* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download 122.1 Schramm
History of biological warfare wikipedia , lookup
Virtual karyotype wikipedia , lookup
Genomic imprinting wikipedia , lookup
Bioinformatics wikipedia , lookup
Community fingerprinting wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Gene nomenclature wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
Gene expression wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Gene therapy wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Gene regulatory network wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Neurogenomics wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Gene prediction wikipedia , lookup
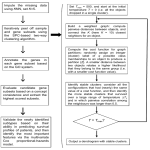
Ref ID: 122.1 Prediction of Clinical Outcome and Biological Subclassification of Primary Neuroblastomas by Expression Profiling Alexander Schramm, Johannes H Schulte, Ludger Klein-Hitpass, Hauke Sieverts, Bernd Berwanger, Holger Christiansen, Thorsten Simon, Frank Berthold, Werner Havers, Patrick Warnat, Benedikt Brors, Jürgen Eils, Roland Eils, Angelika Eggert Children´s Hospital, University Clinic Essen, NRW, Germany. The development of risk-adapted and more effective therapy strategies in neuroblastoma requires further improvements in accurate risk assessment. Although numerous prognostic factors have been identified, precise risk evaluation in individual neuroblastoma patients remains difficult. To define a reliable predictor for event-free survival after first-line therapy and to identify gene signatures characteristic for clinical and biological subgroups, we performed expression profiling of 68 primary neuroblastomas of all stages by Affymetrix HG-U95Av2 arrays. Expression data from subgroups were analyzed using support vector machines with a radial basis function (SVM-rbf), k-nearest neighbors (KNN) algorithms or multiple decision trees. Significance analyses of microarrays (SAM) was applied to search for genes and gene patterns differentially expressed between subgroups. Surprisingly, expression profiles of stage 4 and stage 4s neuroblastomas could not clearly be distinguished by any of the mathematical methods applied. In contrast, MYCN-amplification as well as high expression of the neurotrophin receptor TrkA demonstrated a strong association with specific gene expression patterns. Recurrence of neuroblastoma within two years of diagnosis was correctly predicted in 81% of cases using SVM-rbf or KNN. Sensitivity and specificity of the predictor were 72% and 89%, respectively. Gene patterns associated with eventfree survival were found to be rather homogeneous, while patterns for recurrent disease appeared to be diverse. We conclude, that SVM-rbf and KNN are suitable tools for prediction of clinical outcome and biological subclassification in neuroblastoma. Predictive gene signatures have to be evaluated prospectively in larger study cohorts. Presentation mode(s): ORAL-PRESENTATION – PC-PROJECTOR