* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download WORKING WITH THE FIGURES
Viral phylodynamics wikipedia , lookup
Group selection wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Gene expression programming wikipedia , lookup
Pharmacogenomics wikipedia , lookup
Fetal origins hypothesis wikipedia , lookup
Polymorphism (biology) wikipedia , lookup
Genetic testing wikipedia , lookup
History of genetic engineering wikipedia , lookup
Koinophilia wikipedia , lookup
Genetic engineering wikipedia , lookup
Behavioural genetics wikipedia , lookup
Genetic drift wikipedia , lookup
Public health genomics wikipedia , lookup
Designer baby wikipedia , lookup
Genome (book) wikipedia , lookup
Population genetics wikipedia , lookup
Human genetic variation wikipedia , lookup
Microevolution wikipedia , lookup
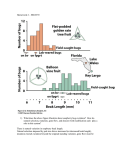
19 The Inheritance of Complex Traits WORKING WITH THE FIGURES 1. Figure 19-9 shows the trait distributions before and after a cycle of artificial selection. Does the variance of the trait appear to have changed as a result of selection? Explain. Answer: The phenotypic variance of the trait does not appear to have changed as a result of selection. Neither the range nor the shape of the phenotypic distribution have changed, so the variance would not have changed because it is a function of these two parameters. Although it might be expected that phenotypic variation would decrease after selection, there are reasons why it might not. First, selection would act on additive genetic effects, which in this case are calculated to account for 50 percent of the phenotypic variance. Phenotypic variance due to dominance and environmental effects would be unchanged and could help to sustain phenotypic variation even after a reduction in additive genetic variation. Second, and probably more important, most populations have a large store of unexpressed additive genetic variation, as shown by the long term responses to most artificial selection efforts. One generation of selection would likely favor the expression of some of this untapped variation, extending the range of phenotypic variation beyond that present in the original population. Together these two factors could lead to a shift in the population mean without a corresponding reduction in phenotypic variation. 2. Figure 19-11 shows the expected distributions for the three genotypic classes if the B locus is a QTL affecting the trait value. a. As drawn, what is the dominance/additive D/A ratio? b. How would you redraw this figure if the B locus had no effect on the trait value? c. How would the positions along the x-axis of the curves for the different genotypic classes of the B locus change if AID= 1.0? 452 Chapter Nineteen Answer: a. As drawn, the Bb distribution is intermediate between bb and BB. This indicates additive gene action and the D/A ratio will be 0. b. If the B locus had no effect on the trait value, all three distributions would coincide and there would be three completely overlapping distributions that looked like the one depicted for bb. low intermediate high c. If the AID ratio= 1.0, this would mean total dominance so that the BB and Bb phenotypes would be the same. There would be two distributions- a small one for bb and a large one for BB and Bb combined. low 3. intermediate high Figure 19-16 shows the results of a QTL fine-mapping experiment. Which gene would be implicated as controlling fruit weight if the mean fruit weight for each line was as follows? Line 1 2 3 4 5 6 7 8 9 10 Fruit wei 181.4 169.3 170.7 171.2 171.4 182.2 180.6 180.7 181.8 169.3 Answer: Lines 1, 6, 7, 8, and 9 all have high fruit weight and Beefmaster chromosomal segments that include the ald2 gene. Lines 2, 3, 4, 5 and 10 all Chapter Nineteen 453 have low fruit weight and Sungold chromosomal segments that include ald2. ald 2 is the only gene that is Beefmaster in high fruit weight lines and Sungold in low fruit weight lines. This indicates that ald2 controls the difference in fruit weight. 4. Figure 19-17 shows a set of haplotypes. Suppose these are haplotypes for a chromosomal segment from 18 haploid yeast strains. On the right edge of the figure, the S and D indicate whether the strain survives (S) or dies (D) at high temperature (40°C). Using the x2 test (see Chapter 3) and Table 3-1, does either SNP 1 or SNP6 show evidence for an association with the growth phenotype? Explain. Answer: These samples may be too small for a meaningful chi-square test, because with samples of this size the differences between the expected and observed outcomes would have to be relatively very large to reach significance. However, a chi-square can be calculated. Overall, there were 10 strains that survived high temperatures. If SNP 1 has no effect, these survivors should be distributed randomly between the A and G variants. Out of 18 strains, 12 were A strains and 6 were G strains, so 2/3 of the survivors, or 6.67 should be A and 113 or 3.33 should be G. This constitutes the expected outcome or null hypothesis. The observed outcome is that 7 survivors were A strain and 3 were G. Using the chi-square formula L (O-E) 2/E, the chi-square test for this data set is 0.049 with 1df, P > 0.05. The null hypothesis cannot be rejected. This data set falls well within the range of normal variation for a sample this size. For SNP 6 half the strains were A and half were G, so the expected number of survivors out of 10 would be 5 each. The observed number of survivors was G = 8 and A = 2. Using the same chi-square formula, a chi-square of 3.6 is obtained. The critical value for a chi-square with 1df is 3.81, so this value is close to significance but also falls within the range of normal variation. Even though G appears to be strongly associated with survival at high temperatures, the null hypothesis cannot be rejected because the sample size is too small to resolve a difference of this magnitude, even if it is "real." A larger sample would be necessary. 5. Figure 19-18a shows a plot of P values (represented by the dots) along the chromosomes of the dog genome. Each P value is the result of a statistical test of association between an SNP and body size. Other than the cluster of small P values near IGFJ, do you see any chromosomal regions with evidence for a significant association between an SNP and body size? Explain. Answer: The only interval with values that exceeds the threshold for significance is that near IGF1, so it is the only region with evidence of significant association with body size. Other regions seem to exceed the usual significance level of 0.05 but are not considered significant in this case. These are not considered significant because in any analysis where many comparisons 454 Chapter Nineteen are made the threshold for significance must be raised to account for the increased chance of a "significant" outcome that was actually due to random chance and would not be repeatable. 6. Figure 19-19 shows plots of P values (represented by the dots) along the chromosomes of the human genome. Each P value is the result of a statistical test of association between an SNP and a disease condition. There is a cluster, or spike, of statistically significant P values (green dots) at the gene HLA-DRBJ for two diseases. Why might this particular gene contribute to susceptibility for the autoimmune diseases rheumatoid arthritis and type 1 diabetes? Answer: HLA is one of the major histocompatibility complex (MHC) genes that regulates immune response. Rheumatoid arthritis and type 1 diabetes are both autoimmune diseases. Mutations in immune response genes are thus associated with diseases that result from abnormal immune responses. BASIC PROBLEMS 7. Distinguish between continuous and discontinuous variation in a population, and give some examples of each. Answer: There are many traits that vary more or less continuously over a wide range. For example, height, weight, shape, color, reproductive rate, metabolic activity, etc., vary quantitatively rather than qualitatively. Continuous variation can often be represented by a bell-shaped curve, where the "average" phenotype is more common than the extremes. Discontinuous variation describes the easily classifiable, discrete phenotypes of simple Mendelian genetics: seed shape, auxotrophic mutants, sickle-cell anemia, etc. These traits show a simple relationship between genotype and phenotype. 8. The table below shows a distribution of bristle number in a Drosophila population. Calculate the mean, variance, and standard deviation for these data. Bristle number 1 2 3 4 5 6 7 Number of individuals 1 4 7 31 56 17 4 Chapter Nineteen 455 Answer: The mean (or average) is calculated by dividing the sum of all measurements by the total number of measurements, or in this case, the total number of bristles divided by the number of individuals. mean= .X= [1 + 4(2) + 7(3) + 31(4) + 56(5) + 17(6) + 4(7)] (1 + 4 + 7 + 31 +56+ 17 +4) = 564;120 = 4.7 average number ofbristlelindividual The variance is useful for studying the distribution of measurements around the mean and is defined in this example as variance= i =average of the (actual bristle count- mean) 2 s2 = IfN L (xi - x)Z = 1/120~]~1- 4.7) 2 + (2- 4.7) 2 + (3- 4.7) 2 + (4- 4.7/ + (5 - 4.7) + (6 - 4.7) + (7- 4.7) 2] =0.26 2 The standard deviation, another measurement of the distribution, is simply calculated as the square root of the variance. standard deviation= s = .J'fj'26= 0.51 9. Suppose that the mean IQ in the United States is roughly 100 and the standard deviation is 15 points. People with IQs of 145 or higher are considered "geniuses" on some scales of measurement. What percentage of the population is expected to have an IQ of 145 or higher? In a country with 300 million people, how many geniuses are there expected to be? Answer: Fifteen IQ points is equal to three standard deviations. Values that fall within ± three standard deviations comprise 99.7 percent of a normally distributed population. In this example that would include all people with IQs of 145 or higher and all those with IQs of 55 and lower. Half of those, or 0.15 percent, will have IQs or 145 or higher. In a country with 300 million people, this would equal 450,000 people. 10. A bean breeder is working with a population in which the mean number of pods per plant is 50 and the variance is 10 pods 2 • The broad-sense heritability is known to be 0.8. Given this information, can the breeder be assured that the population will respond to selection for an increase in the number of pods per plant in the next generation? Answer: Although a response to selection in the next generation is likely, the breeder cannot be assured of it. The response will depend on the type of genetic 456 Chapter Nineteen variation present. Broad-sense heritability consists of both additive and dominant genetic effects. If all genetic variation were due to additive effects, a strong response to selection would occur because the phenotype of selected individuals would correlate strongly with their genotypes and would be transmissable to the next generation. If there were no additive genetic variance (all genetic effects were due to dominant gene action) selected phenotypes would not be correlated with specific genotypes and would not be heritable. Variation due to dominant gene effects is not transmissible to the next generation. The actual response to selection would depend on the relative contributions of additive and dominant genetic effects, the D/A ratio. CHALLENGING PROBLEMS 11. In a large herd of cattle, three different characters showing continuous distribution are measured, and the variances in the following table are calculated: Variance Phenotypic Environmental Additive genetic Dominance genetic Shank length 310.2 248.1 46.5 15.6 Characters Neck length 730.4 292.2 73.0 365.2 Fat content 106.0 53.0 42.4 10.6 a. Calculate the broad- and narrow-sense heritabilities for each character. b. In the population of animals studied, which character would respond best to selection? Why? c. A project is undertaken to decrease mean fat content in the herd. The mean fat content is currently 10.5 percent. Animals with a mean of 6.5 percent fat content are interbred as parents of the next generation. What mean fat content can be expected in the descendants of these animals? Answer: a. Broad heritability measures that portion of the total variance that is due to genetic variance. The equation to use is H 2 = the genetic variance/phenotypic variance where genetic variance = phenotypic variance - environmental variance Narrow heritability measures that portion of the total variance that is due to the additive genetic variation. The equation to use is