* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Supplementary Materials Metabolic Flux Determination in Perfused
Basal metabolic rate wikipedia , lookup
Adenosine triphosphate wikipedia , lookup
Pharmacometabolomics wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Metalloprotein wikipedia , lookup
Microbial metabolism wikipedia , lookup
Photosynthesis wikipedia , lookup
NADH:ubiquinone oxidoreductase (H+-translocating) wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Isotopic labeling wikipedia , lookup
Biochemistry wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
Multi-state modeling of biomolecules wikipedia , lookup
Biosynthesis wikipedia , lookup
Nicotinamide adenine dinucleotide wikipedia , lookup
Citric acid cycle wikipedia , lookup
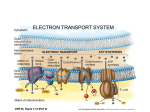
Supplementary Materials Metabolic Flux Determination in Perfused Livers by Mass Balance Analysis: Effect of Fasting Mehmet A. Orman1 Kosuke Arai3 Martin L. Yarmush2,3 Ioannis P. Androulakis1,2 Francois Berthiaume2,* Marianthi G. Ierapetritou1,* 1: Department of Chemical and Biochemical Engineering, Rutgers, The State University of New Jersey, Piscataway, NJ 08854, USA 2: Department of Biomedical Engineering, Rutgers, The State University of New Jersey, Piscataway, NJ 08854, USA 3: Center for Engineering in Medicine/Surgical Services, Massachusetts General Hospital, Harvard Medical School, and the Shriners Hospitals for Children, Boston, MA 02114, USA *: Corresponding authors. Emails: [email protected], [email protected] 1 A. Stoichiometry of Hepatic Metabolic Network The liver metabolic network involving all possible major liver-specific pathways such as gluconeogenesis, glycolysis, urea cycle, fatty acid metabolism, pentose phosphate pathway, TCA cycle, glycogen metabolism and amino acid metabolism is given in Table SI. For more detailed explanations about the enzymes and the number of steps in each reaction, see references (Banta et al. 2007; Chan et al. 2003; Lee et al. 2000; Lee et al. 2003). B. Mathematical Analysis Solution Space of Steady State Fluxes The flux distribution is calculated by using the mass balances of internal metabolites (Varma and Palsson 1994). It is generally assumed that the internal metabolites are at pseudo steady state, since metabolic transients are fast compared to environmental changes (Varma and Palsson 1994). Therefore, the mass balance is rewritten as follows: S .v 0 vi 0 i irreversible reactions (1) where v is the flux distribution vector, and S is the stoichiometric matrix. One of the common problems in solving system (1) to obtain unknown fluxes is that the rank of S is generally smaller than the number of unknown fluxes which leads to an underdetermined system. To overcome this limitation, a range of possible values for each flux is determined using linear programming where each flux is maximized or minimized while allowing all other flux values to vary (Burgard et al. 2001; Llaneras and Picó 2007; Mahadevan and Schilling 2003). To account for experimental errors, the equation is modified such that each of the measured fluxes allow to very between (vmin and vmax) as follows: 2 Maximize / Minimize v j , j unknown fluxes Subject to S .v 0 vmmin vm vmmax , vi 0, m measured fluxes (2) i irreversible fluxes Problem (2) is solved 2*nu times where nu is total number of unknown fluxes and each time, a flux distribution vector is obtained. Monte Carlo Sampling Calculating the range of each flux using problem (2), the solution space of steady state fluxes can be described as follows: S .v 0 v min v v max , v all fluxes (3) To determine a representative sample of flux distributions of the solution space (Equation 3), Monte Carlo Sampling is used. We adopted an algorithm provided by Wiback and her coworkers (Wiback et al. 2004) and modified to add constraints given in equation (3). This method has been widely used to analyze metabolic networks and its applications were recently reviewed by Schellenberger and Palsson (Schellenberger and Palsson 2009). The algorithm is straightforward. After assigning uniformly random values for independent variables within the known ranges, the dependent variables are linearly calculated using the stoichiometric matrix. The results are random flux distributions. Back-calculating the dependent flux values can result in invalid flux distributions (Wiback et al. 2004) since the solution space obtained from sampling analysis can be larger than the actual one defined by equation (3) (see Figure S1) due to the fact that only the independent fluxes are bounded in sampling analysis. The valid flux distribution is one that meets the “range criteria” given in equation (3). In this study, the number of independent 3 variables is 31 since we use a system with 83 fluxes (or variables) and a stoichiometric matrix having a rank of 52. The steady state flux space obtained from the sampling analysis is larger than the actual one which is shown in Figure S2 and the valid flux distributions are the ones in the actual solution space. Singular Value Decomposition A matrix including all valid steady state flux distributions obtained from Monte Carlo sampling is constructed (matrix A). To identify the basic trends in the matrix A, SVD analysis is performed. SVD analysis decomposes the flux distribution matrix into three matrices (Henry and Hofrichter 1992; Klema and Laub 1980): A U V T (4) where U and V are orthonormal matrices with columns corresponding to left and right singular vectors of A, respectively; and Σ is a diagonal matrix containing the singular values (σi) of matrix A, arranged in descending order. The magnitude of the singular values in Σ indicates the relative contribution of the singular vectors in reconstructing the matrix A. U is a combination of the orthonormal bases of the column space and left null space of A. Therefore, in this analysis, the matrix U is further investigated, since each column of U represents a dominant pattern in the flux vector solution space. To determine the relative contribution of each column of U in reconstructing the matrix A, the following weight value is used: wi i i i 1,..., ne (5) i where wi is the weight of eigenvector i, σi is the corresponding singular value, and ne is the number of nonzero singular values or number of eigenvectors. 4 The first eigenvector corresponds to a valid flux distribution since it satisfies both mass and reversibility constraints given in Equation (1). In a solution cone described by a set of vectors, the first eigenvector comes up through the geometric center of the cone, and is close to the part of the cone where the vectors are densely accumulated (Price et al. 2003). Moreover, the first eigenvector contributing most in reconstructing the matrix of flux vectors also gives the dominant flux-pattern. Subsequent eigenvectors define the direction of most variance in the subspace. Although all subsequent eigenvectors obey the mass balance constraints, they do not necessarily satisfy reaction directionality constraints. Each subsequent vector can identify key network branch points or tradeoffs in the network and represent how tradeoff points influence the flux distribution. A more detailed analysis about the physiological meaning of eigenvectors can be found in reference (Price et al. 2003). To simplify the interpretation of initial eigenvectors that capture most of the variation in the matrix of flux samples, basis rotation method can be applied to these eigenvectors (Barrett et al. 2009). They are rotated using the orthogonal varimax rotation procedure (Kaiser 1958) where the variance or sum of the squared vector loadings or coefficients across the column is maximized. The first 5 eigenvectors and their fractional contributions obtained from different number of samples (104 and 105) are given in Table SII. Statistics Comparisons were performed using a 2-tailed Student’s t-test, and p<0.05 was the criterion used for statistical significance. 5 Note: In this work, all optimization problems are solved using MATLAB (Mathworks Inc, Massachusetts) and GAMS with the interface program MAT-GAMS in a Pentium(R) processor at 2.80 GHz with 1.00 GB of memory. Tables Table SI. Hepatic Network Model. Table SII. The weight values of first five eigenvectors. Figure Legends Figure S1. Monte Carlo sampling used to identify the flux vectors within the steady state flux space. Larger square represents the flux space obtained from sampling method since only independent fluxes are bounded. Smaller square (grey color) is actual flux space where all fluxes are bounded. Each dot represents a random solution. Figure S2. The steady state flux space of hepatic metabolism in fasted state (A) and fed sate (B) obtained from the flux spectrum approach (grey space) and sampling analysis (space surrounded by black lines). 6 References Banta S, Vemula M, Yokoyama T, Jayaraman A, Berthiaume F, Yarmush ML. 2007. Contribution of gene expression to metabolic fluxes in hypermetabolic livers induced through burn injury and cecal ligation and puncture in rats. Biotechnology and Bioengineering 97(1):118-137. Barrett CL, Herrgard MJ, Palsson B. 2009. Decomposing complex reaction networks using random sampling, principal component analysis and basis rotation. Bmc Systems Biology 3. Burgard AP, Vaidyaraman S, Maranas CD. 2001. Minimal reaction sets for Escherichia coli metabolism under different growth requirements and uptake environments. Biotechnology Progress 17(5):791-797. Chan C, Berthiaume F, Lee K, Yarmush ML. 2003. Metabolic flux analysis of cultured hepatocytes exposed to plasma. Biotechnology and Bioengineering 81(1):33-49. Henry ER, Hofrichter J. 1992. Singular Value Decomposition - Application to Analysis of Experimental-Data. Methods in Enzymology 210:129-192. Kaiser H. 1958. The varimax criterion for analytic rotation in factor analysis. Psychometrika 23(3):187-200. Klema V, Laub A. 1980. The singular value decomposition: Its computation and some applications. Automatic Control, IEEE Transactions on 25(2):164-176. Lee K, Berthiaume F, Stephanopoulos GN, Yarmush DM, Yarmush ML. 2000. Metabolic Flux Analysis of Postburn Hepatic Hypermetabolism. Metabolic Engineering 2(4):312-327. Lee K, Berthiaume F, Stephanopoulos GN, Yarmush ML. 2003. Profiling of dynamic changes in hypermetabolic livers. Biotechnology and Bioengineering 83(4):400-415. 7 Llaneras F, Picó J. 2007. An interval approach for dealing with flux distributions and elementary modes activity patterns. Journal of Theoretical Biology 246(2):290-308. Mahadevan R, Schilling CH. 2003. The effects of alternate optimal solutions in constraint-based genome-scale metabolic models. Metabolic Engineering 5(4):264-276. Price ND, Reed JL, Papin JA, Famili I, Palsson BO. 2003. Analysis of metabolic capabilities using singular value decomposition of extreme pathway matrices. Biophysical Journal 84(2):794-804. Schellenberger J, Palsson BO. 2009. Use of Randomized Sampling for Analysis of Metabolic Networks. Journal of Biological Chemistry 284(9):5457-5461. Varma A, Palsson BO. 1994. Metabolic Flux Balancing - Basic Concepts, Scientific and Practical Use. Bio-Technology 12(10):994-998. Wiback SJ, Famili I, Greenberg HJ, Palsson BO. 2004. Monte Carlo sampling can be used to determine the size and shape of the steady-state flux space. Journal of Theoretical Biology 228(4):437-447. 8 Tables Table SI. Hepatic Network Model. Enzymes and explanations Reaction No Glycolytic and Gluconeogenic Pathways Reaction 1a,b,* Hexokinasea , Glucose-6-Paseb Glucose + Pi <==> Glucose-6-P Reaction 2a,b,* Phosphoglucose isomerase a,b Glucose-6-P <==> Fructose-6-P a b Reaction 3 a,b,* PFK-1 , Fructose-1,6-Pase-1 Fructose-6-P + Pi <==> Fructose-1,6-P2 Triose P-isomerase, fructose biphosphate Reaction 4 a,b,* aldolase Fructose-1,6-P2 <==> 2 Glyceraldehyde-3-P Glyceraldehyde-P dehydrogenase, 3phosphoglycerate kinase, Reaction 5 a,b,* phosphoglyceromutase, enolase Glyceraldehyde-3-P + Pi + NAD+ + ADP <==> ATP + NADH + PEP Reaction 6 a Pyruvate kinase ADP + PEP ==> ATP + Pyruvate Reaction 7 a PDH NAD+ + Pyruvate ==> CO2 + NADH + Acetyl-CoA Reaction 8 b PEPCK Oxaloacetate + GTP ==> CO2 + PEP + GDP Reaction 9 b Pyruvate carboxylase CO2 + ATP + Pyruvate ==> Pi + ADP + Oxaloacetate Pentose Phosphate Pathway Glucose-6-P dehydrogenase and 3 additional Reaction 10 steps Glucose-6-P + 12 NADP+ + 7 H2O ==> 6 CO2 + 12 NADPH + Pi + 12 H+ 9 Lactic acid production Reaction 11 Lactate dehydrogenase NAD+ + Lactate <==> NADH + Pyruvate TCA cycle Reaction 12 Citrate synthase Oxaloacetate + Acetyl-CoA ==> Citrate + CoA-SH Reaction 13 Aconitase, isocitrate dehydrogenase NAD+ + Citrate <==> CO2 + NADH + a-ketoglutarate Reaction 14 a-ketoglutarate dehydrogenase NAD+ + CoA-SH + a-ketoglutarate ==> CO2 + NADH + Succinyl-CoA Succinyl-CoA synthase and succinate Reaction 15 dehydrogenase Pi + GDP + Succinyl-CoA + FAD <==> GTP + Fumarate + FADH2 Reaction 16 Fumarase Fumarate <==> Malate Reaction 17 Malate dehydrogenase NAD+ + Malate <==> NADH + Oxaloacetate Urea cycle Reaction 18 Arginase Arginine ==> Urea + Ornithine Carbonate dehydratase, carbamoyl-P synthase, Reaction 19 ornithine transcarbamylase CO2 + 2 ATP + Ornithine + NH4+ <==> 2 Pi + 2 ADP + Citrulline Argininosuccinate synthetase, Reaction 20 argininosuccinase. ATP + Citrulline + Aspartate ==> 2 Pi + Fumarate + Arginine + AMP Amino acid metabolism Reaction 21 Alanine aminotranferase NAD+ + Alanine <==> NADH + Pyruvate + NH4+ 10 Reaction 22 Serine dehydratase Serine ==> Pyruvate + NH4+ Transaminase, Reaction 23 3-mercaptopyruvate sulfurtransferase NAD+ + Cysteine + H2SO3 <==> NADH + Pyruvate + NH4+ + H2S2O3 Reaction 24 Threonine 3-dehydrogenase, acetyl-CoA ligase NAD+ + CoA-SH + Threonine ==> NADH + Acetyl-CoA + Glycine Glycine hydroxymethyltranferase, glycine Reaction 25 cleavage system NAD+ + 2 Glycine <==> CO2 + NADH + NH4+ + Serine Methylmalonyl-CoA epimerase, Reaction 26 Methylmalonyl-CoA mutase S-Methylmalonyl-CoA <==> Succinyl-CoA Reaction 27 Lysine metabolism (8 steps) 5 NAD+ + CoA-SH + FAD + Lysine ==> 2 CO2 + 5 NADH + FADH2 + 2 NH4+ + Acetoacetyl-CoA Reaction 28 Phenylalanine hydroxylase Phenylalanine + Tetrahydrobiopterin + O2 ==> Dihydrobiopterin + Tyrosine Reaction 29 Tyrosine metabolism (5 steps) NAD+ + 2 O2 + Tyrosine ==> CO2 + NADH + Fumarate + NH4+ + Acetoacetate Reaction 30 Glutamate dehydrogenase NAD+ + Glutamate <==> NADH + a-ketoglutarate + NH4+ Reaction 31 Glutaminase Glutamine ==> NH4+ + Glutamate Reaction 32 Ornithine metabolism (2 steps) NADP+ + NAD+ + Ornithine ==> NADPH + NADH + NH4+ + Glutamate + H+ Proline oxidase, 1-pyrroline-5-carboxylate Reaction 33 dehydrogenase 0.5 NADP+ + 0.5 NAD+ + 0.5 O2 + Proline ==> 0.5 NADPH + 0.5 NADH + Glutamate Reaction 34 Histidine metabolism (4 steps) Histidine + THF ==> NH4+ + Glutamate + 2-formimino-THF Reaction 35 Methionine metabolism (5 steps) 2 ATP + NAD+ + CoA-SH + Serine + Methionine ==> CO2 + Pi + NADH + NH4+ + Cysteine + PPi + Adenosine + Propinoyl-CoA Reaction 36 Propinoyl-CoA carboxylase CO2 + ATP + Propinoyl-CoA ==> Pi + ADP + S-Methylmalonyl-CoA 11 Reaction 37 Aspartate aminotransferase NAD+ + Aspartate <==> NADH + Oxaloacetate + NH4+ Reaction 38 Asparaginase Asparagine ==> NH4+ + Aspartate Valine metabolism (7 steps) 0.5 NADP+ + 3.5 NAD+ + FAD + 2 H2O + valine ==> 2 CO2 + 0.5 NADPH + 3.5 NADH + FADH2 + NH4+ + Propinoyl-CoA + 3 H+ Reaction 39 Isoleucine Metabolism (6 steps) 0.5 NADP+ + 2.5 NAD+ + FAD + 2 H2O + isoleucine ==> CO2 + 0.5 NADPH + 2.5 NADH + Acetyl-CoA + FADH2 + NH4+ + Propinoyl-CoA + 3 H+ Reaction 40 Leucine Metabolism (6 steps) 0.5 NADP+ + ATP + 1.5 NAD+ + FAD + H2O + leucine ==> 0.5 NADPH + Pi + ADP + 1.5 NADH + Acetyl-CoA + FADH2 + NH4+ + 2 H+ + Acetoacetate Reaction 41 Fatty acid metabolism Reaction 42 Hepatic Lipase Palmitoylglycerol + ATP + NAD+ <==> Pi + 3 Palmitate + ADP + NADH + glycerol Reaction 43 Glycerol-3-P dehydrogenase NAD+ + glycerol <==> Glyceraldehyde-3-P + NADH Reaction 44 Fatty acid oxidation (7x4 steps) ATP + 7 NAD+ + Palmitate + 8 CoA-SH + 7 FAD ==> 2 Pi + 7 NADH + 8 Acetyl-CoA + 7 FADH2 + AMP Reaction 45 Fatty acid synthesis (7x4 steps) 14 NADPH + 7 ATP + 8 Acetyl-CoA + 14 H+ ==> 14 NADP+ + 7 Pi + Palmitate + 7 ADP + 6 H2O Reaction 46 Thiolase (Ketogenesis) 2 Acetyl-CoA <==> 2 CoA-SH + Acetoacetyl-CoA Reaction 47 HMG-CoA synthase and lyase (Ketogenesis) Acetoacetyl-CoA ==> CoA-SH + Acetoacetate Reaction 48 B-OH-butyrate dehydrogenase (Ketogenesis) NADH + Acetoacetate <==> NAD+ + B-OH-butyrate Electron-chain reactions Reaction 49 Electron transport system 3 Pi + 3 ADP + NADH + 0.5 O2 + H+ ==> 3 ATP + NAD+ + H2O Reaction 50 Electron transport system 2 Pi + 2 ADP + FADH2 + 0.5 O2 ==> 2 ATP + FAD + H2O 12 Protein synthesis or degradation Reaction 51** Albumin synthesis or degradation Glycogenesis/Glycogenolysis Reaction 52 Glycogen metabolism (3 steps) glycogen <==> Glucose-6-P External Reactions Reaction 53 glycogen <==> Reaction 54 Palmitoylglycerol <==> Reaction 55 O2 <==> Reaction 56 CO2 <==> Reaction 57 albumin <==> Reaction 58 Glucose <==> Reaction 59 Lactate <==> Reaction 60 Urea <==> Reaction 61 NH4+ <==> Reaction 62 Aspartate <==> Reaction 63 Glutamate <==> Reaction 64 Serine <==> 13 Reaction 65 Asparagine <==> Reaction 66 Glycine <==> Reaction 67 Glutamine <==> Reaction 68 Histidine <==> Reaction 69 Threonine <==> Reaction 70 Arginine <==> Reaction 71 Alanine <==> Reaction 72 Proline <==> Reaction 73 Tyrosine <==> Reaction 74 Cysteine <==> Reaction 75 valine <==> Reaction 76 Methionine <==> Reaction 77 isoleucine <==> Reaction 78 Ornithine <==> Reaction 79 leucine <==> Reaction 80 Lysine <==> Reaction 81 Phenylalanine <==> Reaction 82 Acetoacetate <==> 14 Reaction 83 B-OH-butyrate <==> Notes: a or b stands for a reaction in glycolysis or gluconeogenesis, respectively. * These reactions indicate futile cycle (reversible reaction) composed of two different reactions, one of which is from glycolytic pathway and the other from gluconeogenic pathway. ** Both protein synthesis and degradation are given in one reaction, which results in reversible reaction. 15 Table SII. The weight values of first five eigenvectors. N=104 N=105 Weights Fasted Fed Fasted Fed w1 0.724645 0.723939 0.653732 0.654452 w2 0.085830 0.085985 0.118614 0.118727 w3 0.069688 0.069972 0.074555 0.073390 w4 0.049681 0.049363 0.048616 0.048944 w5 0.037997 0.038441 0.034547 0.034627 ∑ 0.967841 0.9677 0.930064 0.93014 Notes: N is the number of flux samples. ∑ implies fractional contribution of first five eigenvectors to the description of the matrix of flux vector. 16 Figure S1. 17 Figure S2. 18 19

























![Exercise 3.1. Consider a local concentration of 0,7 [mol/dm3] which](http://s1.studyres.com/store/data/016846797_1-c0b17e12cfca7d172447c1357622920a-150x150.png)