* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download The Graph of Life
Pathogenomics wikipedia , lookup
Gene therapy wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Maximum parsimony (phylogenetics) wikipedia , lookup
Gene desert wikipedia , lookup
Genome (book) wikipedia , lookup
Genome evolution wikipedia , lookup
Helitron (biology) wikipedia , lookup
Gene nomenclature wikipedia , lookup
Quantitative comparative linguistics wikipedia , lookup
Gene expression profiling wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Designer baby wikipedia , lookup
Gene expression programming wikipedia , lookup
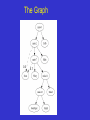
The Graph of Life Dennis Shasha Joint work with Kenneth Birnbaum Treester system by: Matt Olim Phylogenetic Reconstruction (careful: root is bottom-most) Strictly non cyclic •maximum parsimony •maximum likelihood Character Conflict (one feature/two places) Why Conflict? Helianthus petiolaris Helianthus annuus X sunflowers ancient hybridization event ~100,000 years ago Helianthus paradoxus Phylogenetic … Trees? H. paradoxus Adapted from Rieseberg et al. 1991 Phylogenomics •takes many individual gene trees (technically, orthologs) •combines data -- e.g., sequence concatenation •obtains a single tree from combined data -hopefully with high confidence! • Rokas et al. (Nature 2003): 20 trees is enough (based on 8 yeast species). Observation: Individual Trees Vary Example: multiple gene trees of eight sequenced yeast species From Rokas et al. 2003 For this data set Conflict Smoothed by Combining Data 100% Most parsimonious tree from 106 individual trees concatenated 100% 100% 100% 100% S. cerevisiae S. paradoxus S. mikatae S. kudriavzevii S. bayanus S. castellii S. kluyveri C. albicans But then why do the trees vary? •Noise (maybe trees for a given gene aren’t right) •Hybridization (different species have viable offspring) •Horizontal gene transfer, e.g. bacterial orgy •Convergent evolution (think cactus, only at genomic level) Whatever reason: settling on a consensus tree may throw away much information. Finding Hidden Signals •Begin with several trees for each orthologous gene. •Not only the most parsimonious ones but the top few. •1. Find trees in descending order of popularity and see whether genes have interesting commonalities. •2. Generate a network from popular trees. Data SetUp •Using PAUP, generated top 10 most parsimonius trees for each orthologous gene that was present in all eight species. • Popularity contest: loop find most popular tree spit out tree and associated genes remove genes having that tree end loop • Unused Popularity contest: same but don’t remove. Findings •Genes following the Rokas consensus tree are normal in every way. 378 of those. (We expanded analysis) (((((((Scer,Spar),Smik),Skud),Sbay),Scas),Sklu),Calb) [count378] • 46 genes associated with next tree somewhat closer to one another than expected, but not quite at the 5% threshold. (((((((Scer,Spar),Smik),Skud),Sbay),Sklu),Scas),Calb) [count46] •13 genes associated with next tree are well within the 5% threshold of being close to one another and on only 5 of 17 chromosomes. (((((Skud,Sbay),((Scer,Spar),Smik)),Scas),Sklu),Calb) [count13] Other Odd Properties of the 13 • 4 of the 13 genes are annotated as having ATPase or ATP synthase (only 92 out 6,000 genes have similar annotations). • Consensus tree is quite different from 13 gene tree. (((((((Scer,Spar),Smik),Skud),Sbay),Scas),Sklu),Calb) (((((Skud,Sbay),((Scer,Spar),Smik)),Scas),Sklu),Calb) The odd 13 S. cerevisiae S. paradoxus S. mikatae S. kudriavzevii S. bayanus consensus tree S. castellii S. kluyveri C. albicans S. cerevisiae S. paradoxus S. mikatae Remnants of an ancient hybridization event? S. kudriavzevii S. bayanus Parallel gene evolution among ATP-related genes and others ? S. castellii S. kluyveri C. albicans The Graph 0.3 0.7 Network Building •LatTrans: Addario-Berry. Models lateral transfer of genetic information. Makes some assumptions about mutation rates. Always between siblings. •Model horizontal gene transfer: Lake and Rivera. Procaryotic model that tries to distill fundamental tree of life assuming Markov model. Vs. Doolittle Network Building 2 •Moret, Nakleh et al propose "galled networks" which are networks where hybridizations don't intersect. They argue that this limitation makes sense, because there are modest levels of recombination. •Our approach: start with reliable gene trees and build a “conservative” species graph. No statistical assumptions except quality of tree branching. Nomenclature Gene Tree A (gene A and orthologs) A11 A122 A121 A2 extant gene variants A12 missing ancestral gene variants A1 A1 is parent of A12 A Assumptions variant = one of the orthologs of a gene 1. A variant is likely to arise only once in the tree or network (convergent phenotype yes; but not same sequence). 2. If species X has one parent P, then for each gene A, the variant of A in X must be the direct descendent of the variant in P or equal to that variant (e.g., A12-->A121) 3. If species X has more than one parent, then for each gene A, the variant of A must descend/be equal to the variant in exactly one parent. First: characterize species by tree position sp1 sp2 sp1: A1 sp2: A21 sp3: A22 sp1:B22 sp2:B21 sp3:B11 sp3 sp1(A1 B22) sp2(A21 B21) sp3(A22 B11) sp1 In a tree for sp1 and sp2, those species must both be descendants of node N B2 where B2 arises. Further, the split between A1 and A2 must B descend from N. sp3 sp2 Let’s Illustrate Dependencies Species: sp1(A1 B22), sp2(A21 B21), sp3(A22 B11) From sp1 and sp2, B2 arises before A1 splits from A2. Birth(B2) before birth(A2) From sp2 and sp3, A2 arises before B1 splits from B2. Birth(A2) before birth(B2). Shows that tree is not possible. We choose a tree that is consistent with as many species as possible and then add the remaining species using as few edges as possible. Weights indicate number of species. One possible graph sp1 sp2 sp3 sp1: A1 sp2: A21 sp3: A22 sp1:B22 sp2:B21 sp3:B11 m1 (A B) m2 (A B1) sp1(A1 B22) m3 (A2 B2) sp2(A21 B21) sp3 sp2 sp1 m2 stranded taxa sp3(A22 B11) sp3 sp2 sp1 m3 B2 B “base tree” m1 Does it make sense? •The three trees seem quite different: (((((((Scer,Spar),Smik),Skud),Sbay),Scas),Sklu),Calb) (((((((Scer,Spar),Smik),Skud),Sbay),Sklu),Scas),Calb) (((((Skud,Sbay),((Scer,Spar),Smik)),Scas),Sklu),Calb) In particular, Skud seems to move a lot. But our graph showed multiple ancestry for Scas only. Well, maybe •Observe that Scer, Spar, Smik always form same subtree, so let’s replace by a single node xxx. Then remove Scas because we are interested only in whether what remains forms a tree. Here is what we get: ((((((xxx),Skud),Sbay)),Sklu),Calb) ((((((xxx),Skud),Sbay),Sklu)),Calb) (((((Skud,Sbay),(xxx))),Sklu),Calb) Graphing Phylogenies •infer ancestral states for each gene tree •find tree that includes a maximal set of species (“conservative” base tree) •infer parents of remaining species from ancestors (graph) Summary Phylogenomics generates large datasets to overcome conflicting signals in phylogenetic trees, but may cause us to ignore biological signals. •Phylogenetic Networks -- A Graph of Life -- can suggest gene transfers through hybridization or some other reason. • Basic method: start with gene trees, take reliable bifurcations (over 60%) and combine them into a consensus directed graph that suggests possible paths of gene transfer. •Software is general purpose and available. Cause of Conflict? Examined 3 best supported network edges (most gene tree support) Traces of an anicent hybridization •Possible expectation: blocks of genes with common ancestry •none of the genes contributing to network edge are clustered on the chromosome IN C. cerevisiae but 13 oddballs are something of an exception •…total number of genes is 108, so synteny impossible. Will extend this in future. Convergent Evolution •Do genes comprising network edges have a common function? No but still looking. Major Cyclic Edges 12 gene trees S. cerevisiae S. paradoxus S. mikatae S. kudriavzevii 15 gene trees 24 gene trees S. bayanus S. castellii S. kluyveri C. albicans Revised from Rokas et al. 2003