* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download GST SF in E. coli - Institute for Genomic Biology
E. coli long-term evolution experiment wikipedia , lookup
Gene expression profiling wikipedia , lookup
Protein (nutrient) wikipedia , lookup
Magnesium transporter wikipedia , lookup
Protein moonlighting wikipedia , lookup
Secreted frizzled-related protein 1 wikipedia , lookup
Community fingerprinting wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Western blot wikipedia , lookup
Proteolysis wikipedia , lookup
Protein adsorption wikipedia , lookup
Gene regulatory network wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Metabolic network modelling wikipedia , lookup
Amino acid synthesis wikipedia , lookup
List of types of proteins wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
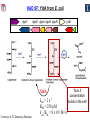
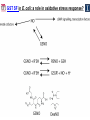
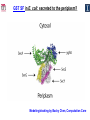
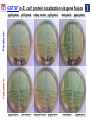
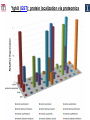
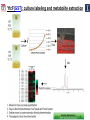
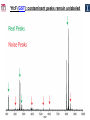
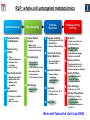
Microbiology/Metabolomics Core John Cronan and Jonathan Sweedler Enzyme Function Initiative (EFI) Advisory Committee Meeting November 30, 2011 Outline • Experimental scope • Infrastructure • Targets • YidA from E. coli (HAD) • YghU, YfcF, and YfcG from E. coli (GST) • RuBisCO-like protein from R. rubrum (ENO) • Future Directions Experimental Scope Phenomics Transcriptomics Conditions for target expression Metabolomics Verification of hypothesized enzyme-catalyzed reaction and/or evidence from relevant pathway (We now do qRT-PCR on each gene of interest) Bochner, B.R. (2003) New technologies to assess genotype-phenotype relationships, Nature Rev. Genetics. 4, 309-314. Infrastructure Personnel Instrumentation • John Cronan (Microbiology) • Jonathan Sweedler (Metabolomics) • Microbiology • • • • • Brad Evans (Metabolomics) McKay Wood (Micro/Meta) Kyuil Cho (Metabolomics) Ritesh Kumar (Micro) Amy Jones (Micro) – Biolog Omnilog phenotype microarray plate reader/incubator – Growth curve-ometer, BioscreenC – E. coli single gene KO collection (Keio collection) • Metabolomics – 11 Tesla LTQ-FT LC-MS – High resolution QTOF LC-MS – Custom XCMS LCMS data analysis platform for untargeted metabolomics Targets from around the EFI • AHS: – E. coli • GST: – E. coli • SsnA, Php, TatD, YahJ, YjjV, HyuA, YcdX, Ade – B. halodurans • LisM-RP • ENO: – E. coli • YfcG, YghU, YqjG, YliJ, YfcF, YncG, YibF, YecN • HAD: – E. coli • YidA, YigB, YbjI, NagD – P. fluorescens • GudX, RspA, YcjG, YfaW – B. cereus • NSAAR – S. enterica • ManD-RP – A. tumefaciens • 1RVK, 2NQL, GlucDRP, Atu0270, Atu4120, Atu3139, Atu4196… • 3M9L • IS: – A. tumefaciens • IspB – C. glutamicum • gi# 19551716 – B. fragilis • gi# 53711383 HAD SF: YidA from E. coli dgoT dgoD dgoA dgoK dgoR yidA YidA Courtesy of D. Dunaway-Mariano kcat = 2 s-1 KM = 250 μM kcat/KM = 8 x 103 M-1s-1 Toxic if concentration builds in the cell! YidA (HAD): no effect after addition of galactonate glycerol + galactonate succinate + galactonate glucose + galactonate YidA KO likely mutated during lag YidA(HAD): long lag when cells are resuspended in galactonate YidA (HAD): LCMS results for KDGP Validated with standard from Hua Huang in the DDM Lab YidA from E. coli (HAD): Results and Conclusions • Phenomics is difficult with HAD SF members, as many are promiscuous housekeeping phosphatases • An abrupt shift from a relatively poor carbon source to galactonate as sole carbon source causes the YidA KO to display a growth lag – The “abruptness” may be important for quickly building levels of the toxic metabolite, KDGP – Growth of YidA following the lag may be due to mutation • Metabolomics efforts so far do not support the connection between YidA KO lag with elevated KDGP levels GST SF in E. coli: a role in oxidative stress response? YfcF and YfcG (GST): NO sensitivity in null mutants GST SF in E. coli: secreted to the periplasm? Modeling/docking by Backy Chen, Computation Core GST SF in E. coli: protein localization via gene fusion yfcG-phoA empty vector yqjG-phoA treA-phoA gapA-phoA yghU-lacZ yfcG-lacZ empty vector yqjG-lacZ treA-lacZ gapA-lacZ Cytoplasm Periplasm yghU-phoA YghU (GST): protein localization via proteomics YfcF(GST): culture labeling and metabolite extraction Ions from WT Ions from mutant YfcF (GST): differential labeling provides higher accuracy YfcF (GST): contaminant peaks remain unlabeled YfcF (GST): affect of nitric oxide on metabolites GST SF: results and conclusions • YfcF and YfcG are implicated in reduction of nitric oxide – NO sensitivity phenotype identified – YfcF metabolomics with cutting-edge labeling protocol allows measurement of small changes in metabolites • Cellular localization is an important aspect of enzyme function – YghU and YfcG appear to remain in the cytoplasm RuBisCO-like protein, RLP, from R. rubrum (ENO) Canonical methionine salvage pathway (e.g. B. subtilis) Seemingly incomplete MSP (R. rubrum) RLP ? Work with Tobias Erb, Gerlt Lab RLP: evidence for novel fate of methionine sulfur Work with Tobias Erb, Gerlt Lab (ENO) RLP: whole cell untargeted metabolomics Work with Tobias Erb, Gerlt Lab (ENO) RLP: whole cell untargeted metabolomics Data Processing Perturbation Exp. LC-MS Analysis Preprocessing (XCMS) Peak detection/alignment Retention time correction Noise filtering Data Quality Control Peak Grouping Isotope Pattern Analysis Mass check Retention time check Intensity ratio check Peak Grouping Primary Peaks Isotope pattern ≥ 20% intensity change Secondary Peaks Isotope pattern < 20% intensity change Retention time filter Adducts/Salt filter Missing value imputation Deisotoping Data Normalization Monoisotopic peaks Time-wise, condition specific Mean-, Z-value … Formula Prediction Formula modeling Primary peaks used first Round Robin Recursive Backtracking Theoretical Isotope Pattern Modeling First order Markov Forward Trellis Bayesian Statistics Isotope pattern comparison experimental v.s theoretical Heuristics Pathway Activity Profiling DB Search 2ppm mass tolerance Top hits formula Seed Metabolites Isotope pattern High intensity change Exist in current DB Pathway Analysis Seed metabolites info. DB Hits mono. peaks Shared pathways detection Activity Profiling Sort detected peaks upon fold change p-values by MSEA Active Pathways Pathways: p < 0.05 Prior prob. for C, N, S 6 Golden rules Potential Target Peaks Top 3 hits Highly up- or downregulated, but not yet annotated peaks Further experiments are needed Work with Tobias Erb, Gerlt Lab (ENO) RLP: whole cell untargeted metabolomics p-value = 0.02 p-value = 4.8 x 10-4 met-salvage pathway p-value = 7.3 x 10-4 Purine metabolism +MTA Glutathione metabolism MTA 8 4 0 MTR-1P 2 1 0 MTRu-1P 0min 10min 20min 1.0 0.5 0 DXP p-value = 1.2 x 10-3 2 1 0 CDP-MEP 8 4 0 Isoprenoid pathway up-regulated pathway down-regulated pathway pathway showing no big difference metabolite 0min 10min 20min c-MEPP 0min 10min 20min Metabolite intensity ( x 106 ) Control p-value = 0.048 16 8 0 +MTA 0min 10min 20min Butanoate metabolism Metabolite intensity ( x 106 ) Control Work with Tobias Erb, Gerlt Lab (ENO) RuBisCO-like protein from R. rubrum RLP Cupin Work with Tobias Erb, Gerlt Lab (ENO) RuBisCO-like protein (ENO): Results and Conclusions • Perfect starting point for Micro./Metabolomics Core – Collaboration with ENO bridging project – Phenotype was known – High profile project (Ashida, et.al. Science, 2003) • Genome context and measured thiol release suggested novel fate of MTA – Key enzymes in known MSP missing from genome – Cell extracts mixed with MTA produced methanethiol • LC-MS-based metabolomics uncovered connection between MTA feeding and isoprenoid biosynthesis – Untargeted metabolite profiling of R. rubrum uncovered: • Predicted MTA degradation products • Unexpected isoprenoid biosynthesis intermediates Taking advantage of existing samples… Noncovalent Protein: Ligand Interactions Measured by Native ESI-MS (from test cases to EFI samples…) Future work will use the samples stored in the Protein / Structure Core Microbiology/Protein/Structure Core Collaboration Micro./Metabolomics Core: future directions • Application of Biolog and custom phenotype microarrays to null mutants of targets from additional organisms • Transcriptional analysis coupled to growth condition screens to gain complementary evidence for when target genes are expressed • Further improvements in XCMS software to better detect metabolites of low abundance • Application of differential labeling and multiple chromatographies for each metabolomics experiment to increase accuracy • Continued and increasing collaboration with the BPs and Cores