* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Selection on quantitative characters
Public health genomics wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Genome (book) wikipedia , lookup
History of genetic engineering wikipedia , lookup
Dual inheritance theory wikipedia , lookup
Designer baby wikipedia , lookup
Koinophilia wikipedia , lookup
Genetic drift wikipedia , lookup
Human genetic variation wikipedia , lookup
Polymorphism (biology) wikipedia , lookup
Behavioural genetics wikipedia , lookup
Group selection wikipedia , lookup
Population genetics wikipedia , lookup
Microevolution wikipedia , lookup
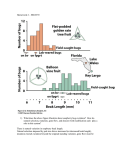
February 5th, 2010 Bioe 109 Winter 2010 Lecture 13 Selection on quantitative traits Selection on quantitative traits - From Darwin's time onward, it has been widely recognized that natural populations harbor a considerably degree of genetic variation. - Darwin came to this conclusion from the experiences of animal and plant breeders of his day and relied on it heavily in developing his theory of evolution by natural selection. - the form of variation envisaged by Darwin to be of fundamental importance for evolutionary change was “continuous”, or what we know call “polygenic” or “quantitative”. - quantitative characters exhibit continuous variation among individuals. - unlike discrete characters, it is not possible to assign phenotypes to distinct groups. - the statistical approach to studying such traits is referred to as quantitative genetics. - the vast majority of morphological, physiological, and behavioral characters are polygenic and thus understanding this class of variation is fundamental for evolutionary biologists. - there are two characteristics of quantitative traits: 1. they are influenced by many genetic loci. 2. they exhibit variation due to both genetic and environmental effects. - we are just now beginning to understand what polygenes are, and just how many such loci are involved in affecting the expression of a quantitative character. - our ignorance is profound. - new insights are being provided by the ability to map QTLs, or "quantitative trait loci". - this is done by melding traditional breeding experiments of quantitative genetics with fineresolution linkage maps. [A linkage map is a detailed "map" of the location of genetic markers dispersed throughout the genome of the species.] - identifying and characterizing QTL loci is going to dominate the field of quantitative genetics for many years to come. - there appears to be nothing magical about QTLs - they possess possess multiple alleles, exhibit varying degrees of dominance, and experience selection and drift. - some QTLs exhibit stronger effects than others – these are called major effect and minor effect genes, respectively. - the total number and relative contributions of major effect and minor effect genes underlies the genetic architecture of the trait. - mapping QTLs is expensive, labor intensive and fraught with statistical problems! What is heritability? - the key to understanding selection on quantitative characters involves the concept of heritability. - heritability estimates the proportion of the total phenotypic variation that is due to genetic rather than environmental effects. - more formally, it is possible to separate the total phenotypic variation of a quantitative trait into two components: environmental and genetic. - let VP = total phenotypic variance VG = genetic variance VE = environmental variance VP = VG + VE - then heritability = VG / VP (broad-sense) - this is called “broad sense” heritability. - we will be considering another type of heritability called “narrow sense” heritability. - this is different from broad sense heritability in incorporating only the additive component of genetic variance. - VG can be partitioned into three components due to additive (VA), dominance (VD), and epistatic (VE) effects. VG = VA + VD + VE heritability = h2 = VA/VP (narrow-sense) - of these three components, it is only the additive component that responds to selection. Example of additive gene action - suppose two genes (A and B) contribute to a quantitative character (say, abdominal bristle number in Drosophila melanogaster) - for alleles that act in an additive fashion, the heterozygote exhibits a phenotype that is exactly intermediate between the two alternative homozygotes: B1B1 A1A1 0 A1A2 2 A2A2 4 B1B2 B2B2 1 3 5 2 4 6 - suppose we selected for increased bristle number in a population with a low mean number of bristles. - this would result in the substitution of the A2 and B2 alleles. Estimating heritability - how do we estimate heritability? - there are many different ways, but the most common is to compare the resemblance between parents and their offspring. - suppose we are interested in obtaining an estimate of heritability for growth rate for a species. - one way to do this would be to perform pair matings between different individuals and then rear the progeny of these crosses under similar conditions (to hold the environmental effect constant among families). - we allow the progeny of these matings to reach the same age as the parents. - we then take the mean of each pair of parents - this is called the midparent value. - we then take the mean of each set of progeny - we can call this the offspring value. Cross F 1 100 2 160 3 150 etc… M “Midparent value” “Offspring value” 140 110 140 120 135 145 132 126 153 - data from many families is preferred. - we can then estimate heritability by regressing the offspring values against the midparent values. - the slope of this regression line is an estimate of heritability. - here are some other comparisons: Comparison Slope Midparent-offspring Parent-offspring Half-sibs First cousins h2 1/2h2 1/4h2 1/8h2 - as the groups become less related, the precision of the h2 estimate is reduced. - once we have an estimate of heritability, we can predict the response to selection by the following prediction equation: R = h2 S - here, R is called the response to selection and S is the selection differential. - the selection differential is equivalent to the strength of selection acting on the trait. - it is quantified by the difference between the mean of the selected group and the original population. - the heritability usually remains constant over a sizable number of generations giving us a constant and predictable response to selection. Example: Beak depth in the large ground finch, Geospiza magnirostris - in this species, the heritability of beak depth is about 0.72. - the initial population mean was 8.82 mm. - in one episode of intense selection, the mean beak depth of survivors was 10.11 mm. Selection differential = S = 10.11 – 8.82 = 1.29 mm Response to selection, = h2S = (0.72)(1.29) = 0.93 mm - thus, beak depth in the next generation is predicted to be 8.82 + 0.93 = 9.75 mm - this assumes that the finch population will not experience drift, mutation, or migration. Heritability estimates in nature - what are heritability estimates in natural populations? - they are commonly very high. - here are some heritability estimates for a population of medium ground finches in the Galapagos studied by Peter Boag (1983) in comparison to a population of N. American mainland song sparrows studied by Smith and Zach (1970): Character Body weight Wing length Tarsus length Bill length Bill depth Bill width Ground Finch 0.91 0.84 0.71 0.65 0.79 0.90 Song Sparrow 0.04 0.13 0.32 0.33 0.51 0.50 - why would the heritabilities of these traits be so much higher in the Galapagos species? - in a review paper, Mousseau and Roff (1987) compared the heritabilities of different kinds of quantitative traits: Trait Life history Physiological Sample size 341 104 Mean heritability 0.262 0.330 Behavioral Morphological 105 570 0.302 0.461 - these studies illustrate the abundance of additive genetic variation in natural populations for most quantitative characters. Selection on quantitative characters - natural selection acting on quantitative traits can take three basic forms: stabilizing, directional, or disruptive. 1. Stabilizing. e.g., gall size in goldenrods caused by the fly, Eurosta altissima. - this is probably the most common form of selection. - here, individuals with intermediate phenotypes have the highest fitness and those with extreme phenotypes the lowest fitness. - selection thus does not change the mean of the character but reduces its variance. - small galls made by E. altissima are preferentially attacked by parasitoid wasps. - large galls are attacked at a higher rate by birds. - the result is a substantial reduction in the variance of gall size with intermediate sized galls surviving at much higher rates. 2. Directional. e.g., selection on corn oil content. - this form of selection increases or decreases the mean of a quantitative character in a consistent manner over a number of generations. - it also reduces the amount of variation in the population, but this depends on the intensity of selection and the heritability of the trait. 3. Disruptive. e.g., seed-cracker finches in Cameroon. - disruptive selection occurs with extreme values of a phenotypic trait have the highest fitness. - unlike stabilizing and directional, this form of selection acts to increase the amount of phenotypic variation of the selected trait but may not change the mean. - it is thought to occur less frequently than the other two.