* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Evolution of the Insulin Receptor Family and
Survey
Document related concepts
Ligand binding assay wikipedia , lookup
Biosynthesis wikipedia , lookup
Proteolysis wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Genetic code wikipedia , lookup
Biochemical cascade wikipedia , lookup
Metalloprotein wikipedia , lookup
Biochemistry wikipedia , lookup
Lipid signaling wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Endocannabinoid system wikipedia , lookup
NMDA receptor wikipedia , lookup
Clinical neurochemistry wikipedia , lookup
Paracrine signalling wikipedia , lookup
Transcript
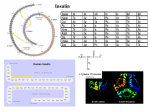
Evolution of the Insulin Receptor Family and Receptor Isoform Expression in Vertebrates Catalina Hernández-Sánchez,* Alicia Mansilla,* Flora de Pablo,* and Rafael Zardoyaà *3D Lab (Development, Differentiation & Degeneration), Department of Cellular and Molecular Physiopathology, Centro de Investigaciones Biológicas, Consejo Superior de Investigaciones Cientı́ficas (CSIC), Ramiro de Maeztu 9, Madrid, Spain; Centro de Investigación Biomédica en Red de Diabetes y Enfermedades Metabólicas (CIBERDEM), Ramiro de Maeztu, 9, Madrid, Spain; and àDepartmento de Biodiversidad y Biologı́a Evolutiva, Museo Nacional de Ciencias Naturales, CSIC, José Gutiérrez Abascal, 2, Madrid, Spain The molecular phylogeny of the vertebrate insulin receptor (IR) family was reconstructed under maximum likelihood (ML) to establish homologous relationships among its members. A sister group relationship between the orphan insulin–related receptor (IRR) and the insulin-like growth factor 1 receptor (IGF1R) to the exclusion of the IR obtained maximal bootstrap support. Although both IR and IGF1R were identified in all vertebrates, IRR could not be found in any teleost fish. The ancestral character states at each position of the receptor molecule were inferred for IR, IRR þ IGF1R, and all 3 paralogous groups based on the recovered phylogeny using ML in order to determine those residues that could be important for the specific function of IR. For 18 residues, ancestral character state of IR was significantly distinct (probability .0.95) with respect to the corresponding inferred ancestral character states both of IRR þ IGF1R and of all 3 vertebrate paralogs. Most of these IR distinct (shared derived) residues were located on the extracellular portion of the receptor (because this portion is larger and the rate of generation of IR shared derived sites is uniform along the receptor), suggesting that functional diversification during the evolutionary history of the family was largely generated modifying ligand affinity rather than signal transduction at the tyrosine kinase domain. In addition, 2 residues at positions 436 and 1095 of the human IR sequence were identified as radical cluster-specific sites in IRR þ IGF1R. Both Ir and Irr have an extra exon (namely exon 11) with respect to Igf1r. We used the molecular phylogeny to infer the evolution of this additional exon. The Irr exon 11 can be traced back to amphibians, whereas we show that presence and alternative splicing of Ir exon 11 seems to be restricted exclusively to mammals. The highly divergent sequence of both exons and the reconstructed phylogeny of the vertebrate IR family strongly indicate that both exons were acquired independently by each paralog. Introduction Insulin and insulin-like growth factors (IGFs) constitute a fundamental family of hormone polypeptides common to all metazoans. These hormones control essential functions including cell growth, metabolism, reproduction, and longevity (Kimura et al. 1997; Efstratiadis 1998; Tissenbaum and Ruvkun 1998; Brogiolo et al. 2001; Nakae et al. 2001; Saltiel and Kahn 2001; Holzenberger et al. 2003; Nef et al. 2003). Dysfunction of these factors in humans is associated to several pathological disorders such as diabetes, dwarfism, and cancer. In invertebrates, insulin and IGF have a general function as mitogenic growth factors (Chan and Steiner 2000). In postnatal vertebrates, the cell proliferation function has been restricted to IGF1 and IGF2, whereas insulin has become a metabolic regulatory hormone mainly controlling homeostasis of different metabolites (most prominently glucose) (Chan and Steiner 2000). However, during embryonic development, insulin action and regulation in vertebrates appear to be reminiscent of those found in invertebrates (Hernandez-Sanchez et al. 2006). Physiological functions of the insulin and IGF polypeptides require specific surface cell receptors, and subtle differences in the structure and function of the receptors can account for important variations in the biological activity of the hormones across metazoans. Although in invertebrates there are several insulin-like peptides, only 1 insulin receptor (IR) protein has been described (Fernandez et al. 1995; Pashmforoush et al. 1996; Kimura et al. 1997; Ruvkun and Hobert 1998). However, in Key words: insulin receptor, alternative splicing, ancestral character states. E-mail: [email protected] Mol. Biol. Evol. 25(6):1043–1053. 2008 doi:10.1093/molbev/msn036 Advance Access publication February 29, 2008 Ó The Author 2008. Published by Oxford University Press on behalf of the Society for Molecular Biology and Evolution. All rights reserved. For permissions, please e-mail: [email protected] vertebrates, 3 distinct receptors that can bind with highaffinity insulin and the IGF were described based on differences in primary structure and function: the IR (Ebina et al. 1985; Ullrich et al. 1985), the type 1 IGF receptor (IGF1R) (Ullrich et al. 1986), and the type 2 IGF receptor (IGF2R) (Morgan et al. 1987). Of these, the IGF2R is in fact the mannose-6-phosphate receptor that, only in mammals, has acquired a binding domain for IGF2, and it is not a signaling receptor (Morgan et al. 1987). In addition, an orphan receptor (with an unknown ligand) termed the insulin receptor-related receptor (IRR) was also described as a member of the IR family based on sequence similarity (Shier and Watt 1989). The IR, IGF1R, and IRR present a rather conserved protein structure (see known domains in fig. 1) (Ullrich et al. 1986; De Meyts 2004) and belong to the larger tyrosine kinase receptor superfamily (Hubbard and Till 2000). Unlike other members of this superfamily, the above-mentioned 3 receptors form dimeric (a2/b2) structures in the cell membrane, which can be either homodimers, composed by 2 identical a/b momomers, or heterodimers formed by 2 different a/b monomers (e.g., IRab/IGF1Rab) (Moxham et al. 1989; Soos and Siddle 1989; Schlessinger 2000; Fernandez et al. 2001). Ligand binding to IR and IGF1R triggers a conformational change that enables autophosphorylation of the receptor cytoplasmic tyrosine residues and initiates a cascade of intracellular signaling events that engender diverse biological responses (metabolism, cell proliferation, cell differentiation, survival, and growth), depending on the cell type and the developmental and functional stage. IR and IGF1R have different but overlapping physiological functions (reviewed in Nakae et al. [2001]). The evolutionary and molecular mechanism through which the functional specialization of each receptor was achieved remains an open question. Thus far, the exact mechanism through which the orphan IRR is activated and its function are unknown. 1044 Hernández-Sánchez et al. The genes encoding IR, IGF1R, and IRR share similar genomic organization. Both the a and b chains are synthesized from a unique mRNA, which is comprised by 22 exons in IR and IRR and by 21 exons in IGF1R (Rosenfeld and Roberts 1999). In both, Ir and Irr, the extra exon with respect to Igf1r is exon 11. Strikingly, exon 11 is constitutive in Irr whereas each of the human and murine Ir exon 11 is alternatively spliced, which results in 2 protein isoforms (IRA and IRB) that differ by the absence or presence of 12 amino acids at the C-terminus of the a subunit, respectively (Ebina et al. 1985; Ullrich et al. 1985; Seino and Bell 1989; Seino et al. 1989). Both IR isoforms display differences in ligand affinity binding, kinase activity, receptor internalization, and recycling as well as intracellular signaling capacity and tissue distribution (Mosthaf et al. 1990; McClain 1991; Vogt et al. 1991; Yamaguchi et al. 1991; Kellerer et al. 1992; Leibiger et al. 2001). In the present study, we reconstructed the molecular phylogeny of the vertebrate IR family in order to establish homologous relationships among its members. We also identified evolutionarily conserved and functionally divergent amino acid residues in the 3 vertebrate receptors, as well as shared derived residues of IR in order to gain insights on the evolutionary mechanisms underlying the functional diversification of the family and to identify those residues that may be responsible for the specific function of IR. In addition, we traced the presence of the alternatively spliced Ir exon 11 in the recovered phylogeny in order to characterize the evolution of this extra exon, and found that it is a novel acquisition of mammals. Materials and Methods Animals Fertilized White Leghorn (Gallus gallus) eggs (Granja Rodrı́guez-Serrano, Salamanca, Spain) were incubated at 38.4 °C and 60–90% relative humidity for the time periods indicated, and the embryos were staged according to (Hamburger and Hamilton 1951). The 10-day posthatching chickens (P) were from Avı́cola Grau (Madrid, Spain). Frogs (Xenopus laevis) were kindly supplied by Dr MJ Delgado (Universidad Complutense de Madrid). The 10-day and 35-day postnatal mice (C57BL/6) (Mus musculus) were from Centro de Investigaciones Biológicas stabularium. All animals were handled according to European Union Guidelines for animal research. RNA Isolation and Reverse Transcriptase–Polymerase Chain Reaction Total RNA from tissues was isolated using Trizol reagent (Invitrogen, Carlsbad, CA). The reverse transcriptase Fig. 1.—Diagram of the a2/b2 quaternaty structure of the IR showing the protein domain organization. L1 and L2, large domains 1 and 2 (leucine-rich repeats); CR, Furin-like cysteine-rich domain; FnIII-1, FnIII-2, FnIII-3, fibronectin type III domains; ID, insert domain in FnIII-2; TM, transmembrane domain; JM, juxtamembrane domain; TK, tyrosine kinase domain; and CT, carboxy-terminal tail. Disulphure bonds are shown. Arrowheads on the left side of the diagram indicate IR shared derived amino acids, whereas lines on the right side of the diagram indicate amino acids conserved in all 3 vertebrate members of the IR family. Evolution of the Insulin Receptor Family in Vertebrates 1045 reaction was typically performed with 5 lg RNA, the Superscript III Kit, and oligo-dT primer (all from Invitrogen), followed by amplification with the Expand High fidelity Polymerase (Roche Diagnostics, Mannheim, Germany). The mouse Ir was amplified using the sense primer 5#GGCCAGTGAGTGCTGCTCATGC-3# (mP1) and the antisense primer 5#-TGTGGTGGCTGTCACATTCC-3# (mP2). The chicken Ir was amplified using the sense primer 5#-CAGAAGGAGCTGGAGGAGTC-3# (cP1) and the antisense primer 5#-TCTGCTCCTCTGCACTCTC-3# (cP2) for the first polymerase chain reaction (PCR) and cP1 sense primer and the antisense (cP4) 5#-GGAGCCCAGGTCTCTTCTCT-3# for the nested PCR. The Xenopus Ir was amplified using the sense primer 5#-ACCTTCATCCAAGTGCTGTC-3# (xP1) and the antisense primer 5#-CAGAGTTCCATTGGCTACTC-3# (xP2) for the first PCR and the sense 5#-GCCTTCCAGAACTTGGACTC-3# (xP3) and the antisense 5#-TGGCTCTGTTTCATCCGGAG-3# (xP4) for the nested PCR. Sequences Molecular databases at the National Center for Biotechnology Information (http://www.ncbi.nlm.nih.gov) were screened for vertebrate IRs using the Blast search program (Altschul et al. 1997) and the human IR (NP_000199.2) as query. In addition, some additional vertebrate IRs were directly retrieved through searches in ENSEBML (http://www.ensembl.org/index.html). Phylogenetic Analysis A total of 39 complete or almost complete IR family proteins were included in the phylogenetic analyses. Sequences were aligned using MUSCLE version 3.633 (Edgar 2004). Multiple alignments were subsequently refined by eye and compared with IR alignments at the Receptors for Insulin and Insulin-like molecules (RILM) database (http://www.biochem.ucl.ac.uk/RILM; Garza-Garcia et al. 2007). Ambiguous alignments in highly variable (gap-rich) regions were excluded from phylogenetic analyses (aligned sequences and the exclusion sets are available from the authors upon request). We used PROTTEST version 1.2.6 (Abascal et al. 2005) to select the substitution model that best fit the empirical data set (JTT þ I þ C; a 5 0.86, I 5 0.09) and PHYML version 2.4.4 (Guindon and Gascuel 2003) to find the maximum likelihood (ML) tree. The robustness of the inferred tree was assessed using bootstrapping (500 pseudoreplicates) as implemented in PHYML. The recovered ML tree was used as framework to identify those protein residues that are shared derived by IR orthologs. In a first round of analysis, each of the protein residues was mapped onto the phylogeny, and the ancestral character states and shared derived amino acid residues were inferred with parsimony using PAUP* version 4.0b10 (Swofford 2002) and MacClade v4.0 (Maddison WP and Maddison DR 1992) and taking into account only those shared derived characters that had a consistency index (CI) (as a measure of the fit of each character to the tree) above 0.5. In a second round of analysis, the likelihood of the an- cestral character state of the identified potential shared derived amino acids of IR was estimated using BayesTraits version 1.0 (Pagel et al. 2004). Finally, the posterior probability for functional cluster-specific (type II; Gu 2006) residues was estimated for IR versus IRR þ IGF1R using Diverge 2.0 (Gu 2006). No attempt of testing for positive selection events based on branch-site likelihood ratio tests (Yang and Nielsen 2002) was made because of the relatively high nucleotide sequence divergences found among vertebrate IRs and the existence of saturation of silent mutations at third codon positions (data not shown). Results Phylogeny of Vertebrate IRs The phylogeny of the vertebrate IR family was reconstructed in order to establish homologous relationships among its members. Phylogenetic analyses were based on an amino acid sequence data alignment including 1,596 positions, of which 335 highly variable positions were excluded due to ambiguity in positional homology assignment. A total of 250 positions were invariant, and 813 characters were parsimony informative. The alignment was used to map sequence divergence along the receptor molecule. Although invariant sites were found throughout the IR sequence (fig. 1), they were particularly abundant around positions 90–110 and 1000–1250 (human IR sequence; fig. 2). In contrast, positions around 650–810, 890–990, and at the carboxy end (above position 1310) were relatively variable and showed few invariant positions. The ML analysis of the molecular data set using the IRs of 2 basal chordates, Ciona intestinalis and Branchiostoma lanceolatum, as outgroup sequences recovered the phylogenetic tree (logL 5 32422.12) shown in figure 3. According to the reconstructed tree, vertebrate IRs could be grouped with maximal bootstrap support into 3 distinct paralogous groups, which correspond to IR, IRR, and IGF1R, respectively. A sister group relationship between IRR and IGF1R to the exclusion of IR obtained maximal bootstrap support. As expected, the phylogeny of vertebrates (teleost fish, (amphibians, (birds, mammals))) was recovered within each paralog. However, although both IR and IGF1R were found in all vertebrates, IRR could not be identified in any teleost fish. Mean (±standard deviation [SD]) amino acid sequence divergences between both outgroups and the 3 vertebrate paralogs, IR, IRR, and IGF1R, were 0.54 ± 0.06, 0.57 ± 0.04, and 0.54 ± 0.05, respectively. Mean (±SD) amino acid sequence divergences within IR, IRR, and IGF1R were 0.15 ± 0.01, 0.21 ± 0.01, and 0.20 ± 0.01, respectively. Mean (±SD) amino acid sequence divergences across placentals for IR, IRR, and IGF1R were 0.07 ± 0.05, 0.12 ± 0.03, and 0.04 ± 0.03, respectively. Ancestral Character State Reconstruction, Shared Derived Characters of IR, and Cluster-Specific Functionally Divergent Sites Ancestral character state reconstruction analyses were used 1) to identify those residues that being shared derived of IR could be responsible for its specific function as well as 1046 Hernández-Sánchez et al. Fig. 2.—Alignment of selected members of the vertebrate IR family. Amino acid positions correspond to the human IR (NP_000199.2). Black boxes represent IR shared derived amino acids, whereas gray boxes indicate amino acids that are conserved in all 3 members of the vertebrate IR family. Main domains of the receptor are shown (see also IR alignments at the RILM database: http://www.biochem.ucl.ac.uk/RILM; Garza-Garcia et al. 2007). 2) to determine their distribution along the receptor molecule. In a preliminary filtering analysis, a total of 41 amino acid residues were identified as potential shared derived characters of IR under the parsimony criterion (CI .0.50) (data not shown). The putative character states of the 41 residues in the common ancestor of IR, IRR þ IGF1R, and all 3 vertebrate paralogs were estimated under the likelihood criterion, respectively (table 1). For 18 Evolution of the Insulin Receptor Family in Vertebrates 1047 Fig. 3.—Phylogenetic analysis of the vertebrate IR family. The ML phylogram is shown. Numbers in nodes represent bootstrap values above 70%. GenBank or Ensembl accession numbers are in brackets. *Ornithorhynchus and Ciona sequences were manually assembled from data available in Ensembl. 1048 Hernández-Sánchez et al. Table 1 Ancestral Character State Reconstruction under the Likelihood Criterion AA position Ir 137a M (0.99) Igf1r þ Irr T K R G Ir þ Igf1r þ Irr (0.64) (0.16) (0.14) (0.06) 192a G (1) E F R S D K (0.36) (0.15) (0.15) (0.15) (0.11) (0.04) 266 E (0.97) P Q R K W T A Q P R W E K (0.33) (0.30) (0.16) (0.07) (0.04) (0.04) (0.04) (0.33) (0.26) (0.15) (0.11) (0.06) (0.05) F L R K L A 283 (0.94) (0.03) (0.39) (0.33) (0.20) (0.08) L F K R (0.39) (0.33) (0.16) (0.07) 284a S (1) 287a (0.51) (0.49) (0.46) (0.27) (0.25) E (0.37) D (0.35) A (0.28) Q Y S G A A (0.53) E (0.34) D (0.10) S (0.63) G (0.23) A (0.1) T (0.97) E (0.90) K (0.04) S (0.02) AA position Ir 735a Y (1) 806 S (0.99) 839 Y (1) 841a S (0.86) T (0.11) 883 L (1) 886a V (1) Igf1r þ Irr F (1) R (0.99) F (0.99) F (0.72) Y (0.27) H (0.99) I (0.92) L (0.08) Ir þ Igf1r þ Irr F(1) S (0.89) F (0.05) R (0.04) Y (0.99) Y (0.86) F (0.12) L (0.86) H (0.14) I (0.97) a 304a Y (1) 364a N (1) 436 R (1) F (0.99) K (0.59) E (0.27) D (0.14) Q (0.99) F (1) K (0.51) E (0.36) D (0.14) C G P A 911 (0.61) (0.28) (0.10) (0.96) C (0.60) G (0.27) P (0.09) H Y F N V R A Q 456a (0.49) (0.51) (0.21) (0.17) (0.16) (0.15) (0.15) (0.15) R (0.99) A V F N Q R (0.29) (0.18) (0.17) (0.15) (0.11) (0.1) 940 T (0.99) 941 Y (0.97) 956 K (0.96) V M I L A (0.27) (0.26) (0.24) (0.19) (0.04) S H F V D (0.28) (0.22) (0.16) (0.15) (0.14) L I M T V (0.43) (0.22) (0.14) (0.11) (0.10) Y H D T V A F (0.58) (0.14) (0.07) (0.06) (0.04) (0.04) (0.03) T M L A V F I K F A L I M T V (0.18) (0.18) (0.16) (0.14) (0.12) (0.11) (0.11) (0.26) (0.23) (0.11) (0.10) (0.09) (0.08) (0.07) (0.06) IR-derived state. residues, the inferred ancestral character state of the IR paralog was significantly distinct (probability .0.95) with respect to the corresponding inferred ancestral character states both of IRR þ IGF1R and of all 3 vertebrate paralogs (table 1). These IR distinct (shared derived) residues were distributed rather evenly along the IR molecule (figs. 1 and 2). Moreover, 14 out of the 18 residues mapped on the extracellular portion of the receptor, whereas the remaining 4 residues were located on the intracellular region (figs. 1 and 2). Given the relative proportions of the extracellular (70%) and intracellular (29%) portions of the receptor, an expected random distribution of the shared derived residues would have been 12 and 6, respectively. The difference between the observed and expected frequencies was not statistically significant according to a chi-square test (P . 0.05). The physicochemical nature of the amino acid change leading to the ancestral IR character state for each of the detected IR shared derived positions was characterized. In 2 instances (positions 137 and 192 of the human IR sequence), the change from the ancestral vertebrate IR character state to the ancestral IR character state implied a replacement of a polar residue by a nonpolar one, whereas in another 2 cases (304 and 735), the change was in the opposite direction. In 3 instances (364, 520, and 1128), a polar residue was substituted by another polar residue, whereas in another 3 cases (628, 640, and 886), a nonpolar residue was replaced by another nonpolar residue. In another 8 cases (284, 287, 456, 531, 841, 995, 1026, and 1141), it was not possible to determine the physicochemical nature of the change (table 1). The inferred shared derived positions of IR do not need to be necessarily conserved in the other paralogous groups of the family. According to the posterior probabilities estimated in the ancestral character state reconstruction analyses, in 10 (192, 284, 287, 364, 456, 520, 531, 628, 1026, and 1128) out of the 18 positions presenting an unambiguous shared derived amino acid state of IR, the inferred amino acid residue was conserved in the ancestor of IR but was variable in the ancestors of the other 2 subsets (IRR þ IGF1R and IR þ IRR þ IGF1R) of homologous genes (table 1). In the remaining 8 (137, 304, 640, 735, 841, 886, 995, and 1141) positions, the ancestors of the other 2 subsets of homologous genes (IRR þ IGF1R and IR þ IRR þ IGF1R) also showed relatively unambiguous character states (table 1). Only 1 (640) out of these 8 positions showed a distinct conserved amino acid in each of the 3 subsets, whereas in the other sites, IRR þ IGF1R and IR þ IRR þ IGF1R shared the same ancestral character state. Changes in the evolutionary conservation at a particular residue may reflect functional divergence after gene Evolution of the Insulin Receptor Family in Vertebrates 1049 Table 1 Extended AA position Ir 520a W (1) 529 G (1) 531a M (1) 566 R (0.97) 620 T (1) 638 I (1) 640a L (1) 698 N (0.80) 721 K (0.98) 731 T (0.90) Igf1r þ Irr Q R H Y (0.46) (0.27) (0.19) (0.08) S (0.99) T I V R (0.51) (0.25) (0.15) (0.10) N (0.91) S (0.09) A (0.97) L V M A L (1) V (0.98) D P S T E (0.28) (0.26) (0.21) (0.13) (0.09) A M R E (0.44) (0.23) (0.19) (0.10) N S V I R (0.76) (0.08) (0.06) (0.04) (0.03) Ir þ Igf1r þ Irr R (0.51) Q (0.40) Y (0.06) G (0.69) S (0.31) T (0.83) V (0.16) R (0.96) T (0.95) A (0.54) M (0.36) V (0.06) I (0.93) L (0.07) P (0.98) M K A E (0.65) (0.17) (0.09) (0.07) T S R I (0.58) (0.21) (0.11) (0.08) 995a P (0.51) S (0.49) 1026a L (1) 1128a R (0.98) 1135 E (0.98) Q (0.41) S (0.20) N (0.15) H (0.07) T (0.07) P (0.04) E (0.04) 1141a A (0.50) V (0.49) AA position Ir 960 G (1.00) 983 Q (0.81) I (0.15) 986 G (0.88) S (0.12) Igf1r þ Irr L (0.86) M (0.06) G (0.04) E (0.33) N (0.33) S (0.33) F (0.66) Y (0.26) S (0.08) P Q R M V R 990 (0.41) (0.39) (0.19) (0.57) (0.39) (0.03) V (1) C S N H M (0.25) (0.23) (0.21) (0.15) (0.13) Q E I L (0.53) (0.27) (0.12) (0.05) R (0.92) E (0.08) G (0.63) S (0.37) Y G S M (0.63) (0.20) (0.10) (0.07) Ir þ Igf1r þ Irr G (0.82) M (0.15) Q (0.55) I (0.38) E (0.04) G (0.54) S (0.45) R Q M P (0.47) (0.16) (0.15) (0.13) V (1) M N S C (0.40) (0.31) (0.18) (0.08) E (0.82) Q (0.12) I (0.04) E (0.76) R (0.21) D (0.03) S (0.87) G (0.13) T Y G S M (0.33) (0.31) (0.19) (0.09) (0.07) duplication. Using a statistical approach that compares amino acid changes between IR and IRR þ IGF1R, we found that most sites of the IR molecule were predicted to be unrelated with cluster-specific (type II) functional divergence. Only 2 residues (positions 436 and 1095 of the human IR sequence) received the highest posterior ratio score (2.54) and were identified as radical cluster-specific sites (posterior probability of 0.72). However, neither position 436 (R in IR and Q in IRR þ IGF1R) nor 1095 (K in IR and Q in IRR þ IGF1R) correspond to any of the sites identified as shared derived of IR (table 1). 628a P (0.96) (0.37) (0.36) (0.17) (0.10) 1362 T (0.99) only isoform IRB and in brain only isoform IRA were detected (Fig. 4B). Interestingly, when the equivalent approach was taken for chicken and Xenopus tissues, only a single isoform was detected (fig. 4C and D) after 35 cycles of PCR amplification. Sequencing analysis of the amplified PCR bands showed that they corresponded to the isoform IRA. To discard the possibility that the isoform IRB could be expressed at so low levels that could not be detected in a single round of amplification, we performed a nested PCR. As shown in figure 4E and F, only a single amplification product was again obtained strongly, suggesting that only 1 isoform is expressed in the analyzed chicken and Xenopus tissues. Evolution of the Splicing of Exon 11 The human and murine Ir are alternatively spliced in a tissue-specific manner and produce 2 isoforms, IRA and IRB. In order to trace the origin of this splicing mechanism, we analyzed the presence of 2 isoform transcripts in nonmammalian species by reverse transcriptase–polymerase chain reaction (RT-PCR) analysis using the upstream primer directed against the 3# end of exon 10 and the downstream primer against the 5# end of the putative exon 12 (fig. 4A). In agreement with previous reports, we found differential distribution of both Ir isoform transcripts in mouse tissues in the 2 ages analyzed. Adipose tissue and muscle expressed both isoform transcripts to different degree, whereas in liver Discussion In this study, we provide for the first time a robust phylogenetic framework to understand the molecular evolution and functional diversification of IRs in vertebrates. According to the reconstructed phylogeny, the 3 described vertebrates IR, IRR, and IGF1R conform each a monophyletic group and correspond to 3 distinct paralogous groups. A first duplication of the receptor gene led to the Ir paralog and the ancestor of Igf1r and Irr, which were both subsequently generated in a second round of duplication. The presence of 3 or more paralogs in vertebrates but only 1 gene copy in nonvertebrates is a common pattern to other 1050 Hernández-Sánchez et al. Fig. 4.—IR isoform expression. (A) Schematic representation of the mouse IR gene. White boxes indicate noncoding regions, whereas gray boxes represent the coding exons, which are numbered. Solid lines represent constitutive splicing, and dashed lines represent alternative RNA processing. Primers (P) used in PCR are indicated. IR RT-PCR with P1 and P2 primers of RNA from different tissues of postnatal day 10 (P10) and 35 (P35) mouse (B) embryonic day 19 (E19) and postnatal day 10 (P10) chicken (C) and adult Xenopus (D). Nested PCR using P1 and P4 primers of P10 and P35 chicken (E) and P3 and P4 of adult Xenopus (F) tissues. A, adipose tissue; M, skeletal muscle; L, liver; and B, brain. (G) IR and IRR exon 11 alignments. Amino acids encoded together by exon 11 and its flanking exons are shown in red. protein families (e.g., hedgehog; Zardoya et al. 1996) and could be the result of independent duplication events in each protein family. However, there is increasing evidence that 2 rounds of whole-genome duplications occurred early in vertebrate evolution and could be responsible for having generated the observed higher paralog number of vertebrate protein families (Meyer and Schartl 1999; Dehal and Boore 2005), in general, and of the IR family, in particular. Another genome duplication has been proposed in teleost fishes (Meyer and Van de Peer 2005; Brunet et al. 2006), and, for example, Tetraodon nigroviridis presents indeed 2 gene copies of Ir (CAG08022.1 and CAG07190.1) and of Igf1r (CAG13078.1 and CAG03114.1). Strikingly, Irr was missing in teleost fish. This may reflect that Irr orthologs in teleosts might have highly divergent sequences and thus might have not been detected yet through similarity searches. Alternatively, it may be possible either that Irr was lost at least in the common ancestor of teleosts or that Irr was a novel acquisition of tetrapods. Sequencing the genomes of basal actinopterygian (e.g., bichir and sturgeon) and sarcopterygian (e.g., lungfishes and coelacanth) fishes would help in discerning among these competing hypotheses. As previously reported for mammals (Ullrich et al. 1986; Shier and Watt 1989; Rosenfeld and Roberts 1999), primary structure was relatively highly conserved among paralogs of the IR family across vertebrates (22% invariant sites). Conserved sites are distributed throughout the molecule, and conserved stretches are particularly abundant around the tyrosine kinase domain. In fact, some sites of this domain are also conserved in less related members of the tyrosine kinase superfamily (Hubbard and Till 2000; Ward et al. 2007) such as the epidermal growth factor receptor and the plate-derived growth factor receptor (data not shown). The overall evolutionary conservation of the primary structure of the vertebrate IR indicates that a relatively high proportion of the molecule is under strong selection pressure because it is likely needed to maintain the general IR and signal transduction functions. In agreement with this observation, only few sites (2%) were identified as being shared derived by IR. These positions are maintained by purifying selection and not need to be fully conserved in all IR orthologs. They characterize IR and may be particularly important for its related but distinct function. Our results show that the relative distribution of IR shared derived sites was even along the receptor (i.e., the rate of generation of shared derived sites along the molecule was uniform). However, because the extracellular portion of the receptor more than doubles the intracellular portion, most IR shared derived sites were located in the extracellular portion, Evolution of the Insulin Receptor Family in Vertebrates 1051 which may suggest that subtle differences in function among paralogs of the family are evolutionarily achieved more through changes in ligand-binding affinity (Schaefer et al. 1990; Brandt et al. 2001) than by modifications of the intracellular signal transduction at the tyrosine kinase domain. Shared derived residues are best candidates for sitedirected mutagenesis in order to identify which evolutionary changes are responsible for functional divergence of IR (Jimenez-Jimenez et al. 2006). One of the identified shared derived IR characters (position 364) corresponds to a potential site for N-glicosilation. This posttranslational modification is critical for the correct assembling of tertiary and quaternary IR structures (Olson et al. 1988). Two other potential N-glicosilation sites (positions 105 and 651) are conserved among all paralogs across vertebrates. However, individual mutation of the 18 N-glicosilation sites of the human IR showed high functional redundancy of those sites (reviewed in Adams et al. [2000]). Other important residues in maintaining the quaternary structure of the receptor are 6 cysteines involved in the formation of disulphure bonds. These cysteines are highly conserved in all paralogs across vertebrates and in Branchiostoma. However, 4 out of these 6 cysteines are not conserved in Ciona. A similar pattern is found in Drosophila (Fernandez et al. 1995) where the receptor assembles into a quatenary structure but only 2 out of the 6 cysteines described in human IR are conserved. Amino acid residues that are highly conserved in IR not need to be necessarily conserved in the other paralogs and reflect site-specific shift of evolutionary rate (Gu 2006). However, in 2 positions (436 and 1095), cluster-specific residues (different between paralogs IR and IRR þ IGF1R, but otherwise highly conserved within each homologous group) were identified with statistical support. These residues evidenced radical shifts of amino acid property (type II functional divergence) (Gu 2006). According to the phylogeny, IR retains the ancestral character state in these 2 positions, whereas IRR þ IGF1R feature a shared derived character state. Again, these sites should be straightforward targets for site-directed mutagenesis, in this case, to characterize IRR þ IGF1R functional divergence. Both Ir and Irr have an extra exon (namely exon 11) with respect to Igf1r. The Irr exon 11 can be traced back to amphibians, whereas Ir exon 11 is found exclusively in mammals (fig. 4). The highly divergent sequence of both exons (fig. 4) and the reconstructed phylogeny of the vertebrate IR family strongly indicate that both exons were acquired independently by each paralog rather than being present in the ancestor of the vertebrate protein family and lost multiple times. Although most functional diversification of the vertebrate IR family is achieved through gene duplication, alternative splicing is also found to be an important mechanism that has generated functional diversification during the evolutionary history of the family. In this study, we show that alternative splicing of Ir exon 11 seems to be restricted exclusively to mammals. The physiological outcome of the evolutionary acquisition of Ir exon 11 is not fully understood to date. The novel IR isoform (IRB) shows a decreased affinity for IGF2 resulting in a more specific receptor for insulin and restricting IGF2 signaling. In this regard, 2 additional evolutionary novelties appeared in mam- mals to fine-tune IGF2 activity. First, a novel receptor for IGF2, namely IGF2R, was evolved. This receptor is devoid of signal transduction capabilities, and it acts as a clearance receptor that modulates levels of circulating IGF2. Second, imprinting of the Igf2 gene was evolved to prevent expression of the maternal allele. Alterations of the strict control of IGF2 bioavailability as well as IRA expression have been associated to malignant process (reviewed in Denley et al. [2003]). A second selective advantage derived from the evolutionary acquisition of Ir exon 11 could be specialization of IRB as a more metabolic receptor. This isoform is predominantly expressed in insulin target tissues (Seino and Bell 1989; Mosthaf et al. 1990) that are responsible of glucose homeostasis. Furthermore, patients with myotonic dystrophy type 1 present a 70% decrease in insulin sensitivity in skeletal muscle that is associated with a switch in alternative splicing from IRB to IRA (Savkur et al. 2001). The insulin signaling pathway is most complex in vertebrates with both insulin and IGF having acquired important and diversified metabolic functions beyond their original growth-stimulating function in nonvertebrates. Our study shows that an important element to understand the functional diversification of these hormones is the corresponding functional diversification of the vertebrate IRs with respect to their nonvertebrate counterparts. Specificity of the ligands of the three vertebrate IR paralogs seems to have been acquired mostly through gene duplication of the gene products, as well as through a mechanism of alternative splicing in mammals. Functional divergence among vertebrate IR paralogs is centered in few amino acid residues along the molecule and future site-directed mutagenesis essays on these residues will be key in disentangling the complex evolution of new functions within the protein family and, in particular, in deciphering the unknown function of the orphan IRR. Acknowledgments We thank Ms C. Murillo for her excellent technical support. We thank Dr MJ Delgado (Universidad Complutense de Madrid) for providing the adult Xenopus and Dr R. Martı́nez-Álvarez for performing preliminary experiments. We thank Dr J. Rozas and 2 anonymous reviewers for insightful comments on an earlier version of the manuscript. The studies were financed partially by the grants BFU2004–2352 and BFU2007-61055 from the Spanish Ministry of Education and Science (MEC) and the ‘‘Red de Grupos’’ RGDM G03/212 from the ‘‘Instituto de Salud Carlos III’’ from MSC (Spain) to F.d.P. and the grant CGL2004-00401 from MEC to R.Z.; C.H.S. was a holder of a ‘‘Ramón y Cajal’’ contract and A.M. had a predoctoral fellowship, both from MEC (Spain). Literature Cited Abascal F, Zardoya R, Posada D. 2005. ProtTest: selection of best-fit models of protein evolution. Bioinformatics. 21:2104–2105. Adams TE, Epa VC, Garrett TP, Ward CW. 2000. Structure and function of the type 1 insulin-like growth factor receptor. Cell Mol Life Sci. 57:1050–1093. 1052 Hernández-Sánchez et al. Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ. 1997. Gapped BLAST and PSIBLAST: a new generation of protein database search programs. Nucleic Acids Res. 25:3389–3402. Brandt J, Andersen AS, Kristensen C. 2001. Dimeric fragment of the insulin receptor alpha-subunit binds insulin with full holoreceptor affinity. J Biol Chem. 276:12378–12384. Brogiolo W, Stocker H, Ikeya T, Rintelen F, Fernandez R, Hafen E. 2001. An evolutionarily conserved function of the Drosophila insulin receptor and insulin-like peptides in growth control. Curr Biol. 11:213–221. Brunet FG, Crollius HR, Paris M, Aury JM, Gibert P, Jaillon O, Laudet V, Robinson-Rechavi M. 2006. Gene loss and evolutionary rates following whole-genome duplication in teleost fishes. Mol Biol Evol. 23:1808–1816. Chan SJ, Steiner DF. 2000. Insulin through the ages: phylogeny of a growth promoting and metabolic regulatory hormone. Am Zool. 40:213–222. De Meyts P. 2004. Insulin and its receptor: structure, function and evolution. Bioessays. 26:1351–1362. Dehal P, Boore JL. 2005. Two rounds of whole genome duplication in the ancestral vertebrate. PLoS Biol. 3:e314. Denley A, Wallace JC, Cosgrove LJ, Forbes BE. 2003. The insulin receptor isoform exon 11 (IR-A) in cancer and other diseases: a review. Horm Metab Res. 35:778–785. Ebina Y, Ellis L, Jarnagin K, et al. (12 co-authors). 1985. The human insulin receptor cDNA: the structural basis for hormoneactivated transmembrane signalling. Cell. 40:747–758. Edgar RC. 2004. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32:1792–1797. Efstratiadis A. 1998. Genetics of mouse growth. Int J Dev Biol. 42:955–976. Fernandez AM, Kim JK, Yakar S, Dupont J, HernandezSanchez C, Castle AL, Filmore J, Shulman GI, Le Roith D. 2001. Functional inactivation of the IGF-I and insulin receptors in skeletal muscle causes type 2 diabetes. Genes Dev. 15:1926–1934. Fernandez R, Tabarini D, Azpiazu N, Frasch M, Schlessinger J. 1995. The Drosophila insulin receptor homolog: a gene essential for embryonic development encodes two receptor isoforms with different signaling potential. EMBO J. 14:3373–3384. Garza-Garcia A, Patel DS, Gems D, Driscoll PC. 2007. RILM: a web-based resource to aid comparative and functional analysis of the insulin and IGF-1 receptor family. Hum Mutat. 28:660–668. Gu X. 2006. A simple statistical method for estimating type-II (cluster-specific) functional divergence of protein sequences. Mol Biol Evol. 23:1937–1945. Guindon S, Gascuel O. 2003. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol. 52:696–704. Hamburger V, Hamilton HL. 1951. A series of normal stages in the development of the chick embryo. J Morphol. 88:49–92. Hernandez-Sanchez C, Mansilla A, de la Rosa EJ, de Pablo F. 2006. Proinsulin in development: new roles for an ancient prohormone. Diabetologia. 49:1142–1150. Holzenberger M, Dupont J, Ducos B, Leneuve P, Geloen A, Even PC, Cervera P, Le Bouc Y. 2003. IGF-1 receptor regulates lifespan and resistance to oxidative stress in mice. Nature. 421:182–187. Hubbard SR, Till JH. 2000. Protein tyrosine kinase structure and function. Annu Rev Biochem. 69:373–398. Jimenez-Jimenez J, Zardoya R, Ledesma A, Garcia de Lacoba M, Zaragoza P, Mar Gonzalez-Barroso M, Rial E. 2006. Evolutionarily distinct residues in the uncoupling protein UCP1 are essential for its characteristic basal proton conductance. J Mol Biol. 359:1010–1022. Kellerer M, Lammers R, Ermel B, Tippmer S, Vogt B, Obermaier-Kusser B, Ullrich A, Haring HU. 1992. Distinct alpha-subunit structures of human insulin receptor A and B variants determine differences in tyrosine kinase activities. Biochemistry. 31:4588–4596. Kimura KD, Tissenbaum HA, Liu Y, Ruvkun G. 1997. daf-2, an insulin receptor-like gene that regulates longevity and diapause in Caenorhabditis elegans. Science. 277:942–946. Leibiger B, Leibiger IB, Moede T, Kemper S, Kulkarni RN, Kahn CR, de Vargas LM, Berggren PO. 2001. Selective insulin signaling through A and B insulin receptors regulates transcription of insulin and glucokinase genes in pancreatic beta cells. Mol Cell. 7:559–570. Maddison WP, Maddison DR. 1992. MacClade: analysis of phylogeny and character evolution. Sunderland (MA): Sinauer Associates Inc. McClain DA. 1991. Different ligand affinities of the two human insulin receptor splice variants are reflected in parallel changes in sensitivity for insulin action. Mol Endocrinol. 5:734–739. Meyer A, Schartl M. 1999. Gene and genome duplications in vertebrates: the one-to-four (-to-eight in fish) rule and the evolution of novel gene functions. Curr Opin Cell Biol. 11:699–704. Meyer A, Van de Peer Y. 2005. From 2R to 3R: evidence for a fish-specific genome duplication (FSGD). Bioessays. 27:937–945. Morgan DO, Edman JC, Standring DN, Fried VA, Smith MC, Roth RA, Rutter WJ. 1987. Insulin-like growth factor II receptor as a multifunctional binding protein. Nature. 329:301–307. Mosthaf L, Grako K, Dull TJ, Coussens L, Ullrich A, McClain DA. 1990. Functionally distinct insulin receptors generated by tissue-specific alternative splicing. EMBO J. 9:2409–2413. Moxham CP, Duronio V, Jacobs S. 1989. Insulin-like growth factor I receptor beta-subunit heterogeneity. Evidence for hybrid tetramers composed of insulin-like growth factor I and insulin receptor heterodimers. J Biol Chem. 264:13238–13244. Nakae J, Kido Y, Accili D. 2001. Distinct and overlapping functions of insulin and IGF-I receptors. Endocr Rev. 22:818–835. Nef S, Verma-Kurvari S, Merenmies J, Vassalli JD, Efstratiadis A, Accili D, Parada LF. 2003. Testis determination requires insulin receptor family function in mice. Nature. 426:291–295. Olson TS, Bamberger MJ, Lane MD. 1988. Post-translational changes in tertiary and quaternary structure of the insulin proreceptor. Correlation with acquisition of function. J Biol Chem. 263:7342–7351. Pagel M, Meade A, Barker D. 2004. Bayesian estimation of ancestral character states on phylogenies. Syst Biol. 53:673–684. Pashmforoush M, Chan SJ, Steiner DF. 1996. Structure and expression of the insulin-like peptide receptor from amphioxus. Mol Endocrinol. 10:857–866. Rosenfeld RG, Roberts CT. 1999. The IGF system: molecular biology, physiology and clinical applications. Totowa (NJ): Humana Press. Ruvkun G, Hobert O. 1998. The taxonomy of developmental control in Caenorhabditis elegans. Science. 282:2033–2041. Saltiel AR, Kahn CR. 2001. Insulin signalling and the regulation of glucose and lipid metabolism. Nature. 414:799–806. Savkur RS, Philips AV, Cooper TA. 2001. Aberrant regulation of insulin receptor alternative splicing is associated with insulin resistance in myotonic dystrophy. Nat Genet. 29:40–47. Schaefer EM, Siddle K, Ellis L. 1990. Deletion analysis of the human insulin receptor ectodomain reveals independently Evolution of the Insulin Receptor Family in Vertebrates 1053 folded soluble subdomains and insulin binding by a monomeric alpha-subunit. J Biol Chem. 265:13248–13253. Schlessinger J. 2000. Cell signaling by receptor tyrosine kinases. Cell. 103:211–225. Seino S, Bell GI. 1989. Alternative splicing of human insulin receptor messenger RNA. Biochem Biophys Res Commun. 159:312–316. Seino S, Seino M, Nishi S, Bell GI. 1989. Structure of the human insulin receptor gene and characterization of its promoter. Proc Natl Acad Sci USA. 86:114–118. Shier P, Watt VM. 1989. Primary structure of a putative receptor for a ligand of the insulin family. J Biol Chem. 264:14605–14608. Soos MA, Siddle K. 1989. Immunological relationships between receptors for insulin and insulin-like growth factor I. Evidence for structural heterogeneity of insulin-like growth factor I receptors involving hybrids with insulin receptors. Biochem J. 263:553–563. Swofford DL. 2002. PAUP*: phylogenetic analysis using parsimony (*and other methods). Version 4.0b 10. Sunderland (MA): Sinauer Associates, Inc. Tissenbaum HA, Ruvkun G. 1998. An insulin-like signaling pathway affects both longevity and reproduction in Caenorhabditis elegans. Genetics. 148:703–717. Ullrich A, Bell JR, Chen EY, et al. (15 co-authors). 1985. Human insulin receptor and its relationship to the tyrosine kinase family of oncogenes. Nature. 313:756–761. Ullrich A, Gray A, Tam AW, et al. (14 co-authors). 1986. Insulinlike growth factor I receptor primary structure: comparison with insulin receptor suggests structural determinants that define functional specificity. EMBO J. 5:2503–2512. Vogt B, Carrascosa JM, Ermel B, Ullrich A, Haring HU. 1991. The two isotypes of the human insulin receptor (HIR-A and HIR-B) follow different internalization kinetics. Biochem Biophys Res Commun. 177:1013–1018. Ward CW, Lawrence MC, Streltsov VA, Adams TE, McKern NM. 2007. The insulin and EGF receptor structures: new insights into ligand-induced receptor activation. Trends Biochem Sci. 32:129–137. Yamaguchi Y, Flier JS, Yokota A, Benecke H, Backer JM, Moller DE. 1991. Functional properties of two naturally occurring isoforms of the human insulin receptor in Chinese hamster ovary cells. Endocrinology. 129:2058–2066. Yang Z, Nielsen R. 2002. Codon-substitution models for detecting molecular adaptation at individual sites along specific lineages. Mol Biol Evol. 19:908–917. Zardoya R, Abouheif E, Meyer A. 1996. Evolutionary analyses of hedgehog and Hoxd-10 genes in fish species closely related to the zebrafish. Proc Natl Acad Sci USA. 93:13036–13041. Norihiro Okada, Associate Editor Accepted January 29, 2008