* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Parallel Chemical Genetic and Genome
Survey
Document related concepts
Deoxyribozyme wikipedia , lookup
Microevolution wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Point mutation wikipedia , lookup
Primary transcript wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
History of genetic engineering wikipedia , lookup
Mir-92 microRNA precursor family wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
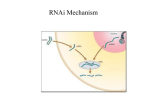
RNA silencing wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Transcript
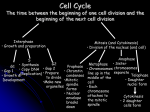
Parallel Chemical Genetic and Genome-Wide RNAi Screens Identify Cytokinesis Inhibitors and Targets Ulrike Eggert, Amy Kiger, Constance Richter, Zachary Perlman, Norbert Perrimon, Tomothy Mitchison, Christine Field Pat Ekpanyaskun Mar 3, 2005 To study protein function Small molecule inhibitors – directly inhibit protein fx. – search by phenotypic screening -> i.d. targets / pathway – binding of irrelevant prot. RNA interference RNAi – dsRNA targets destruction of RNA – inhibit prot. fx in a genome-wide basis Mitosis • daughter cells are separated by constriction of a cleavage furrow This paper Goal Complete list of prot. required for cytokinesis in Drosophila cells Approach Genome-wide RNAi screening & small molecule inhibitors Screening dsRNA screening Small molecule screening - 51,000 inhibitors - >90% annotated genome dsRNAs - incubate ~4 d 2d Immunofluorescence -dsRNAs of 214 genes -1 new gene (Borr/CG4454) -50 inhibitors (commercial lib, bioactive, nat prod. extract) Predicted Functional Annotations of 214 Genes Associated with RNAi Binucleate Phenotypes Penetrance of binucleate phenotype 24% 40% 32% 44% 28% Inhibitor RNAi 6% 20% 29% 51% 74% Phenotypic classes 24% 40% 28% Binucleate binucleate w large, diffuse DNA 8% Inhibitor RNAi 12% Low cell count 29% 51% Low cell count Binucleate with MT extension Two phenotypes correlate in both methods RNAi inhibitors dsRNA targeting Act5C OR Cytochalasin D Two phenotypes correlate in both methods •Binucleate with diffuse DNA •Chromosomal passenger protein that co-localize with Aurora B throughout mitosis & cytokinesis Aurora B Andrews et al 2003 Chromosomal passenger prot Localize on centromeres (prophase-metaphase) Relocalize to spindle midzone and cortex (anaphase) Disruption causes loss of phosphorylation on histone H3 and failure in cytokinesis Two phenotypes correlate in both methods •Chromosomal passenger protein that co-localize with Aurora B throughout mitosis & cytokinesis Borr/CG4454 co-localizes with Aurora B •Borr is required for INCENP localization Aurora B RNAi and Binucleine 2 shares same phenotype and affect localization of INCENP Cells treated to aurora B dsRNA, borr/CG4454 dsRNA or Binucleine 2 results in an absence of H3 Summary i.d. new prot. involved in cytokinesis and new small molecule that inhibits it 214 prot. important for cytokinesis in Drosophila Borr/CG4454 is a chromosomal passenger prot that co-localize with Aurora B Depletion had a profound effect on cytokinesis Binucleine 2 as a cancer drug ? Message Integration of data from different tools to answer biological questions Comment Subjective to visual inspection