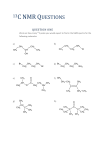
* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Nuclear Magnetic Resonance
Superconducting magnet wikipedia , lookup
Electromagnetism wikipedia , lookup
Relativistic quantum mechanics wikipedia , lookup
Magnetic monopole wikipedia , lookup
Magnetometer wikipedia , lookup
Earth's magnetic field wikipedia , lookup
Giant magnetoresistance wikipedia , lookup
Electromagnetic field wikipedia , lookup
Magnetotactic bacteria wikipedia , lookup
Neutron magnetic moment wikipedia , lookup
Electromagnet wikipedia , lookup
Force between magnets wikipedia , lookup
Magnetoreception wikipedia , lookup
Multiferroics wikipedia , lookup
Magnetohydrodynamics wikipedia , lookup
Magnetotellurics wikipedia , lookup
History of geomagnetism wikipedia , lookup
Electron paramagnetic resonance wikipedia , lookup
Magnetochemistry wikipedia , lookup
Ferromagnetism wikipedia , lookup
Nuclear magnetic resonance wikipedia , lookup
Two-dimensional nuclear magnetic resonance spectroscopy wikipedia , lookup
Nuclear magnetic resonance spectroscopy wikipedia , lookup
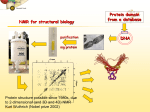
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Nuclear Magnetic Resonance Spectroscopy 核磁共振光譜 蘇士哲 19/Nov/2013 NMR, the Lord of spectrometers • • • • • Resolution to atomic level. Enable to determine structure. Dynamic information in wide time scale. Chemical modification is not required. In solution condition, more close to biology. e Pieter Zeeman Zeeman effect • This splitting of spectral line into several components in the presence of magnetic field is called Zeeman (1902, Nobel Prize in physics) effect. Nuclear has Spin, too • In 1945 the groups of both Bloch (Stanford) and Purcell (Harvard) succeeded in detecting nuclear magnetic resonance absorption in bulk matter. – The energy absorption was observed by irradiating the sample with radiofrequency field and varying the strength of the magnetic field (continue wave). – 1952 Nobel prize in physics. Felix Bloch Edward Mills Purcell Nuclear Spin Nuclear spin is the total nuclear angular momentum quantum number (spin number), characterized by a quantum number I. Different nuclear has different number I, which only can be integral, half integral or zero. The spin number I represents the magnetic quantum number mI of - I, I + 1, ….+ I. Different mI represents one energy state in magnetic field. The number of possible orientations is give by 2I +1. Not all nuclei have “spin”. Think about I = 0, there is only one the energy state. 12C, 16O, 32S When I = 1/2, there are two energy states (only one energy transition). 1H, 13C, 15N, 31P When I > 1/2, there are 2I + 1 states. 2H (I = 1), 14N (I = 1), 17O (I = 5/2) The energy gap • The two states have magnetic quantum numbers mI = 1/2 and mI = -1/2. 1 1 E [( ) H] [( ) H] H 2 2 h H H 2 The strength of magnetic field H B0 2 H = 26.7519 x 107 (T-1 S-1) • If 300 MHz for 1H, that is ~ 7 Tesla (~ 7 x 104 Gauss). The strength of the magnetic field of the Earth is only ~ 50 uT (0.5 Gauss) • 13C • Super magnet to enlarge the energy gap and 15N: 13C = 6.7283 x 107 and 15N = -2.712 x 107 ? The location of NMR signal High energy Low energy • Richard R. Ernst - The Nobel Prize in chemistry 1991, “for his contribution to the development of high resolution nuclear magnetic resonance (NMR) spectroscopy” • Develop Fouier transform techniques in NMR (FT-NMR) Richard R. Ernst Free induction decay (FID) and Fouier transform (FT) (A) One high energy pulse (B) FT transform, from time domain data to frequency domain Jean Baptiste Fourier 1768-1830 The advantages of FT-NMR • Fast • Multiplex • Informative: frequency, intensity and line width • Signal averaging Modern Fourier transform NMR spectrometer Coil and superconductor LN2 and LHe2 tank Typical protein proton 1D spectrum Typical DNA proton 1D spectrum Chemical Shift • When an external field is applied by a magnet, the different protons in a molecule will see different “effective” magnetic fields because of the other fields induced in the molecule itself. • At constant external field, the protons absorb at different frequencies depending on local variation in the field. • The frequency of resonance is given by: obs (B0/2)(1) shielding constant shielded by electron cloud , withdraw electron cloud In practice, we use a reference compound to define the zero point, such as tetramethylsilane (TMS, Si(CH3)4) or sodium 2,2-methyl-2silapentane-5-sulfonate (DSS). The protons is highly shielded. Downfield Less shielded Low electron density Higher magnetic strength Higher frequency Upfield Shielded High electron density Lower magnetic strength Lower frequency Multidimensional NMR spectroscopy 1D Why do we go beyond one dimension? • To resolve the crowded signals in 1D spectrum by spreading them into other dimensions. • Homo- and hetero- nuclear NMR. • 2D, 3D and 4D spectra. • To elucidate the “through-bond” and “through-space” relationships between the spins in the molecules. Three most well-known Two-dimensional homonuclear NMR spectra TOCSY Mix NMR signal on the evolution period t1 A example of 2D COSY: 1D & 2D 2D COSY v.s. 2D TOCSY: through bond N 2D NOESY: through space Nuclear Overhauser Effect (NOE) – A change in signal intensity for one type of nucleus is caused by irradiation of another type of nucleus that is nearby. – Duo to the local magnetic fluctuation. – The NOE is a dipole-dipole interaction that depends on the distance between two nuclei as r -6. – The NOE can be used to measure interproton distance up to 5-6 Å. – Most important information for structure determination by NMR. Ready for Heteronuclear NMR: Isotopelabeling of proteins (15N, 13C labeling) • Grow proteins on minimal media (M9) with 15NH4Cl as the sole nitrogen source and 13C-glucose as the sole carbon source. Pulse program of 2D Heteronuclear Single Quantum Coherence 2D 1H-15N HSQC 2D 1H-13C HSQC -13CH3 -13CH -13CH2 - 2D 1H-15N HSQC 15N (ppm) 1H (ppm) Case study A a.a. 1-240 a.a. 1-100 a.a. 101-240 Case study B Case study C From 2D to 3D: one block combines another block Pair-wise backbone experiment and sequential assignment Assignment Strategy Experiments for structural determination Three pairs of spectra for backbone assignment: HNCA/HN(CO)CA CBCANH/CBCA(CO)NH HNCO/HN(CA)CO More Spectra for side chain assignment: HCC(CO)NH/CC(CO)NH HBHA(CO)NH 15N HSQC-TOCSY HCCH-TOCSY ……. More NOESY Spectra for structural determination 15N NOESY-HSQC 13C NOESY-HSQC Example for Modern NMR pulse program, CBCANH NOESY for obtaining NOE constrains One cross peak represents one distance constrain from a proton pair. Protein Secondary structure elements -helix v.s. -sheet Structure determination 1. 2. 3. 4. Preparation of protein sample NMR data acquisition Resonance assignment, NOE assignments, and collection of other geometric restraints. Structure modeling and refinement NMR and Dynamics Protein-Protein complex interaction Li et al. JMB (2004) 335:371-381 NMR in drug discovery by line broadening & relaxation Time Fejzo et al. Chemistry & Biology, 1999, 6:755-769 Fragment-based NMR screening (SAR by NMR, Fesik) Shuker et al. Science, 1996, 274:1531-1534 Metabonomics Nature review In drug discovery 2002 Chemical exchange 1 1 A k , if k < obs T2 T2 1 1 2 , if k > obs T2 T2 8k | A B | 2 1 1 T2 2 ka A kb B ionization States and pH A-+H+ k1 k-1 0 [H ] [H ] K a 0 When , K a [H ] 2 AH