* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download ppt - Sol Genomics Network
Gene regulatory network wikipedia , lookup
Gene desert wikipedia , lookup
Promoter (genetics) wikipedia , lookup
DNA sequencing wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Non-coding DNA wikipedia , lookup
Ridge (biology) wikipedia , lookup
X-inactivation wikipedia , lookup
Genomic imprinting wikipedia , lookup
Community fingerprinting wikipedia , lookup
Gene expression profiling wikipedia , lookup
Copy-number variation wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Exome sequencing wikipedia , lookup
Molecular evolution wikipedia , lookup
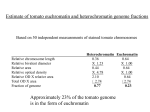
Expanding the Tool Kit for BAC Extension Summary of completion criteria developed for NSF Tomato Sequencing Workshop January 14, 2007 Resources for Additional Anchor BACs • 1) Mapping of random low copy BACs to create a pool of new seed BACs that could serve all 12 chromosomes. - low copy BACs identified by the group of Tabata et al. - BAC end sequences mapped to S. lycopersicum X S. pennellii ILs. - India, Korea, others • 2) Sequencing of 2 million plus reads derived from low copy BACs defined by Dr. Tabata et al. • 3) identification of low copy cosmids (US) that would also be end sequenced by Japan. • 3) BAC OVERGO screen of MboI library – – – – Large inserts 130 kb avg. FPC data generated by Sanger 10 - 20 probes per country Screening in next several months 4) Industry anchored BACs Goals of the International Tomato Genome Sequencing Project Estimate of tomato euchromatin and heterochromatin genome fractions Based on 50 independent measurements of stained tomato chromosomes Relative chromosome length Relative bivalent diameter Relative area Relative optical density Relative OD X relative area Total OD X area Fraction of genome Heterochromatin Euchromatin 0.36 0.64 X 1.23 X 1.00 0.44 0.64 X 4.78 X 1.00 2.10 0.64 / 2.74 / 2.74 0.77 0.23 Approximately 23% of the tomato genome is in the form of euchromatin Mb size of tomato euchromatin based on cytogenetic measurements 0.95 pg / tomato genome X 0.23 (euchromatin fraction) = 0.22 pg 965 X 106 pb/pg = 2.12 x 108 bp or 212 Mb (705 Mb heterochromatin) Estimate of tomato euchromatin size based on available EST and genome sequence 15.5 Mb available sequence (Fall 2006) 8,097 high quality unigene set - all available full-length tomato genes in GENBANK - TIGR full-length cDNA sequences (redundantly sequenced) - SGN unigene contigs with 5 or more ESTs - redundnacy correction 456 of 8,097 genes found in available genome sequence (5.6%) Correcting for 85% expectation yields 6.6% of target gene space 15.5/0.066 = 239 Mb tomato euchromatin target Estimate of gene space missed in this approach: Genes missed in centromere (rice chromosome 8 - 86 genes) 12 x 86 = 1032 centromere genes Exelisis heterochromatin BACs - 2 BACs representing 200 kb were sequenced and one gene identified. 705,000 kb in herterochromatin (slide 2) 705,000 / 200 = 3525 heterochromatin genes 35,000 estimated tomato genes - 1032 - 3525 = 30,500 genes (87%) Correcting for 3% euchromatin gaps (as in rice) results in 85% of total tomato gene space is anticipated to be recovered under the International Tomato Genome Sequencing Project. Sequencing standards A “finished BAC” is defined as…… • it contains an error rate of less than 1:10,000 bases and continuous sequence across the entire BAC (HTGS phase 3) • has an average of 8-fold redundancy in sequencing coverage with a minimum of one high quality read in both directions at any specific sequence • all reasonable state of the art approaches available at the time for gap filling will be used Tomato euchromatin completion criteria: 1) complete sequencing of the major euchromatin “arms” flanking each of the 12 tomato chromosomes 2) to a degree of completion comparable to the standards of completion used to guide the international rice genome sequencing project (IRGSP, 2005) ---- e.g. anticipate 4 - 6 gaps per chromosome. Furthermore: 1) Sequence to at least the closest mapped marker to the euchromatin / heterochromatin border . 2) Attempt to walk until characteristic heterochromatin repeats are identified and at minimum define the size of the remaining gap In summary, the target of the international genome sequencing effort is sequencing of the euchromatin arms of all twelve tomato chromosomes which we estimate will represent approximately 85% of the tomato gene space.