* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download DNA analysis - Madeira City Schools
Zinc finger nuclease wikipedia , lookup
Gene therapy wikipedia , lookup
Gene expression profiling wikipedia , lookup
Metagenomics wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Gene prediction wikipedia , lookup
Real-time polymerase chain reaction wikipedia , lookup
Non-coding DNA wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Restriction enzyme wikipedia , lookup
DNA vaccination wikipedia , lookup
DNA supercoil wikipedia , lookup
Molecular cloning wikipedia , lookup
Genome editing wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Designer baby wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Genetic engineering wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
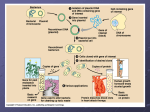
We’re going to look at the following: DNA analysis: this makes it possible to examine variation among individuals. Why would you want to do this? Genetic engineering: manipulating DNA or organisms to perform practical tasks or provide useful products I. DNA analysis: Result of an electrophoresis A. Most common uses are: 1. analyze a person’s genes (looking for diseases) 2. compare the sequence of nitrogen bases among individuals (paternity and crime scenes) B. Use restriction enzymes 1. A restriction enzyme is an enzyme that cuts DNA at specific “recognition sites” 2. In nature restriction enzymes protect bacteria against intruding DNA (viruses) by cutting the intruders DNA (can’t function) C. Use an electrophoresis apparatus to separate the cut DNA 1. Fancy way to do chromatography D. Process of DNA analysis 1. DNA is obtained (hair, blood, skin cells…) 2. Copies of DNA are made using the PCR method (polymerase chain reaction). This is just a complex way to make a ton of DNA copies 3. Restriction enzymes are added to cut the DNA into fragments at very specific sequences. 4. DNA has an overall negative charge 5. Put DNA into wells of gel at the negative end of the chamber 6. Turn on power supply 7. Smaller fragments will travel faster through gel, therefore they will travel the farthest in the allotted time. Gel that is loaded with DNA and reading to “run” No DNA in these wells DNA is present in these wells II. Genetic Engineering A. An overview: 1. Use bacteria plasmids - small circular DNA that replicate within the bacterial cell. These are isolated. 2. The plasmid and gene of choice are both cut using the same restriction enzyme (therefore cutting at the same recognition site) b. this produces what we call “sticky ends” 3. The plasmid and gene of choice are put in a test tube together 4. DNA ligase (an enzyme) is added fuse the two strands together 5. Plasmid is put back into bacteria 6. Bacteria reproduces 7. Bacteria will express the inserted gene by making the protein GAATTC CTTAAG G CTTAA AATTC G B. What you will be doing in your lab: 1. Gene from a bioluminescent jellyfish has been isolated a. it produces a green fluorescent protein (GFP) 2. It has been inserted into a bacterial plasmid. a. the recombinant DNA molecule is called “pGLO” plasmid 3. The plasmid will (hopefully) transform into bacteria that will then express the inserted gene and therefore glow. a. Genetic transformation = change caused by genes C. Things you need to know to understand what you are doing 1. Inserting the gene is not enough…you have to have something to turn it on 2. Turning on and off genes (gene expression) is carefully regulated to allow for adaptation to different conditions a. prevents wasteful production of unneeded proteins b. digestive enzymes are highly regulated 3. Bacteria regulates their gene expression through “operons” a. an operon is a cluster of genes controlled by one promoter The Arabinose Operon araC araB araA araD OFF araC araB araA araD ON The pGLO plasmid araC GFP ON Glowing 4. An ampicillin resistant gene is also added to your pGLO plasmid a. this is a gene that makes a protein that allows the bacteria to grow in the presence of the antibiotic therefore making it “resistant” ampicillin, b. This insures that only the bacteria that was transformed will grow…nothing else. c. It is a way to control your experiment and not add more variables. GFP – makes Origin – where transcription starts The pGLO plasmid green flourescent protein Bla gene – makes bacteria resistant to ampicillin What you will see in lab: LB = Luria broth (food for bacteria) AMP = Ampicillin (antibiotic -- kills bacteria) ARA= Arabinose (sugar -- turns on the pGLO gene) LB LB/AMP LB/AMP LB/AMP/ ARA Plasmid added --- --- + + Purpose Control for bacterial growth Control for effectiveness of ampicillin Control for effectiveness of transformation End result Does bacteria gRow Does bacteria gLow