* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Mechanism of Succinyl
Catalytic triad wikipedia , lookup
Multi-state modeling of biomolecules wikipedia , lookup
Adenosine triphosphate wikipedia , lookup
NADH:ubiquinone oxidoreductase (H+-translocating) wikipedia , lookup
Photosynthetic reaction centre wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Photosynthesis wikipedia , lookup
Proteolysis wikipedia , lookup
Fatty acid metabolism wikipedia , lookup
Fatty acid synthesis wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Biochemistry wikipedia , lookup
Metalloprotein wikipedia , lookup
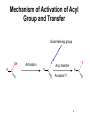
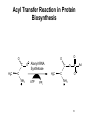
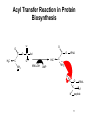
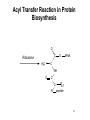
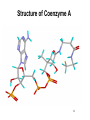
King Saud University College of Science Department of Biochemistry Disclaimer • The texts, tables and images contained in this course presentation are not my own, they can be found on: – References supplied – Atlases or – The web Part 3 Coenzymes-Dependent Enzyme Mechanism Professor A. S. Alhomida 1 2 Coenzyme A (CoA or HS-CoA) 3 Biosynthesis of CoA 4 Coenzyme A (CoA or HS-CoA) • How do living systems synthesize the amide bonds found in proteins, or the ester functional groups found in lipids and other natural products? • The general strategy is to make an activated acyl derivative containing a good leaving group, and then to carry out an acyl transfer reaction 5 Mechanism of Activation of Acyl Group and Transfer Good leaving group OH R X Activation R C O Y Acyl transfer R C O Acceptor Y C O 6 Acyl Carrier Protein Thiol group is the point of attachment to the acyl group being transferred, forming a thioester linkage 7 Acyl Transfer Reaction in Protein Biosynthesis • The aminoacyl group of amino acids is activated and transferred during the assembly of the polypeptide chains of proteins by ribosomes • Amino acid activation is carried out by ATPdependent aminoacyltransfer RNA (tRNA) synthetases 8 Acyl Transfer Reaction in Protein Biosynthesis • The aminoacyl-tRNA ester is then bound to the ribosome and the free amine used to form the next amide bond in the sequence of the protein 9 Acyl Transfer Reaction in Protein Biosynthesis O C H3C O O O Alanyl-tRNA Synthetase C C H3C NH3 ATP PPi C O P O NH3 10 Ad Acyl Transfer Reaction in Protein Biosynthesis O O C H3C C O P O C Ad H3C O NH3 tRNA-OH AMP C O tRNA .. NH2 O C O C N tRNA R n-1 peptide 11 Acyl Transfer Reaction in Protein Biosynthesis O O C Ribosome tRNA C H3C NH O C C N R n-1 peptide 12 Structure of Coenzyme A Thiol group is the point of attachment to the acyl group being transferred, forming a thioester linkage 13 Structure of Coenzyme A 14 Functions of Coenzyme A • CoA is well suited to carry out acyl transfer reactions, since thoils are inherently more nucleophilic than alcohols or amines • Thiols are also better leaving groups (pKa 89), which explains why the hydrolysis of thioesters under basic conditions is more rapid than ester hydrolysis 15 Functions of Coenzyme A • Derived from the vitamin pantothenate (Vit B3) • Participates in acyl-group transfer reactions with carboxylic acids and fatty acids • CoA-dependent reactions include oxidation of fuel molecules and biosynthesis of carboxylic acids and fatty acids • Acyl groups are covalently attached to the SH of CoA to form thioesters 16 Thioester vs Oxyester • The carbonyl carbon atom has more positive charge that the carbonyl in the oxygen esters • To explain this, we should consider the resonance for an oxyester and thioester • The contribution from form II of oxyester tends to decrease the positive charge on the carbon, whereas forms I an II tend to increase the positive charge comparing with forms of thioester 17 Thioester vs Oxyester • Positive charge on carbon will make it easier for a nucleophilic group, such as a carbanion, to attack the carbonyl group • It will also make it easier to remove a proton from the adjacent carbon atom to form a carbanion 18 Resonance O R C O R C .. .. .. ..O .. ..S O R1 R C R 1 R .. II .. .. C I .. ..S O C O R1 I O R1 .. O R .. O R R O .. C S R1 II O R1 + III O R1 C .. R C + .. ..S III 19 R1 Classification of Mechanism of CoA 20 1. Head Activation • This reaction involving attack of nucleophilic groups at the acyl carbonyl carbon atom with transfer of the acyl function to the attacking group and release of CoA • This mechanism is called head activation because the end of acyl function nearest to the CoA becomes attached to the nucleophile 21 Head Activation Mechanism O R C O S CoA R C S Nu + S .. Nu 22 CoA Examples for Head Activation Mechanism • • • • Nu = phosphate: succinly-CoA synthetase Nu = Amine: glucosamine acyl transferase Nu = Water: acetyl-CoA hydrolase Nu = Alcohol: glycerophosphate acetyltransferase • Nu = Thiol: lipoate transferase • Nu = Hydride: acyl-CoA reductase • Nu = Carbanion: b-ketothiolase 23 2. Tail Activation • This is reaction involving condensation of the alkyl carbon of the acyl-CoA by the alkyl carbon by formation of its carbanion • It is called tail activation because the target group is attached to the acyl function by the end furthest from the CoA 24 2. Tail Activation • This is reaction involving condensation of the alkyl carbon of the acyl-CoA by the alkyl carbon by formation of its carbanion • It is called tail activation because the target group is attached to the acyl function by the end furthest from the CoA 25 Tail Activation Mechanism O O O C OH O C CH3CH C S CoA O .. CH3CH C S CoA 26 Tail Activation Mechanism • The carbanion on the a-C of the propionlyCoA attacks the bicarbonate to make methylmalonyl-CoA • The facile character of this reaction is attributed to the increased acidity of the thioester compared to the oxyester • Thioester is 100 – 1000 times more acid which means that it has a much greater tendency to undergo proton dissociation at the methylene function immediately adjacent to the sulfur 27 Tail Activation Mechanism • Negative charge that is produced by this dissociation is stabilized by delocalization over the carbonyl group and by the polarizability of the sulfur • Example: Citrate synthetase 28 3. Siamese Twin Reaction • Two molecules of acyl-CoA react together • One acyl-CoA undergoes head activation and other undergoes tail activation 29 4. Addition Reaction • Reactions involving additions to CoA group • Example: Enoyl-CoA hydratase 30 5. Acyl Group Interchage Reaction • Reactions involving acyl group interchage • Example: Acetoacetyl-CoA transferase 31 Mechanism of Succinyl-CoA Synthetase (Succinyl Thiokinase ) (Head Activation Mechanism) 32 Reaction of Succinyl-CoA Synthetase + DG = - 2.9 kJ/mol 33 Reaction of Succinyl-CoA Synthetase • The conversion of succinyl-CoA to succinate by succinyl CoA synthetase involves use of the high-energy thioester of succinyl-CoA to drive synthesis of a high-energy nucleotide phosphate, by a process known as substrate-level phosphorylation • A high energy succinyl-phosphate intermediate is formed, with the phosphate subsequently being transferred to GDP 34 Reaction of Succinyl-CoA Synthetase • Mitochondrial GTP is used in a transphosphorylation reaction catalyzed by the mitochondrial enzyme nucleoside diphospho kinase to phosphorylate ADP, producing ATP and regenerating GDP for the continued operation of succinyl CoA synthetase • Enzyme has two isoforms in mammals, one which specifically uses ATP for synthesis, and one which utilizes GTP 35 Reaction of Succinyl-CoA Synthetase • Succinyl-CoA is a high energy thioester compound • The D G = - 35.5 kJ/mol for the hydrolysis of succinly-CoA is comparable to that of ATP (30.5 kJ/mol) • How does the enzyme couple the exergonic cleavage of succinly-CoA to the endergonic formation of GTP? 36 Mechanism of Energy Coupling 37 Experimental Evidence for Mechanism of Energy Coupling • By using isotopically labeled ADP • In the absence of succinly-CoA, the enzyme catalyzes the transfer of the g-phosphoryl group from ATP to [14C]ADP, producing [14C]ATP • This isotope-exchange reaction suggests the participation of a phosphoryl-enzyme intermediate that mediates the reaction sequence 38 Experimental Evidence for Mechanism of Energy Coupling 39 Experimental Evidence for Mechanism of Energy Coupling • The enzyme catalyzes two different steps; the favorable oxidation and unfavorable phosphorylation reactions are coupled by the phosphoryl-enzme intermediate • The formation led to the isolation of a kinetically active phosphoryl-enzme in which the phosphoryl group is covalently bound to the N3 position of a His residue 40 Structure of Succinyl-CoA Synthetase • The enzyme is an a2b2 heterodimer; the functional units is one ab pair 41 Mechanism of Succinyl-CoA Synthetase (Head Activation Mechanism) Head activation SCoA C O O P OH Pi O His O His (C H2 )2 O COO SCoA OH H O C O (CH2 ) 2 COO N Succinyl-CoA O P N H N N Tetrahedral intermediate It is the displacement of CoA by Pi which generates another high energy compound, succinly-phosphate (phosphoester) 42 BH Mechanism of Succinyl-CoA Synthetase (Head Activation Mechanism) O O P CoASH O O OH SuccinlylPhosphat C O His C O P (CH2 ) 2 COO His O O N N (CH2 ) 2 OH N BH COO NH B: phosphohistidine His removes the phosphoryl group with the concomitant generation of succinate and phosphohistidine 43 Mechanism of Succinyl-CoA Synthetase (Head Activation Mechanism) His BH O O P O GDP GM O P COO CH2 OH O N N phosphohistidine OH GDP CH2 COO Succinate 44 Mechanism of Succinyl-CoA Synthetase (Head Activation Mechanism) His GTP H N N 45 Mechanism of Citrate Synthtase (Tail Activation Mechanism) 46 • Note that the enzyme catalyst enables the coupling of two chemically independent reactions: – The aldol condensation (with D G = zero) to the very favorable hydrolysis of the CoA thiol ester bond which drives the overall reaction far towards product – Unfortunately the resulting citrate is a tertiary alcohol which cannot be readily oxidized Citrate OAA Aldol condensation Hydrolysis of thioester 47 • • • • Citrate Synthase is an enzyme that catalyzes the first step in the citric acid cycle. Oxaloacetate and acetyl-CoA bind to Citrate Synthase, which then catalyzes the reaction which joins the two compounds together In eukaryotes, Citrate Synthase is a dimer, meaning that it is a protein which is composed of two separate amino acid chains which are not covalently bonded to each other The conformation for Citrate Synthase changes when oxaloacetate (OAA) binds to citrate synthase. This conformational change creates the acetyl-CoA binding site. Once OAA is bound, the binding constant for acetyl-CoA is increased by a factor of 20. To see OAA, click After OAA is bound to the enzyme, acetyl-CoA binds. The enzyme then catalyses the following reactions: enolization of the acetyl-CoA by removing the methyl group, a Claisen condensation which joins the enolated acetyl-CoA and the oxaloacetate creating citryl-CoA, then a thioester hydrolysis which creates citrate and CoA. The monomer of Citrate Synthase, pictured in the lower frame of the left side of this screen shows the citrate synthase enzyme bound to the two products citrate (click 48 49 50 51 Mechanism of Citrate Synthtase (Tail Mechanism) SCoA SCoA C O C O His H C H OAA H Acetyl-CoA N B: N H His CH H carbanion H N COO C N O BH CH2 H 2O COO OAA H O Acetyl-CoA His O H SCoA N C HO C B: N H CH2 COO CH2 COO CoASH His SCoA O C OH N N H B: COO H CH2 C COO HO CH2 COO Citrate CH2 HO C His H N N BH COO CH2 COO Tertahedral Intermediate 52 Mechanism of Thiolase (Siamese Twin Mechanism) SCoA SCoA C H Acetyl-CoA O C O His H C H Acetyl-CoA carbanion H N SCoA N N H His C H2 B: C N O BH CH3 His SCoA O H C N N BH CH2 O C CH3 SCoA SCoA Tertahedral Intermediate O C CH2 + CoASH O C CH3 SCoA Acetoaceyl-CoA His N N H B: 53 Mechanism of thiolase (Acyl Group Interchange Mechanism) SCoA O C His CoASH AcAc-CoA CoASH C N CoAS C H2 His H N O CH3 N H B: N AcAc-CoA SCoA C O His C H2 O C H N SCoA CH3 Tertahedral Intermediate N BH SCoA 2 C O CH3 Acetyl-CoA His N N H B: 54 Thiamine Pyrophosphate (TPP) • TPP is a derivative of thiamine (Vit B1) • Reactive center is the thiazolium ring (with a very acidic hydrogen atom at C-2 position) • TPP participates in reactions of: (1) Decarboxylation (2) Oxidative decarboxylation (3) Transketolase enzyme reactions 55 Thiamine (Vitamin B1) and TPP 56 Mechanism of Pyruvate Dehydrogenase Complex B: R1 CH3 CH3 N C H Enz 1 S R2 R1 CH3 N S R2 TPP C CH3 C O C O R1 BH C S R2 TPP O Pyruvate CH3 CH3 N C OH C O O C O Pyr DH S SH CO2 Enz Acetyl-lipoic acid CH3 Dihydrolipoyltransacetylase S R2 B: CH3 CH3 R1 CH3 N O S S C OH HETPP S S BH Enz Lipoic Acid SH ACetyl-CoA HS C H C C O R2 SCoA CH3 N CoASH C R1 Enz SH Dihydrolipoyl DH Enz FAD FADH2 NAD NADH + H 57 Mechanism of Transketolase B: R1 CH3 CH3 N S R2 C H Enz 1 R1 CH2 OH N C S TPP R2 C O S R2 TPP CH2 OH N BH OH C H H C R1 CH3 C OH C OH C H OH H C CH2O P OH CH2O P Xylulose-5 P O H C H C CH2 OH C OH C (H C OH CH2O P O R1 CH3 H CH2OH N OH )3 S R2 C C OH HETPP CH2O P Sedoheptulose-7 P O B: CH3 R1 N S R2 CH2 OH H C C O OH C (H C H OH )3 BH H C (H C OH )3 CH2O P Ribose-5 P CH2O P 58 Pyridoxal Phosphate (PLP) • PLP is derived from Vit B6 family of vitamins (deficiencies lead to dermatitis and disorders of protein metabolism) • Vitamin B6 is phosphorylated to form PLP • PLP is a prosthetic group for enzymes catalyzing reactions involving amino acid metabolism (isomerizations, decarboxylations, side chain eliminations or replacements) 59 B6 Vitamins and pyridoxal phosphate (PLP) 60 PLP Reactions HN O P C COO CH CH2 O H O O OH O H .. O N CH3 CH2 O C COO + NH3 + PLP Transamination H O CH3 C HN COO + NH3 O + Racem ization a P COO OH O O PLP C CH CH2 O CH3 N H a-b-elimination H ab HN C COO c O CH CH2 CH2 HO O H3N C COO O + OH O P HN H a O O CH3 N PLP P C COO CH CH2 O O H OH O .. O N H CH3 H PLP-Ala Schiff H HN + O P O decarboxylation CH2OH b c H deformylation HN CH O CO2 C O .. N H H HOCH2CHO CH3 + + CH2O + O P COO CH O OH C O O OH .. N PLP CH3 H + NH3 Glycine + PLP 61 Mechanism of Amino Acid Transaminase-1 Lys-258 Ala H2O Lys :N O ENZ OH O P O OH O N H CH3 N O P H O CH O C H O O NH2 H2 N C H H PLP CH3 H COO PLP-Enz Schiff (Aldimine) CH3 Ala B: : NH Lys 2 H HN O OH O P O P H HC H O PLP-Ala Schiff OH O O N H2 CH3 N CH3 N NH2 O COO CH CH3 O BH C Lys H Pyridoxamine (PMP) O H3C C COO H3C N H3 NH CH O OAA O P Lys COO C Pyr OH O .. O CH3 N H Quinonoid B: H3C BH P O H C COO :N Lys H 2O H2 NH H CH O O O H3C OH O N C :N H2 HC H O H O P O Lys COO NH CH3 O OH O H N H CH3 H B: Ketimine Ketimine 62 Mechanism of Amino Acid Transaminase-2 COO NH2 H CH O O P O O C :N Lys COO H2 CH2 CH2 O COO N CH3 OAA BH H 2O PMP O :N H2 NH HCH O H Lys COO C OH OH O P O CH3 N H PLP-OAA Schiff (Aldimine) COO CH2 H C N H3 COO Lys NH CH O O P B: OH O O COO CH2 CH3 N H2N H C COO H Asp H N CH O O P O Lys OH O N CH3 BH H PLP-Enz Schiff base (Aldimine) 63 Biotin • Biotin is required in very small amounts because it is available from intestinal bacteria • Avidin (raw egg protein) binds biotin very tightly and may lead to a biotin deficiency (cooking eggs denatures avidin so it does not bind biotin) • Biotin (a prosthetic group) enzymes catalyze: (1) Carboxyl-group transfer reactions (2) ATP-dependent carboxylation reactions 64 Enzyme-bound biotin • Biotin is linked by an amide bond to the e-amino group of a lysine residue of the enzyme • The reactive center of biotin is the N-1 (red) 65 Reaction catalyzed by pyruvate carboxylase Two step mechanism (next slide) Step 1: Formation of carboxybiotin-enzyme complex (requires ATP) Step 2: Enolate form of pyruvate attacks the carboxyl group of carboxybiotin forming oxaloacetate and regenerating biotin 66 Mechanism of PEP Carboxylase (ATP-Independent Biotin Carboxylase) O O CH2 O + HO C C O COO Bicarbonate P O B: OH R S Biotin-Enz HN PEP N CH2 C H H O B: HN N O O O C O P CH2 HN N O BH R S S O O P Carboxyl-phosphate OH COO R O O C O H O C O O H COO Pyruvate enolate Biotin-Enz O O O O P Pi O O BH O R S C HN BH R S OH N HN CH2 O C O N O C OH O C O COO COO OAA CH2 Carboxyl-biotin-Enz Pyruvate enolate COO 67 Mechanism of Pyr Carboxylase (ATP-Dependent Biotin Carboxylase) O O B: Biotin-Enz O + HO C AD O P O Bicarbonate OH R S Pyruvate HN N H H B: ATP CH2 O O O C O P O C H O O Carboxyl-phosphate OH COO Pyruvate CH2 C O BH R S COO S HN B: R N HN H N O O O C O P O H CH2 O O O Biotin-Enz C O COO Pyruvate enolate O O P BH O R S BH R S C HN O O Pi OH N HN CH2 O C O N O C OH O C O COO COO OAA CH2 Carboxyl-biotin-Enz Pyruvate enolate COO 68 Folic acid 69 Tetrahydrofolate (THF) • Vitamin folate is found in green leaves, liver, yeast • The coenzyme THF is a folate derivative where positions 5,6,7,8 of the pterin ring are reduced • THF contains 5-6 glutamate residues which facilitate binding of the coenzyme to enzymes • THF participates in transfers of one carbon units at the oxidation levels of methanol (CH3OH), formaldehyde (HCHO), formic acid (HCOOH) 70 Tetrahydrofolate is an important cofactor in nitrogen metabolism Biotin transfers carbon in its most oxidized state - carbon dioxide SAM (S-adenosylmethionine) can transfer carbon in its most reduced state methyl groups (but the methyl group comes from 5-methyl-tetrahydrofolate) Tetrahydrofolate transfers one-carbon groups in intermediate oxidation states and sometimes as methyl groups 71 Note the differences between folate and tetrahydrofolate Close up views of how the different oxidation states of carbon are carried by folate 72 Different Forms of Folate Note: these are positions 5,6,7,9 and 10 73 Pterin, folate and tetrahydrofolate (THF) 74 Formation of tetrahydrofolate (THF) from folate 75 • One-carbon derivatives of THF Continued next slide 76 77 THF, Vitamin B12 and SAM His Sources of carbon (1 - 5) Epinephrine 2 Gly Formimino-Glu CO2 + NH4+ 3 4 Glu + NH4+ Glucose Formaldehyde Gly 1 Formate 5 Ser THF DHF + NADP NADPH THF-C dTMP A dUMP B12- CH3 Purine precursors B12 Homocysteine D SAH CH3 Trp B Ser C Gly Purines (C2 and C8) Meth SAM Recipients of carbon (A - D) Norepinephrine Epinephrine Guanidinoacetate Creatine Nucleotides Methyated nucleotides Phosphatidylethanolamine Phosphatidylcholine Acetylserotonin Melatonin 78 Metabolic reactions involving synthesis, interconversions and utilization of single carbon adducts of tetrahydrofolate Fig. 20.17 Note following reactions: 9 methionine synthase 10, 11 = serine and glycine catabolism 79 12 thymidylate synthase Mechanism of Thymidate Synthase BH+ H N H2N N N Cys CH2 N H R Cys O H R R H NH O OH O P NH OH O P N H2C O S H B: CH2 N H Cys R N H2C N N R CH2 N O S N5, N10-methylene-THF H N R N N H2N N H O HC 2 S H H N H2N .. OCH2 N O O O OCH2 N O O H H H OH H H O H H H OH H H BH+ N H2N Cys H N S CH2 N O OCH2 H H NH N O O H H OH H R O N H2N Cys H R N B: OH H N H2C O H O P dUMP B: H N N S N O BH+ R S R N H H O H H2C NH OH O P Cys H CH2 OCH2 N O O O H H H OH H H H N H2N H H N N CH2 N O N R R H 7,8-DHF + O H3C NH OH O P OCH2 N O O O H H H OH H H dTMP 80 Mechanism of Thymidate Synthase Inhibition BH+ H N H2N N .. N Cys CH2 N H S R Cys R N O Cys R H NH O OH F O P R F H NH OH N H2C O S H B: CH2 N H R N H2C N N R CH2 N O S N5, N10-methylene-THF H N N H2N N H O HC 2 H H N H2N O P OCH2 N O O O OCH2 N O O H H H OH H H O H BH+ H H OH H H 5-F-dUMP H N H2N Cys H N S N CH2 N O F N H2C O B: H NH OH O P R R OCH2 N O O O H H H OH H H 81 Mechanism of Inhibition of Biosynthesis of dTMP O H3C H3N C H H H N H2N COO N R N N H CH2OH O CH2NH N H O P OCH2 + N O O O CH2NH O H NH OH N R N NADP Serine N HN DHF Reductase 2 H H NADPH THF H OH Thymidylate synthetase 5-F-dUMP H dTMP H Methotrexate O H H3N C H N H2N COO N N H H2 O HC 2 Glycine NH OH N CH2 N O P OCH2 R N O O O R H N5, N10-methylene-THF H H OH H H dUMP O F NH OH O P OCH2 N O N O COO O H H OH H H H 5-Fluoro-2'-deoxyuridylate monophosphate (5-F-dUMP) OOCCH2CH2CHNHC O NCH2 N CH3 N NH2 N NH2 Methotrexate 82 Relationship between THF, Vit B12 and SAM ATP Methionine 3P THF-CH3 THF B12 S-adenosylmethionine ( SAM ) Precursor CH3 - product B12 -CH3 S-adenosylhomocysteine ( SAH ) Homocysteine Adenosine 83 Mechanism of Serine Hydroxymethyltransferase-1 Lys NH O H H B: OH O P O N H N C H CH O HO H2C CH3 C COO NH H COO Lys :HN2 CH O O CH2 BH P OH O O OH Ser CH3 N H PLP-Sub Schiff (Aldimine) BH OH H 2C C COO N H3 NH CH O O P Lys OH O .. O N CH3 H Next Page Quinonoid THF H2O H N H2N N N O CH2NH C H2 C COO P O O .. N N H2N CH3 H PLP-THF Schiff (Aldimine) N C COO R N N THF O CH2NH H O P O N H3 NH CH O B: OH C H H HC O O Lys N H3 BH NH H H R N H OH O N CH3 H Ketimine 84 Mechanism of Serine Hydroxymethyltransferase-2 H N H2N N R N N H O CH2NH B: HC H H C COO P P H O N N H2N OH O H OH .. N CH3 N N CH2 C N H2 R O 5 O H N CH3 H O Lys N H3 CH O HC O C COO NH NH O H BH Lys N H3 Quinonoid H2O N, 10N-methylene-THF Lys O P O H C H BH C H O O COO N H3 OH O CH O N Lys N H3 NH CH3 O H P O PLP OH O N H O H H CH3 B: COO H C H NH2 Gly 85 Folic acid deficincies Early 1990s: Epidemiological studies demonstrated correlations between folate deficiencies and increased risk of myocardial infarctions heart attacks These same individuals also had elevated levels of homocysteine Working hypothesis: homocysteine accumulates in folate deficient individuals because of a decrease in the ability of the methionine synthase reaction to function (due to lack of THF) Homocysteine causes heart damage by an unknown mechanism Folate deficiencies during embryogenesis cause a significant proportion of neural tube defects and consequent failure of the nervous system to develop properly This is most likely due to inability to synthesize adequate amounts of thymine nucleotides 86 HOMOCYSTINURIA AND HEART DISEASE some patients with arteriosclerosis have elevated homocysteine low plasma B vitamins and low dietary intake show increased risk for heart disease folate/B12 deficiency common in high risk populations (elderly, smokers) B vitamins decrease plasma homocysteine homocysteine may oxidatively damage lipoproteins and endothelia of vessel walls controversy remains 87 3 S-Adenosylmethionine Homocysteine-SH methyl-B12 THF 4 CYTOPLASM Methionine synthase ATP Methionine-SCH3 ↑ in methyl trap with B12 deficiency N5-methyl THF 1 Hydroxy B12 X From diet 2 Adenosyl B12 Succinyl CoA Methylmalonyl CoA mutase Methylmalonyl CoA increase Propionyl CoA carboxylase TCA cycle biotin MITOCHONDRION Propionyl CoA Figure 5. Metabolism of cobalamin and associated disorders (homocystinuria and methylmalonic aciduria) 88 Homocysteine Homocysteinuria • Rare; deficiency of cystathionine b-synthase • Dislocated optical lenses • Mental retardation • Osteoporosis • Cardiovascular disease death High blood levels of homocysteine associated with cardiovascular disease • May be related to dietary folate deficiency • Folate enhances conversion of homocysteine to methionine 89 Dihydrofolate reductase converts folate to tetrahydrofolate by two successive reductions Dihydrofolate reductase is the target of a number of clinically important drugs called antimetabolites: synthetic compounds that are structural analogs of a normal metabolite sulfa drugs in bacterial infections Anticancer agents 90 Aminopterin Tetrahydrofolate in the metaboblism of one-carbon units Single carbon groups can be carried on N-5, N-10, or bridged between N-5 and N-10 methyl Carbon units are obtained from a variety of sources BUT most activated single carbon units are obtained from the beta carbon of serine Once a single carbon unit has been activated by attachment to tetrahydrofolate it can be used directly in a biosynthetic reaction or it can undergo interconversions to different oxidation states methylene formyl 91 Cobalamin (Vitamin B12) • Coenzymes: methylcobalamin, adenosylcobalamin • Cobalamin contains a corrin ring system and a cobalt (it is synthesized by only a few microorganisms) • Humans obtain cobalamin from foods of animal origin (deficiency leads to pernicious anemia) • Coenzymes participate in enzyme-catalyzed molecular rearrangements in which an H atom and a second group on the substrate exchange places 92 Cobalamin (Vit B12) and its coenzymes (a) Cobalamin. Corrin ring (black) 93 (b) Abbreviated structure of cobalamin coenzymes 94 Intramolecular rearrangements catalyzed by adenosylcobalamin enzymes (a) Rearrangement of an H and substituent X on an adjacent carbon 95 (b) Rearrangement of methylmalonyl CoA 96 Methylcobalamin participates in the transfer of methyl groups 97 Methyl-Vitamin B12 OH OH CH2 O Amenine Con+ DMB 5'-Adenosyl-B12 (5'-Adenosylcobalamine) CH3 Con+ DMB Methyl-B12 (methylcobalamin) CH3 Con+ DMB Methyl B12 as originally isolated (cyanocobalamin) n = 1, 2 or 3 98 Mechanism of Methymalonyl-CoA Mutase H 1. 2. Homolytic cleavage of the CCo2+ bond Ado H . Co2+ Abstraction of hydrogen atom from methylmalonyl-CoA COO CH H C Ado O S H C + H C H CoA + H C H + . Co2+ COO . CH C H Methyl-malonyl-S-CoA H C O S CoA . 2 DMB Rearrangement DMB 3 1 Ado H Co2+ . H C Ado + H C H + H H COO C CH H 3. Carbon skeleton rearrangement CoA 5'-Adenosyl-B12 (5'-Adenosylcobalamine) Abstraction of hydrogen atom from 5`-deoxyadenosine . Ado + H H C + H H H COO C . CH C O S CoA DMB Hypothetical intermediates 5 4 Release of succinly-CoA and reformation of Vit B12 S Cyclopropyloxy radical H 5. O DMB Co2+ 4. . C DMB Co3+ H C C S COO CH O H CoA Co2+ Succinyl-CoA . Ado + H . C H + H H COO C CH C O H S CoA DMB 99 Lipoamide • Coenzyme lipoamide is the protein-bound form of lipoic acid • Animals can synthesize lipoic acid, it is not a vitamin • Lipoic acid is an 8-carbon carboxylic acid with sulfhydryl groups on C-6 and C-8 • Lipoamide functions as a “swinging arm” that carries acyl groups between active sites in multienzyme complexes 100 Lipoamide • Lipoic acid is bound via an amide linkage to the e-amino group of an enzyme lysine • Reactive center of the coenzyme shown in red 101 Transfer of an acyl group between active sites • Acetyl groups attached to the C-8 of lipoamide can be transferred to acceptor molecules • In the pyruvate dehydrogenase reaction the acetyl group is transferred to coenzyme A to form acetylSCoA 102 Lipid Vitamins • Four lipid vitamins: A, D, E, K • All contain rings and long, aliphatic side chains • All are highly hydrophobic • The lipid vitamins differ widely in their functions 103 Vitamin A (Retinol) • Vit A is obtained from liver, egg yolks, milk products or b-carotene from yellow vegetables • Vit A exists in 3 forms: alcohol (retinol), aldehyde and retinoic acid • Retinol and retinoic acid have roles as protein receptors • Rentinal (aldehyde) is a light-sensitive compound with a role in vision 104 Formation of vitamin A from b-carotene 105 Vitamin D • A group of related lipids involved in control of Ca2+ utilization in humans • Vitamin D3 and 1,25-dihydroxycholecalciferol 106 Vitamin E (a-tocopherol) • A reducing reagent that scavenges oxygen and free radicals • May prevent damage to fatty acids in membranes Fig 7.29 Vitamin E (a-tocopherol) 107 Vitamin K (phylloquinone) • Required for synthesis of blood coagulation proteins • A coenzyme for mammalian carboxylases that convert glutamate to g-carboxyglutamate residues • Calcium binds to the g-carboxyGlu residues of these coagulation proteins which adhere to platelet surfaces • Vitamin K analogs (used as competitive inhibitors to prevent regeneration of dihydrovitamin K) are given to individuals who suffer excessive blood clotting 108 (a) Structure of vitamin K (b) Vit K-dependent carboxylation 109 Ubiquinone (Coenzyme Q) • Found in respiring organisms and photosynthetic bacteria • Transports electrons between membraneembedded complexes • Plastoquinone (ubiquinone analog) functions in photosynthetic electron transport 110 (a) Ubiquinone, (b) Plastoquinone • Hydrophobic tail of each is composed of 6 to 10 five-carbon isoprenoid units • The isoprenoid chain allows these quinones to dissolve in lipid membranes 111 • Three oxidation states of ubiquinone • Ubiquinone is reduced in two one-electron steps via a semiquinone free radical intermediate. Reactive center is shown in red. 112 Protein Coenzymes • Protein coenzymes (group-transfer proteins) contain a functional group as part of a protein or as a prosthetic group • Participate in: (1) Group-transfer reactions (2) Oxidation-reduction reactions where transferred group is a hydrogen or an electron • Metal ions, iron-sulfur clusters and heme groups are commonly found in these proteins 113 Stereo view of oxidized thioredoxin • Cystine group is on the surface (sulfurs in yellow) 114 Cytochromes • Heme-containing coenzymes whose Fe(III) undergoes reversible one-electron reduction • Cytochromes a,b and c have different visible absorption spectra and heme prosthetic groups • Electron transfer potential varies among different cytochromes due to the different protein environment of each prosthetic group 115 (a) Heme group of cyt a 116 (b) Heme group of cyt b 117 (c) Heme group of cyt c 118 Absorption spectra of oxidized and reduced cytochrome c • Reduced cyt c (blue) has 3 absorbance peaks: a,b,g • Oxidized cyt c (red) has only a g (Soret) band 119