* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download 6-Methoxyadenine Residue Forms a Watson
Homologous recombination wikipedia , lookup
DNA repair protein XRCC4 wikipedia , lookup
DNA sequencing wikipedia , lookup
DNA profiling wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
DNA replication wikipedia , lookup
Microsatellite wikipedia , lookup
United Kingdom National DNA Database wikipedia , lookup
DNA polymerase wikipedia , lookup
DNA nanotechnology wikipedia , lookup
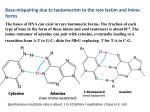
Article No. jmbi.1999.3303 available online at http://www.idealibrary.com on J. Mol. Biol. (1999) 294, 1215±1222 Crystallographic Studies on Damaged DNAs. I. An N 6-Methoxyadenine Residue Forms a Watson-Crick Pair with a Cytosine Residue in a B-DNA Duplex Toshiyuki Chatake1, Akira Ono2, Yoshihito Ueno3, Akira Matsuda3 and Akio TakeÂnaka1* 1 Graduate School of Bioscience and Biotechnology, Tokyo Institute of Technology Nagatsuda, Midori-ku Yokohama, 226-8501, Japan 2 Graduate School of Science Tokyo Metropolitan University Hachioji, Minamiosawa, Tokyo, 192-0364, Japan 3 Graduate School of Pharmaceutical Sciences Hokkaido University, Nishi 6, Kita 12, Kita-ku, Sapporo, 060-0812, Japan Oxyamines such as hydroxylamine and methoxylamine disturb DNA replication and act as potent mutagens, causing nucleotide transition from one purine to another or one pyrimidine to another. In order to investigate mismatch base-pairing in DNA damaged with oxyamines, a dodecamer with the sequence d(CGCGmo6AATCCGCG), where mo6 A is 20 -deoxy-N6-methoxyadenosine, was synthesized and its crystal structure determined. No signi®cant conformational changes are found between the present dodecamer and the original undamaged B-form dodecamer. Electron density maps clearly show that the mo6A residue forms a basepair with a 20 -deoxycytidine residue through hydrogen bonds similar to a Watson-Crick G C base-pair. For these hydrogen bonds to be made, N6-methoxyadenine must chemically take the imino form. The methoxylation thus enables the adenine base to mimic a guanine base. As a result, misincorporation of 20 -deoxycytidine instead of thymidine, or 20 -deoxyadenosine instead of 20 -deoxyguanosine, can occur in DNA replication. # 1999 Academic Press 6 *Corresponding author Keywords: N -methoxyadenine; mutagenesis; X-ray structure; mismatch; damaged DNA Introduction In all organisms, the very high accuracy of DNA replication is achieved by using the Watson-Crick pairs between adenine and thymine residues and between guanine and cytosine residues as an absolute principle. However, oxyamines such as hydroxylamine and methoxylamine disturb this rule and act as potent mutagens, causing nucleotide transitions from one purine to another purine, or from one pyrimidine to another pyrimidine (Singer & Kusmierek, 1982). These chemicals predominantly attack and modify the exocyclic amino groups of nucleic acid bases (Kochetkov & Budowsky, 1969). Substitutions at the N4 amino group of cytosine and at the N6 amino group of adenine have been identi®ed and considered to be Abbreviations used: dATP, 20 -deoxyadenosine 50 triphosphate; TTP, thymidine 50 -triphosphate; dGTP, 20 deoxyguanosine 50 -triphosphate; dCTP, 20 -deoxycytidine 50 -triphosphate; mo6A, 20 -deoxyN6-methoxyadenosine; mo6dATP, 20 -deoxy-N6methoxyadenosine 50 -triphosphate. E-mail address of the corresponding author: [email protected] 0022-2836/99/501215±8 $30.00/0 the origin of the mutation (Budowsky et al., 1975). Matsuda and co-workers (Nishio et al., 1992) found that when adenine residues of a template DNA strand were methoxylated at N6, the 20 -deoxycytidine 50 -triphosphate (dCTP) as well as the complementary thymidine 50 -triphosphate (TTP) were incorporated into the newly synthesized DNA strand at the opposite site to 20 -deoxy-N6-methoxyadenosine (hereafter 20 -deoxy-N6-methoxyadenosine is designated as mo6A). On the other hand, when 20 -deoxyadenosine 50 -triphosphate (dATP) is methoxylated (mo6dATP), mo6dATP is incorporated at the opposite sites to both 20 -deoxycytidine and thymidine residues of a template strand (Singer & Spengler, 1982; Abdul-Maish & Bessman, 1986; Hill et al., 1998). It is interesting to reveal how such a non-Watson-Crick nucleotide is incorporated, despite the fact that the DNA polymerase accepts only Watson-Crick base-pairs (Kiefer et al., 1998). Crystallographic studies have been reported on the structural properties of methoxylated adenine. Two crystal structures of N6-methoxyadenine derivatives, N9-benzyl-N6-methoxyadenine (Fujii et al., 1990) and N6-methoxy-20 ,30 ,50 -tri-O-methyl# 1999 Academic Press 1216 adenosine (Birnbaum et al., 1984), indicate that the N6-methoxylated base prefers the imino form. This fact is consistent with the results from proton nuclear magnetic resonance (Stolarski et al., 1984, 1987), and ultraviolet and infrared spectroscopy studies (Fujii et al., 1987). In the crystals, the methoxy groups take a syn conformation around the N6-C6 bond to the N1 atom, so that the two purine moieties are associated through three hydrogen bonds including a C-H N interaction. From these results, Birnbaum et al. (1984) proposed that mo6A with a syn conformation pairs with a 20 deoxycytidine through two hydrogen bonds, between N1(mo6A) and N3(C) and between the methoxy oxygen atom and N4(C). The geometry of the base-pair, however, will be largely distorted from that of the regular Watson-Crick pairing. Such a distortion might prevent incorporation of the modi®ed adenine base into the active site of DNA polymerase, because the incoming nucleotides are strictly selected for their ability to form a Watson-Crick base pairing (Kiefer et al., 1998). In order to investigate structurally the mismatch base-pairing, X-ray analysis of DNA oligomers containing mo6A should be an ideal approach. Unfortunately, this has not been possible so far: no X-ray structures of DNA oligomers containing N6-methoxyadenine or N6-hydroxyadenine have been reported. Only the crystal structure of a DNA hexamer containing 20 -deoxy-N4-methoxycytidine with the sequence d(CGCGXG) (X is 20 -deoxy-N4-methoxycytidine) was reported (Meervelt et al., 1990). The N4-methoxy group takes the imino form, consistent with solution studies (Brown et al., 1968; Morozov et al., 1982). However, the hexamer adopts a Z-form and the methoxy group is in a syn conformation around the N4-C4 bond, forming a wobble base-pair with the guanine moiety on the opposite strand. Similar structures have also been found in other hexamers containing 20 -deoxy-b-D-ribo furanosyl-(6H,8H-3,4-dihydro-pyrimido[4,5-c][1,2] oxazin-7-one) as X (Moore et al., 1995). These structural features are unacceptable to the DNA polymerase (Kiefer et al., 1998). In addition, only the wobbled base-pairing has been found between cytosine bases and adenine bases in the protonated form (Schuerman et al., 1998; Hunter et al., 1986). To understand the structural basis of mutation mechanism, it is necessary to clarify whether the modi®ed adenine moiety can form a Watson-Crick pair with a cytosine base. Therefore, DNA dodecamer containing 20 -deoxy-N6-methoxyadenosine, with the sequence d(CGCGmo6AATCCGCG) (hereafter Dmo6A C), has been synthesized and its crystal structure determined. This dodecamer was designed so that mo6A interacts with C upon duplex formation, since it is known that the original sequence d(CGCGAATTCGCG) (referred to hereafter as the original dodecamer) prefers to take a B-form conformation (Dickerson & Drew, 1981; Shui et al., 1998). The X-ray structure reported here shows that the methoxy group has no signi®cant effects on the overall DNA conformation. Electron Watson-Crick type N6-Methoxyadenine Cytosine Pair density maps clearly show that the mo6A residue forms a Watson-Crick-like base-pair with the cytosine residue on the opposite strand. This is the ®rst example of such a pairing between N6-methoxyadenine and a cytosine base. We describe below the details of the novel hydrogen-bonding scheme thus revealed in B-form duplex DNA. Results and Discussion Overall structure The crystal structure of Dmo6A C is isomorphous to that of the original dodecamer (Dickerson & Drew, 1981; Shui et al., 1998). The two Dmo6A C strands form a duplex, as expected (Figure 1). The overall structures of the two dodecamers with and without methoxy groups are very similar to each other, their root-mean-square displacement being Ê on superimposition of non-hydrogen atoms. 0.31 A All torsion angles and local helical parameters calculated by the program NUPARM (Bansal et al., 1995) are given in the Supplementary Material (Tables 1 and 2). Figure 2 shows plots of some representative helical parameters along the nucleotide sequence. These parameters ¯uctuate around average values close to those of typical B-form DNA; their variations along the two sequences are almost the same. Thus, the methoxylation of adenosine residues does not affect the DNA conformation signi®cantly. Crystal packing Two duplexes related by a crystallographic 21 symmetry along the c-axis in the crystal form columns in a head-to-tail fashion. The guanine residue at one end of one duplex forms a base-pair with the terminal guanine base at the other end of the symmetry-related duplex, through two Figure 1. A stereo view of the ``damaged'' DNA dodecamer containing 20 -deoxy-N6-methoxyadenosine. The diagram was drawn with the program MOLSCRIPT (Kraulis, 1991). The mo6A residues are shown in red and the cytosine bases paired with mo6A are in green. 1217 Watson-Crick type N6-Methoxyadenine Cytosine Pair Figure 3. The guanine-guanine hydrogen bonds between two symmetry-related duplexes (see the text). The nucleotides are numbered from the 50 end independently in the two strands, a and b. Figure 2. A comparison of the helical parameters of Dmo6A C (circles) with those of the original dodecamer (boxes) (Shui et al., 1998). The top, middle and bottom diagrams, represent the helical rises, the helical twist angles and the displacements of the two duplexes, respectively, plotted along the nucleotide sequence to show their ¯uctuations. These parameters were calculated by the program NUPARM (Bansal et al., 1995). In the right, shaded column, their average values are indicated, together with the typical values for standard A and B-DNA. N2-H N3 hydrogen bonds (see Figure 3). As reported in a previous paper (Chatake et al., 1999b), during data collection, a phase transition of the Dmo6A C crystal with changing humidity at room temperature was found. Water molecules being incorporated into the crystal cause this change. As a result, the contact angle between two neighboring duplexes is changed from 132 to 144 . The present crystal structure has been determined at 110 K. The corresponding angle in the present crystal is 146 , and the unit cell is slightly shrunk, though the variation is within experimental error. An octahedrally hydrated magnesium cation is located in the major groove of one duplex and contacts the ribose-phosphate backbone of another duplex, the position being almost the same as that found in the original dodecamer. The cation links the two duplexes laterally through hydrogen bonds, as shown in Figure 4. Three of the Mg2coordinating water molecules form hydrogen bonds with guanine bases, and the other three are hydrogen bonded to the oxygen atoms of the phosphate groups of another duplex, related by a 21 symmetry along the b-axis (Figure 4). Other water molecules (with the exception of those around the methoxy groups) are also located at similar positions to those of the original dodecamer, and form spines in the grooves of the duplex. Some of the water positions in the minor groove might be partially occupied by sodium cations, as proposed by Shui et al. (1998). The secondary spine in the minor groove rides ``piggyback'' on the primary spine. Tautomerism of mo6A and hydrogenbonding scheme Final 2jFoj ÿ jFcj maps of the individual basepairs are shown in Figure 5. At the ®fth and eighth base-pairs, the densities are well resolved and clearly reveal the positions of the two mo6A residues. The most interesting feature is that both methoxylated adenines pair with cytosine bases in a geometry very similar to the Watson-Crick adenine thymine pair. The average temperature Ê 2 and factors of these two base-pairs (12.6 A Ê 2) are comparable to those of the other base13.4 A pairs, indicating that the two base-pairs, mo6Aa5 Cb8 and mo6Ab5 Ca8 (see the nucleotide numbering in Figure 5) each occupy a single con®guration. Their closest atomic distances and the atomic valence angles are given in Table 1; those of the other hydrogen bonds are given in the Supplementary Material (Table 3). They are in an acceptable range for the N-H N hydrogen bond. To form base-pairs through these hydrogen 1218 Watson-Crick type N6-Methoxyadenine Cytosine Pair mation may be induced to stabilize the base-pairing with cytosine. Biological significance Figure 4. The octahedrally hydrated magnesium cation which links two neighbouring duplexes through hydrogen bonds. Ow indicates water molecules. Ê. Distances are given in A bonds, the chemical structure of the methoxylated adenine base must be the imino form, as shown in Figure 6, since it is known that the cytosine base exists only in the amino form (Pieber et al., 1973). This is consistent with the structures found in N6methoxyadenine derivatives (Birnbaum et al., 1984; Fujii et al., 1990), which take the imino form. Although many types of base-pairs have been reported, only the wobble base-pairing has been found between cytosine and adenine bases in the protonated form (Schuerman et al., 1998; Hunter et al., 1986). Therefore, the present structure is the ®rst example of a Watson-Crick pair between N6methoxyadenine and cytosine. It clearly reveals the structural basis of the mutation mechanism associated with the damaged DNA. The second striking feature is that both methoxy groups adopt an anti conformation to the N1 atom around the C6-N6 bond. This conformation is quite different from those found previously in N6-methoxyadenine derivatives (Birnbaum et al., 1984; Fujii et al., 1990). In the latter crystals, the derivatives are in a syn conformation, forming a hydrogenbonded ribbon involving C-H O and C-H N interactions. Although the tautomers are in the same imino form as here, the conformations of the methoxy groups are different. To change the conformation from syn to anti or vice versa, the adenine moiety must pass through the amino form by tautomerization. Since it is thought that N6-methoxyadenine derivatives are in equilibrium between the amino and the imino forms in solution (Stolarski et al., 1984), the imino form with the anti confor- During replication of DNA, the polymerase accepts only Watson-Crick type base pairs (Kiefer et al., 1998). To form such a pair with a cytosine residue, the adenine moiety must be in the imino form. The unmodi®ed adenine residue never adopts such a tautomer (Wolfenden, 1969). On the other hand, in the structures of N6-methoxyadenine derivatives (Fujii et al., 1990; Birnbaum et al., 1984), including the present dodecamer, the adenine moieties are all in the imino form. The donor/acceptor sites for hydrogen bonds are changed, so that the methoxylated adenine can mimic guanine in hydrogen bond formation. This mimicry makes it possible to form a Watson-Crick type base-pair with a cytosine residue. In the imino form of mo6A, there are two possible conformations for the methoxy group around the C6-N6 bond. Although Birnbaum et al. (1984) proposed a pairing between mo6A in a syn conformation and a cytosine residue, the corresponding geometry is largely distorted from that of regular Watson-Crick pairs. Such a distortion might prevent incorporation of the modi®ed adenine residue into the active site of DNA polymerase (Kiefer et al., 1998). In contrast, mo6A in the present structure is in an anti conformation. When the mo6A C pairs found in the present investigation are superimposed on the canonical A T pairs of the original dodecamer, the base positions are almost the same. This structural isomorphism will almost certainly favor recognition by the polymerase. In addition, the polymerase has an open, solvent accessible space in the major groove of the bound DNA, so that the methoxy groups on the damaged DNA will not interfere with its binding to the polymerase. Through these structural characteristics, an mo6A residue on the template strand makes it possible for dCTP to be incorporated at the opposite site in the newly synthesized DNA strand (Nishio et al., 1992). In the same way, mo6dATP can be incorporated at the site opposite to a cytosine residue on the template strand (Singer & Spengler, 1982; Abdul-Maish & Bessman, 1986; Hill et al., 1998). The equilibrium between the amino and the imino forms of N6-methoxyadenine changes with the polarity of the solution (Stolarski et al., 1984). In addition, when a cytosine derivative is added to a solution containing an mo6A derivative, the imino form is increased, whereas when a uracil derivative is added, the amino form is increased (Stolarski et al., 1987). Therefore, it is expected that an mo6A residue in the amino form will make a Watson-Crick base-pair with a thymine residue. The crystal structure of a DNA dodecamer containing such base-pairs will be discussed in the following paper (Chatake et al., 1999a). Watson-Crick type N6-Methoxyadenine Cytosine Pair 1219 Figure 5. Final re®ned 2jFoj ÿ jFcj electron density maps superimposed on all base-pairs. The maps were contoured at the 1s level by the program O (Jones et al., 1991). The nucleotides are numbered from the 50 end independently in the two strands, a and b. Experimental Procedures Synthesis and crystallization A DNA dodecamer of Dmo6A C was synthesized and puri®ed by the reported method (Nishio et al., 1992). Crystallization was carried out at 4 C by the hanging drop vapor diffusion method in a 4 ml droplet containing equal volumes of 1.5 mM DNA solution in water and reservoir solution. Suitable crystals of Dmo6A C were obtained from 35 % (v/v) 2-methyl-2,4-pentanediol, 18 mM magnesium acetate, 6 mM spermine tetrahydrochloride and 10 mM sodium cacodylate (pH 7.0). 1220 Watson-Crick type N6-Methoxyadenine Cytosine Pair Ê ) and angles (deg.) of mo6A C base-pairs Table 1. Hydrogen bond distances (A mo6Aa5 Cb8 Ca8 mo6Ab5 N1 N3 C2-N1 N3 N1 N3-C2 N3 N1 C2-N3 N1 N3 N1-C2 2.76 123 112 2.80 109 123 N6 N4 C6-N1 N3 N1 N3-C4 N4 N6 C4-N3 N1 N3 N1-C6 Data collection The crystals were ¯ash-frozen in a stream of nitrogen at 110 K. X-ray data were collected at 110 K on the Sakabe-Weissenberg camera (Sakabe, 1991) with synchrotron radiation at the Photon Factory (BL-6B) in Tsukuba. To compensate the blind region, another crystal with a different orientation was used and the X-ray diffraction was collected at 110 K on a Rigaku RAXIS IIc with CuKa radiation generated at 50 kV and 80 mA. Diffraction patterns recorded on imaging plates were processed by the program DENZO (Otwinowski & Minor, 1997) and intensity data were merged by the programs SCALA and AGROVATA in CCP4 (Collaborative Computational Project Number 4, 1994). A total of 8017 independent re¯ections with an Rmerge of 6.0 % were obtained from 32,780 observed re¯ections. The completeÊ resolution ness of the data was 87.1 % in the 100-1.6 A Ê resolution shell. range and 72.1 % for the outer 1.69-1.6 A Ê , b 39.9 A Ê, The cell dimensions are a 25.1 A Ê , the space group being P212121. Statistics of c 65.8 A data collection and crystal data are summarized in Table 2. Structure determination Initial phases were determined by the molecular replacement method using the structure of the original dodecamer (Dickerson & Drew, 1981; Shui et al., 1998) with the program AMoRe (Navaza, 1994). The molecular structure was constructed and modi®ed on a graphic workstation by inspecting jFoj ÿ jFcj omit maps at every nucleotide residue with the programs O (Jones et al., 1991) and QUANTA (distributed by Molecular Simulations, Inc.). The jFoj ÿ jFcj omit maps clearly showed that the mo6A residues form base-pairs with cytosine residues, the methoxy groups taking an anti conformation to the N1 atom around the N6-C6 bond. From the hydrogen-bonding scheme, the mo6A residues were assumed to take the imino form. The stereochemical parameters employed for structural re®nement with the program X-PLOR (BruÈnger, 1992b) were those in the dictionary ®le dna-rna.param. For the present re®nements, this ®le was modi®ed to include the 20 -deoxy-N6-methox- 2.87 112 128 3.10 131 113 122 114 C4-N4 N6 N4 N6-C6 112 120 O6-N6 N4 125 N4 N6-O6 127 yadenosine residue with the imino form. Its structural parameters were derived from the crystal structures of mo6A derivatives (Fujii et al., 1990; Birnbaum et al., 1984). The crystal structure was re®ned through a combination of simulated annealing and positional re®nements, followed by interpretation of omit maps at every nucleotide residue. During the above re®nement, no hydrogen bonding restraints were applied between paired nucleotides. A total of 173 peaks were assigned as water molecules. One magnesium cation coordinated octahedrally by six water molecules was found. The ®nal Ê resolution data R-factor was 19.6 % for 5.0 1.6 A (Rfree 23.1 % for 10 % of the observed data). Statistics of the structure determination are summarized in Table 2. Data Bank accession codes The atomic coordinates have been deposited in the Nucleic Acid Database (NDB) (entry code BD0009). Table 2. Crystal data, data collection and structure determination Space group Ê) Unit cell (A Asymmetric unit (duplexes) Ê) Resolution (A Measured reflections Unique reflections Completeness (%) In the Outershell (%) Rmerge a (%) Structure refinement Resolution range Used reflections Final model Number of non-hydrogen atoms Number of water molecules Number of magnesium atoms Ê 2) Average B-factors (A DNA Water molecules Magnesium atom R-factorb (%) Rfree c (%) r.m.s. deviation Ê) Bond lengths (A Bond angles (deg.) Improper angles (deg.) Ê) Average coordinates errord (A a P212121 a 25.1, b 39.9, c 65.8 4 100 1.6 32,780 8017 87.1 Ê) 72.1(1.69 1.6 A 6.0 5.0 1.6 7676 488 173 1 17.3 37.9 15.8 19.6 23.1 0.009 1.4 1.6 0.20 Rmerge 100 hkljjIhklj ÿ hIhklij/hklhIhkli. R-factor 100 jjFoj ÿ jFcjj/jFoj, where jFoj and jFcj are the observed and calculated structure factor amplitudes, respectively. c Calculated using a random set containing 10 % of observations that were omitted during re®nement (BruÈnger, 1992a). d Estimated from a Luzzati plot (Luzzati, 1952). b Figure 6. Chemical structure of mo6A with possible hydrogen bonds. C6-N6 N4 N6 N4-C4 Watson-Crick type N6-Methoxyadenine Cytosine Pair Acknowledgments We thank N. Sakabe and N. Watanabe for facilities and help during data collection at the Photon Factory (Tsukuba), and T. Simonson for proofreading of the manuscript. This work was supported in part by a grant for the RFTF(97L00503) (Research For The Future) from the Japanese Society for the Promotion of Science, by Grants-in-Aid for Scienti®c Research on Priority Area (Nos. 08124203 and 07250205) from the Ministry of Education, Science, Sport and Culture of Japan, and by the Sakabe project of TARA (Tsukuba Advanced Research Alliance), University of Tsukuba. References Abdul-Masih, M. T. & Bessman, M. J. (1986). Biochemical studies, 6-N-hydroxyaminopurine. J. Biol. Chem. 261, 2020-2026. Bansal, M., Battacharyya, D. & Ravi, B. (1995). NUPARM and NUCGEN: Software for analysis and generation of sequence dependent nucleic acid structures. Comput. Appl. Biosci. 13, 281-287. Birnbaum, G. I., Kierdaszuk, B. & Shugar, D. (1984). Tautomerism and conformation of the promutagenic analogue N6-methoxy-20 ,30 ,50 -tri-O-methyladenosine. Nucl. Acids Res. 12, 2447-2460. Brown, D. M., Hewlins, M. J. E. & Schell, P. (1968). The tautomeric state of N(4)-hydroxy- and of N(4)amino-cytosine derivatives. J. Chem. Soc. sect. C, 1968, 1925-1929. BruÈnger, A. T. (1992a). Free-R value: a novel statistical quantity for assessing the accuracy of crystal structures. Nature, 355, 472-475. BruÈnger, A. T. (1992b). X-PLOR Version 3.1. A System for X-ray Crystallography and NMR, Yale University Press, New Haven, CT. Budowsky, E. I., Sverdlov, E. D., Spasokukotskaya, T. N. & Koudelka, J. (1975). Mechanism of the mutagenic action of hydroxylamine. VIII. Functional properties of the modi®ed adenosine residues. Biochim. Biophys. Acta, 390, 1-13. Chatake, T., Hikima, T., Ono, A., Ueno, Y., Matsuda, A. & TakeÂnaka, A. (1999a). Crystallographic studies on damaged DNAs. II. N6-Methoxyadenine can present two alternate faces for Watson-Crick base-pairing, leading to pyrimidine transition mutagenesis. J. Mol. Biol. 294, 1223-1230. Chatake, T., Ono, A., Ueno, Y., Matsuda, A. & TakeÂnaka, A. (1999b). Crystallization and preliminary X-ray analysis of a DNA dodecamer of d(CGCGmo6AATCCGCG) containing 20 -deoxy-N6methoxyadenosine: Change in crystal packing with different humidity. Acta Crystallog. sect. D, 55, 873876. Collaborative Computational Project Number 4 (1994). The CCP4 suite: programs for protein crystallography. Acta Crystallog. sect. D, 50, 760-763. Dickerson, R. E. & Drew, H. R. (1981). Structure of a BDNA dodecamer II. In¯uence of base sequence on helix structure. J. Mol. Biol. 149, 761-786. Fujii, T., Saito, T., Itaya, T., Kizu, K., Kumazawa, Y. & Nakajima, S. (1987). Purines. XXIX. Syntheses of 9-alkyl-2-deuterio-N6-methoxyadenine and 2deuterio-N6-9-dimethyladenine: Tautomerism in 9-substituted N6-alkoxyadenines. Chem. Pharm. Bull. 35, 4482-4493. 1221 Fujii, T., Saito, T., Date, T. & Nishibata, Y. (1990). Purines. XXXIX. The crystal structure of 9-benzylN6-methoxyadenine. Chem. Pharm. Bull. 38, 912-916. Hill, F., Williams, D. M., Loakes, D. & Brown, D. M. (1998). Comparative mutagenicities of N6-methoxy2,6-diaminopurine and N6-methoxyaminopurine 20 -deoxyribonucleosides and their 50 -triphosphates. Nucl. Acids Res. 26, 1144-1149. Hunter, W. N., Brown, T., Anand, N. N. M. & Kennard, O. (1986). Structure of an adenine:cytosine base pair in DNA and its implications for mismatch repair. Nature, 320, 552-555. Jones, T. A., Zou, J. Y., Cowman, S. W. & Kjeldgaard, M. (1991). Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallog. sect. A, 47, 110-119. Kiefer, J. R., Mao, C., Braman, J. C. & Beese, L. S. (1998). Visualizing DNA replication in a catalytically active Bacillus DNA polymerase crystal. Nature, 391, 304307. Kochetkov, N. K. & Budowksy, E. I. (1969). The chemical modi®cation of nucleic acids. Prog. Nucl. Acid Res. Mol. Biol. 9, 403-438. Kraulis, P. J. (1991). MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallog. 24, 946-950. Luzzati, P. V. (1952). Traitement statistique des erreurs dan la determination des structures cristallines. Acta Crystallog. 5, 802-810. Meervelt, L. V., Moore, M. H., Lin, P. K. T., Brown, D. M. & Kennard, O. (1990). Molecular and crystal structure of d(CGCGmo4CG): N4-methoxycytosine: guanine base-pairs in Z-DNA. J. Mol. Biol. 216, 773781. Moore, M. H., Meervelt, L. V., Salisbury, S. A., Lin, P. K. T. & Brown, D. M. (1995). Direct observation of two base-pairing modes of a cytosine-thymine analogue with guanine in a DNA Z-form duplex: signi®cance for base analogue mutagenesis. J. Mol. Biol. 251, 665-673. Morozov, Y. V., Savin, F. A., Chekhov, V. O., Budowsky, E. I. & Yakovlev, D. Y.-U. (1982). Photochemistry of N6-methoxyadenosine and of N4hydroxycytidine and its methyl derivatives I: Spectroscopic and quantum chemical investigation of ionic and tautomeric forms: syn-anti isomerization. J. Photochem. 20, 229-252. Navaza, J. (1994). AMoRe: an automated package for molecular replacement. Acta Crystallog. sect. A, 50, 157-163. Nishio, H., Ono, A., Matsuda, A. & Ueda, T. (1992). The synthesis and properties of oligodeoxyribonucleotides containing N6-methoxyadenine. Nucl. Acids Res. 20, 777-782. Otwinowski, Z. & Minor, W. (1997). Processing of X-ray diffraction data collected in oscillation mode: Macromolecular crystallography Part A. Methods Enzymol. 276, 307-326. Pieber, M., Kroon, P. A., Prestegard, J. H. & Chan, S. I. (1973). Erratum. Tautomerism of nucleic acid bases. J. Am. Chem. Soc. 95, 3408. Sakabe, N. (1991). X-ray diffraction data collection system for modern protein crystallography with a Weissenberg camera and an imaging plate using synchrotron radiation. Nucl. Instrum. Methods Phys. Res. ser. A, 303, 448-463. Schuerman, G. S., Meervelt, L. V., Loakes, D., Brown, D., Lin, P. K. T., Moore, M. H. & Salisbury, S. A. 1222 (1998). A thymine-like base analogue forms wobble pairs with adenine in a Z-DNA duplex. J. Mol. Biol. 282, 1005-1011. Shui, X., McFail-Ison, L., Hu, G. G. & Williams, L. D. (1998). The B-DNA dodecamer at high resolution reveals a spine of water on sodium. Biochemistry, 37, 8341-8355. Singer, B. & Kusmierek, J. T. (1982). Chemical mutagenesis. Annu. Rev. Biochem. 52, 655-693. Singer, B. & Spengler, S. (1982). Reaction of O-methylhydroxylamine with adenosine shifts tautomeric equilibrium to cause transitions. FEBS Letters, 139, 69-71. Stolarski, R., Kierdaszuk, B., Hagberg, C. E. & Shugar, D. (1984). Hydroxyamine and methoxyamine mutagenesis: Displacement of the tautomeric equilibrium of the promutagen N6-methoxyadenosine by complementary base pairing. Biochemistry, 23, 2906-2913. Stolarksi, R., Kierdaszuk, B., Hagberg, C. E. & Shugar, D. (1987). Mechanism of hydroxyamine mutagenesis: tautomeric shifts and proton exchange between the promutagen N6-methoxyadenosine and cytidine. Biochemistry, 26, 4332-4337. Watson-Crick type N6-Methoxyadenine Cytosine Pair Wolfenden, R. V. (1969). Tautomeric equilibria in inosine and adenosine. J. Mol. Biol. 40, 307-310. Edited by I. Tinoco (Received 19 April 1999; received in revised form 6 October 1999; accepted 12 October 1999) http://www.academicpress.com/jmb Supplementary Material comprising three Tables is available from JMB Online