, right). According to this tree, Fungi diverged before the Animals, and before the gene duplication event that resulted in the emergence of type I and type II subunits. The points of divergence of Mollusca (Aplysia), Arthropoda (Drosophila), and Nematoda (C. elegans) conformed to accepted phylogeny (Davidson et al. 1995). The presence of single type I and type II subunits in Drosophila suggests that the subsequent gene duplications that resulted in the appearance of subtypes ␣ and  occurred after Arthropoda diverged. The branching point of Dictyostelium has been the subject of ample controversy. Most phylogenetic analysis based on rRNA (McCarrol et al. 1983; Hasegawa et al. 1985; Herzog and Maroteaux 1986; Hendricks et al. 1991; Douglas et al. 1991) and some based on protein sequence analysis (Kuma et al. 1995; Baldauf and Doolittle 1997; Baldauf et al. 2000) place Dictyostelium as outgroup to Fungi and Animals. Conversely, other models based on protein sequence analysis suggest that Fungi diverged from the line leading to Animals before the divergence of Dictyostelium and Animals (Loomis and Smith 1990, 1995; Roger et al. 1996; Kalhor et al. 1999; Norian et al. 1999; Swigart et al. 2000). Our analysis is consistent with the latter model. It suggests that Dictyostelium is more closely related to Animals than to Fungi and that it branched from the line leading to Metazoa after Fungi. Signature sequences are of great diagnostic and practical value for an immediate assignment of newly sequenced subunits to a common classification scheme. We have identified a global signature sequence (Table 2) 28 common to all PKA-R members and suitable for discriminating R subunits from other cyclic nucleotidebinding proteins. Since the general structure of the PBC is very similar between R and other cAMP-binding proteins, such as the bacterial regulator CAP (McKay et al. 1982), it is remarkable that such short signature sequence is capable of discriminating between members of the PKA-R family and other cAMP-binding regulators such as the NFR, CAP, or PKG families. Characteristically, CAP has a PBC with an extra residue that lies within the restrained, solvent-accessible loop of the PBC, i.e., in the X1–X2–X3 region of the signature sequence (Fig. 7). That part of the PBC, the most distant from the phosphate of the cAMP molecule, is also the most variable between the R subfamilies. A notable exception is Tyr in position 8 of PBC A. That residue is invariant in all PBC A, whereas it corresponds to the most variable position in PBC B. This is consistent with structural information that indicates that this tyrosine lies at the center of a complex network of contacts responsible for interdomain communication. The residue in position 14 of the signature sequence is extremely important to determine cyclic nucleotide specificity. Replacement of an Ala residue in that position with a Thr produces a drastic change in selectivity for cGMP vs cAMP (Weber et al. 1989). Residue 14 is a conserved Ala in all PBC A, but only in 27 of 31 PBC B. The sequences that do not have an Ala in that position, and could potentially be activated by both cAMP and cGMP, are Blastocladiella (V), S. pombe (N), and Saccharomyces (Q) (Fig. 7). Experimental evidence indicates that the subunit from Saccharomyces can be activated by both cAMP and cGMP (Cytrynska et al. 1999). In conclusion, this study shows that R subunits can be classified taking into consideration their phylogeny and that the PBC regions can be used to establish family- and subfamily-specific signatures. Our observations have profound implications regarding the use of biological model systems to study cAMP-mediated signaling. Accordingly, Drosophila could provide an ideal model for the study of type I and type II subunit-mediated regulation of PKA, whereas Dictyostelium constitutes a suitable model for type I subunit-related studies. Attempts to extrapolate from observations in yeast should take into consideration that the R subunits from Fungi belong to a separate type. Most studies classify the R subunits from yeast as type II, when in fact those R subunits are distinct and related to both type I and type II subunits. The variability in the residue responsible for cAMP–cGMP specificity in several fungal subunits also suggests that caution should be used in the exclusive attribution of PKA-mediated cellular responses to cAMP. We are confident that this study will contribute to establishing a standardized categorical classification and nomenclature of the PKA-R family and stimulate comparative studies on the evolution of this protein family, in itself or in the global context of the evolution of cyclic nucleotide-mediated protein kinase signaling. Acknowledgments. This work was supported in part by USPHS Grant GM34921 to S.S.T. Sequence data for Plasmodium falciparum chromosome 12 was obtained from the Stanford DNA Sequencing and Technology Center (http://www-sequence.stanford.edu/group/malaria). Sequencing of Plasmodium falciparum chromosome 12 was accomplished as part of the Malaria Genome Project with support by the Burroughs Wellcome Fund. Sequence data for Candida albicans were generated at the Stanford DNA Sequencing and Technology Center with the support of the NIDR and the Burroughs Wellcome Fund. We wish to thank Anna Canaves for her assistance in manuscript preparation and graphics design. References Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res 25:3389–3402 Appel RD, Bairoch A, Hochstraser DF (1994) A new generation of information retrieval tools for biologists—The example of the Expasy WWW server. Trends Biochem Sci 19:258–260 Baldauf SL, Doolittle WF (1997) Origin and evolution of the slime molds (Mycetozoa). Proc Natl Acad Sci USA 94:12007–12012 Baldauf SL, Roger AJ, Wenk-Siefert I, Doolittle WF (2000) A kingdom-level phylogeny of eukaryotes based on combined protein data. Science 290:972–977 Carlson GL, Nelson DL (1996) The 44 kDa regulatory subunit of Paramecium cAMP-dependent protein kinase lacks a dimerization domain and may have a unique autophosphorylation site sequence. J Eukaryot Microbiol 43:347–356 Corbin JD, Keely SL, Park CR (1975) The distribution and dissociation of cyclic adenosine 3⬘:5⬘-monophosphate-dependent protein kinases in adipose, cardiac, and other tissues. J Biol Chem 250:218– 225 Cytrynska M, Wojda I, Franjt M, Jakubowicz T (1999) PKA from Saccharomyces cerevisiae can be activated by cyclic AMP and cyclic GMP. Can J Microbiol 45:31–37 Davidson EH, Peterson KJ, Cameron RA (1995) Origin of bilaterian body plans: Evolution of developmental regulatory mechanisms. Science 270:1319–1325 Dell’Acqua ML, Scott JD (1997) Protein kinase A anchoring. J Biol Chem 272:12881–12884 Diller TC, Madhusudan, Xuong NH, Taylor SS (2001) Molecular basis for regulatory subunit diversity in cAMP-dependent protein kinase: Crystal structure of the type II beta regulatory subunit. Structure 9:73–82 Døskeland SO, Øgreid D (1984) Characterization of the interchain and intrachain interactions between the binding sites of the free regulatory moiety of protein kinase I. J Biol Chem 259:2291–2301 Douglas SE, Murphy CA, Spencer DF, Gray MW (1991) Cryptomonad algae are evolutionary chimaeras of two phylogenetically distinct unicellular eukaryotes. Nature 350:148–151 Eron L, Arditti R, Zubay G, Connaway S, Beckwitt JR (1971) An adenosine 3⬘:5⬘-cyclic monophosphate-binding protein that acts on the transcription process. Proc Natl Acad Sci USA 68:215–218 Fantozzi DA, Harrotunian AT, Wen W, Taylor SS, Feramisco JR, Tsien RY, Meinkoth JL (1994) Thermostable inhibitor of cAMPdependent protein kinase enhances the rate of export of the kinase catalytic subunit from the nucleus. J Biol Chem 269:2676–2686 Felsenstein J (1985) Confidence limits on phylogenies: An approach using the bootstrap. Evolution 39:783–791 First EA, Bubis J, Taylor SS (1988) Subunit interaction sites between 29 the regulatory and catalytic subunits of cAMP-dependent protein kinase: Identification of a specific interchain disulfide bond. J Biol Chem 263:5176–5182 Gonnet GH, Cohen MA, Benner SA (1994) Analysis of amino acid substitution during divergent evolution—The 400 by 400 dipeptide substitution matrix. Biochem Biophys Res Commun 199:489–496 Hall TA (1999) BioEdit: A user-friendly biological sequence alignment editor and analysis package program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98 Hasegawa M, Iida Y, Yano T, Takaiwa F, Iwabuchi M (1985) Phylogenetic relationships among eukaryotic kingdoms inferred from ribosomal RNA sequences. J Mol Evol 22:32–38 Hendricks L, De Baere R, Van de Peer Y, Neefs J, Goris A (1991) The evolutionary position of rhodophyte Porphyra umbilicalis and the basidiomycete Leucosporidium scottii among other eukaryotes as deduced from complete sequences of small ribosomal subunit RNA. J Mol Evol 32:167–177 Henikoff S, Henikoff JG (1992) Amino acid substitution matrices from protein blocks. Proc Natl Acad Sci USA 89:10915–10919 Herzog M, Maroteaux L (1986) Dinoflagelate 17S rRNA sequence inferred from the gene sequence: Evolutionary implications. Proc Natl Acad Sci USA 83:8644–8648 Kalderon D, Rubin GM (1988) Isolation and characterization of Drosophila cAMP-dependent protein kinase genes. Genes Dev 2:1539– 1556 Kalhor HR, Niewmierzycka A, Faull KF, Yao X, Grade S, Clarke S, Rubenstein PA (1999) A highly conserved 3-methylhistidine modification is absent in yeast actin. Arch Biochem Biophy 370:105– 111 Kawasaki H, Springett GM, Mochizuki N, Toki S, Nakaya M, Matsuda M, Housman DE, Graybiel AM (1998) A family of cAMP-binding proteins that directly activate Rap1. Science 282:275–279 Kimura M (1983) The neutral theory of molecular evolution. Cambridge University Press, Cambridge, UK Kuma K, Nikoh N, Iwabe N, Miyata T (1995) Phylogenetic position of Dictyostelium inferred from multiple protein data sets. J Mol Evol 41:238–246 Loomis WF, Smith DW (1990) Molecular phylogeny of Dictyostelium discoideum by protein sequence comparison. Proc Natl Acad Sci USA 87:9093–9097 Loomis WF, Smith DW (1995) Consensus phylogeny of Dictyostelium. Experientia 51:1110–1115 Ludwig J, Margalit T, Eismann E, Lancet D, Kaupp UB (1990) Primary structure of cAMP-gated channel from bovine olfactory epithelium. FEBS Lett 270:24–29 McCarrol R, Olsen GJ, Stahl YD, Woese CR, Sogin ML (1983) Nucleotide sequence of the Dictyostelium discoideum small-subunit ribosomal ribonucleic acid inferred from the gene sequence: Evolutionary implications. Biochemistry 22:5858–5868 McKay DB, Weber IT, Steitz TA (1982) Structure of catabolite gene activator protein at 2.9 A resolution: Incorporation of amino acid sequence and interactions with cyclic AMP. J Biol Chem 257: 9518–9524 Nakamura T, Gold GH (1987) A cyclic nucleotide-gated conductance in olfactory receptor cilia. Nature 325:442–444 Nicholas KB, Nicholas HB Jr, Deerfield DW II (1997) GeneDoc: Analysis and visualization of genetic variation. EMBNET.NEWS 4:14 Norian L, Dragoi IA, O’Halloran T (1999) Molecular characterization of rabE, a developmentally regulated Dictyostelium homolog of mammalian rab GTPases. DNA Cell Biol 18:59–64 Øgreid D, Ekanger R, Suva RH, Miller JP, Døskeland SO (1989) Comparison of the two classes of binding sites (A and B) of type I and type II cyclic AMP-dependent protein kinases by using cyclic nucleotide analogs. Eur J Biochem 181:19–31 Page RDM (1996) TREEVIEW: An application to display phylogenetic trees on personal computers. CABIOS 12:357–358 Robinson-Steiner AM, Corbin JD (1983) Probable involvement of both intrachain cAMP binding sites in activation of protein kinase. J Biol Chem 258:1032–1040 Roger AJ, Smith MW, Doolittle RF, Doolittle WF (1996) Evidence for the Heterobolosea from phylogenetic analysis of genes encoding glyceraldehyde-3-phosphate dehydrogenase. J Euk Microbiol 43: 475–485 Rosen OM, Erlichman J (1975) Reversible autophosphorylation of a cyclic 3⬘:5⬘-AMP-dependent protein kinase from bovine cardiac muscle. J Biol Chem 250:7788–7794 Saitou N, Nei M (1987) The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425 Schneider TD, Stephens RM (1990) Sequence logos: A new way to display consensus sequences. Nucleic Acids Res 18:6097–6100 Shabb JB, Corbin JD (1992) Cyclic nucleotide-binding domains in proteins having diverse functions. J Biol Chem 267:5723–5726 Su Y, Dostmann WRG, Herberg FW, Durick K, Xuong N-h, Ten Eyck L, Taylor SS, Varughese KI (1995) Regulatory subunit of protein kinase A: Structure of deletion mutant with cAMP binding domains. Science 269:807–813 Swigart P, Insall R, Wilkins A, Cockcroft S (2000) Purification and cloning of phosphatidylinositol transfer proteins from Dictyostelium discoideum: Homologues of both mammalian PITPs and Saccharomyces cerevisiae sec14p are found in the same cell. Biochem J 347:837–843 Takio K, Smith SB, Krebs EG, Walsh KA, Titani K (1982) Primary structure of the regulatory subunit of type II cAMP-dependent protein kinase from bovine cardiac muscle. Proc Natl Acad Sci USA 79:2544–2548 Taylor SS, Buechler JA, Yonemoto W (1990) cAMP-dependent protein kinase: framework for a diverse family of regulatory enzymes. Annu Rev Biochem 59:971–1005 Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, positions-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680 Weber IT, Steitz TA (1987) Structure of a complex of catabolite gene activator protein and cyclic AMP refined to 2.5A resolution. J Mol Biol 198:311–326 Weber IT, Shabb JB, Corbin JD (1989) Predicted structures of the cGMP binding domains of the cGMP dependent protein kinase—A key alanine threonine difference in evolutionary divergence of cAMP and cGMP bindings sites. Biochemistry 28:6122–6127 Zheng Z, Schäffer AA, Miller W, Madden TL, Lipman DJ, Koonin EV, Altschul SF (1998) Protein sequence similarity searches using patterns as seeds. Nucleic Acids Res 26:3986–3990