* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Transcriptional Control
Survey
Document related concepts
Transcript
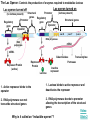
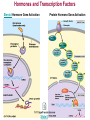
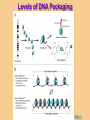
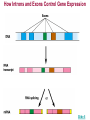
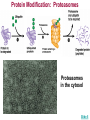
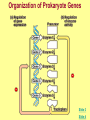
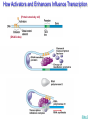
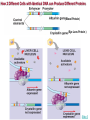
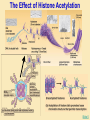
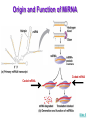
Gene Regulation: The control of protein production Chapter 18 pages 360 -389 I. Differences between gene regulation of prokaryotes and eukaryotes Prokaryotes A. Small circular genome B. Unicellular Eukaryotes A. Large genome, many strands, genes randomly distributed among the strands B. Multicellular (DNA instructions for every cell type of the organism) (DNA instructions for only one cell type) C. Most DNA as “junk DNA” C. Most DNA codes for protein (Repetitive DNA that make introns, centromeres, telomeres) D. Most of the genome is expressed D. Little of the genome is expressed (20% max) E. Mechanism: The Operon E. Mechanisms: 1. Chromosome Structure 2. Transcriptional Control 3. Post-Transcriptional Control 4. Translational Control 5. Post-Translational Control II. The Operon System Parts of the Operon A. Structural Genes: The sequence of genes required to produce the desired product. Many are part of the same metabolic pathway and are in a specific order. B. Promoter: Area of the DNA that allow attachment to the of RNA polymerase C. Operator: Part of the DNA that when bound to a specific protein will prevent the attachment of RNA polymerase D. Regulatory gene: Produces a protein that interacts with the operator E. Types 1) Inducible operons example: Lac operon 2) Repressible operons example: Tryp operon The Lac Operon: Controls the production of enzymes required to metabolize lactose Lac operon turned on Lac operon turned off Structural Promoter genes Regulatory Promoter gene Operator Operator (lactose present) (no lactose present) Regulatory gene Structural genes Lac Z Lac Y Lac A RNA polymerase RNA polymerase mRNA mRNA Galactosidase Repressor Protein (active) Repressor Protein Transacetylase Permease Lactose Inactive repressor 1. Active repressor binds to the operator 1. Lactose binds to active repressor and deactivates the repressor 2. RNA polymerase can not transcribe structural genes 2. RNA polymerase bonds to promoter allowing the transcription of the structural genes Why is it called an “inducible operon”? Video Slide 2 The Tryp Operon: Controls the enzymes required to build tryptophan (amino acid) Operon turned off Operon turned on (Tryptophan present) Promoter Structural genes Regulator gene (Tryptophan absent) Promoter Regulator gene Operator Operator RNA Polymerase RNA Polymerase mRNA Active Repressor Tryp E Tryp D Tryp C Tryp B mRNA Tryp A mRNA Inactive repressor Active Repressor Inactive repressor Enzymes required for the synthesis Tryptophan Tryptophan 1. Inactive repressor is made active in the presence of tryptophan 2. Tryptophan is not synthesized since RNA polymerase cannot bond to promoter 1. Inactive repressor can not bond to the operator 2. RNA polymerase bonds to DNA so the enzymes for the synthesis of tryptophan are transcribed Why is this called a “repressible operon”? Slide 2 Areas of Control in Eukaryotic Cells Control Type Methods 1. Histone Acetylation Chromosome Structure 2. DNA Methylation Transcriptional Control Post Transcriptional Control Translational Control 1. Spreads out nucleosomes to facilitate transcription 2. Makes genes inaccessible (Barr Bodies and Imprinting) 1. Transcription factors bond at promoter site to increase RNA polymerase affinity 1. Hormones & Signal Transduction 2. Activator proteins bind to DNA Enhancers to bend DNA to form transcription initiation complex 2. Formation of transcription complex 1. mRNA processing introns and exons Albumin (liver) vs. Crystalin (lens) 1. Intron and exon splicing will modify the protein produced 2. mRNA export 2. 5’ Cap and Poly A tail can prevent export to cytoplasm 3. Long lived or short lived mRNA 3. RBC have long lived mRNA 1. Translation Repressor Protein binds to 5’ Cap preventing translation 2. Non-Coding RNAs a. MicroRNA (miRNA) Post Translational Control Function / Examples b. Small Interfering RNA (siRNA) 1. Protein Processing modifies the activity of proteins 2. Selective degradation 1. mRNA prevented from translation in egg until after fertilization 2. Non-coding RNA that binds with protein that degrades or prevents the translation of coding RNA 1. Insulin and Digestive Enzymes 2. Targeting by a) ubiquitin b) Proteasomes Hormones and Transcription Factors Steroid Hormone Gene Activation Protein Hormone Gene Activation Slide 5 Slide 1 Levels of DNA Packaging Slide 5 How Introns and Exons Control Gene Expression Slide 5 Protein Modification: Proteasomes Proteasomes in the cytosol Slide 5 Organization of Prokaryote Genes Slide 3 Slide 4 Slide 2 Slide 1 Androgens are illegal because they work!! Slide 6 How Activators and Enhancers Influence Transcription (Proteins made by cell) (DNA Codes) Slide 5 How 2 Different Cells with Identical DNA can Produce Different Proteins (Blood Protein) (Eye Lens Protein ) Slide 5 The Effect of Histone Acetylation Slide 5 Origin and Function of MiRNA Coded mRNA Coded mRNA Slide 5