* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Lecture_12_2005
Vectors in gene therapy wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Minimal genome wikipedia , lookup
Point mutation wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Genome evolution wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
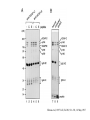
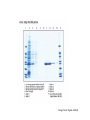
• Protein domains vs. structure domains - an example. Genome of the week • Nanoarchaeum equitans - archaea – Hyperthermophile – Diverged early in evolution from other archaea – New kingdom of archaea? • Obligate symbiont with Ignicoccus • Smallest completely sequenced genome – <500kB – Genome reduction observed in symbionts (Schmidt) – Is N. equitans a “primitive” archaea or is the genome undergoing reductive evolution? • N. equitans lacks genes necessary for many aspects of central metabolism. – Can’t make lipids, vitamins, amino acids, etc. – Parasite, not symbiont? First archaea • Genome is quite compact. – 95% of genome codes for genes. 552. • Not primitive. – Has complete set of information pathway and cell cycle genes found in archaea. • No longer undergoing reductive evolution. – Normally would find pseudogenes - not found. Protein complexes - why? • Proteins often function as large, multisubunit complexes. – ribosomes • Can get clues about the function of a protein by knowing what other proteins it contacts. Protein:protein interactions • Genetic approach – Yeast 2-hybrid • Biochemical approach – Co-immunoprecipitation – Fusion protein affinity chromatography • Cell-biology – FRET - fluorescence resonance energy transfer • Computational – Rosetta Stone – Co-regulation – Phylogenetic analysis Yeast 2-hybrid approach • Based on the fact some transcriptional activators have separable DNA binding (BD) and transcriptional activation domains (AD). – GAL4, LexA • Protein you are studying = Bait – Fused to the DNA binding domain of GAL4 • Protein(s) you are screening = Fish or Prey – Fused to the activation domain of GAL4 • Transform Bait and Fish plasmids into yeast, measure the expression of a reporter gene. – Usually a gene can be selected for when expressed. Image from: http://www.bioteach.ubc.ca/MolecularBiology/AYeastTwoHybridAssay/ Yeast 2-hybrid on a genome wide scale • Clone every gene in your genome into both the “bait” and “fish” vectors. • Systematically screen each gene for interactions. – Mate individual yeast strains. • Many false positives. Interactome Term to define all of the protein interactions that take place in the cell. Book example - predicting human interactions. Based on data that only 10% of the measured interactions are physiological Yeast 2-hybrid • False-positives – Some baits are “sticky” leading to non-functional interactions • False negatives – Binding not tight enough to detect interaction – Fusion proteins often do not fold correctly • Works best when comparing two proteins suspected of interacting • Bacterial 2-hybrid systems Co-immunoprecipitation • Using an antibody to isolate and purify a protein from a whole cell lysate. • Normally you will only purify the protein the antibody recognizes. • Any additional proteins that co-purify are candidates for interacting proteins. Hirano et al, 1997 Cell, Vol 89, 511-521, 16 May 1997 Fusion protein affinity chromatography • Express the protein of interest as a fusion protein. – 6-8X His residues – Glutathione S-transferase (GST) – Other “tags” • Bind and purify the protein of interest – Poly His residues will bind Ni2+ – GST will bind glutathione Image from: Sigma-Aldrich Fusion proteins - identifying interactions. • In vivo - express fusion protein in vivo – Purify complexes from the cell • In vitro - overexpress protein in vitro – Bind fusion protein to a column and run whole cell lysate through the column. Identify proteins that “stick” to the fusion protein. Difficulties when using biochemical approaches • Stability of protein:protein interactions. – Many are not stable enough to survive purification. • Is the fusion protein functional? – Many times fusions will not be functional. • Quality of the antibody. – Is it good enough to precipitate enough protein for analysis? Computational methods • Rosetta stone analysis – Search for proteins that are separate in one organism but are fused into one protein in another organism. Computational methods • Co-expression – Genes that are in operons are often functionally linked. (not always true). – Determine if the structure of an operon is conserved, indicating co-expression. – Candidates for interaction. – Not a great method. Phylogenetic analysis • Search for the presence of a protein in all organisms. • Determine the distribution. • Identify other proteins that also show this distribution. • Functionally interact? Physically? PLEX • Protein Link EXplorer. • Uses phylogenetic profiles to predict possible associations.