* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Quantitative Genetics: Traits controlled my many loci Quantitative
Inbreeding avoidance wikipedia , lookup
Genetics and archaeogenetics of South Asia wikipedia , lookup
Dual inheritance theory wikipedia , lookup
Pharmacogenomics wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
History of genetic engineering wikipedia , lookup
Designer baby wikipedia , lookup
Polymorphism (biology) wikipedia , lookup
Genetic engineering wikipedia , lookup
Dominance (genetics) wikipedia , lookup
Genome (book) wikipedia , lookup
Genetic testing wikipedia , lookup
Selective breeding wikipedia , lookup
Public health genomics wikipedia , lookup
Medical genetics wikipedia , lookup
Genetic drift wikipedia , lookup
Koinophilia wikipedia , lookup
Microevolution wikipedia , lookup
Human genetic variation wikipedia , lookup
Behavioural genetics wikipedia , lookup
Population genetics wikipedia , lookup
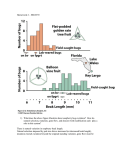
Quantitative Genetics: Traits controlled my many loci Quantitative Genetics I: Traits controlled my many loci So far in our discussions, we have focused on understanding how selection works on a small number of loci (1 or 2). Learning Objectives: 1. To describe how segregation at multiple loci can produce a pattern of quantitative variation in a trait. However in many cases, evolutionary biologists ask questions about traits or phenotypes (for example…) 2. To define the breeding value (A) and relate it to the average effects of alleles. Many factors may affect a trait, including the action of alleles at one or more loci, and the environment in which an individual exists. 3. To define and differentiate broad and narrow sense heritability. 4. To describe the components of trait (phenotypic) variation and describe how and why additive genetic variation is the key component of variation relevant to narrow sense heritability and the response to selection. Quantitative genetics provides the framework for understanding how evolutionary forces act on complex traits. 336-9 Readings: Chapter 9 in Freeman 1 Quantitative genetics vs. population genetics 336-9 2 Sir Ronald Fisher (1890-1962): Linking quantitative traits variation and Mendelian genetics • In 1918, Fisher showed that a large number of Mendelian factors (genes) influencing a trait would cause a nearly continuous distribution of trait values. Therefore, mendelian genetics can lead to an approximately normal distribution 336-9 3 336-9 4 Wheat kernel colour variation 336-9 Wheat kernel colour variation This figure shows measured phenotypes in a population of F2 plants from parents that differ in kernel colour. We can see that more than two or three phenotypes are seen in the F2. This pattern is explained by the action of three loci. 5 Population genetics 336-9 With three loci, each with two alleles, six phenotypic classes are obtained, and the distribution of phenotypes begins to look like a normal curve. 336-9 6 Quantitative genetics 7 What are the conditions that will lead to a shift in the mean value of a trait under selection? 336-9 8 Breeder’s equation Human Height: An example of a quantitative trait 336-9 9 The first big question in quantitative genetics: Breeder’s equation: R = h2S 10 Building a Quantitative Genetics Model: lessons from agriculture • How much phenotypic variation among individuals is due to the presence and interaction of different alleles, and how much is due to differences in the environment? • Answers to this question will determine the degree to which traits can respond to selection. 336-9 336-9 In quantitative genetics, the phenotypic value (P) of an individual (e.g. height) is attributed to the genotype of the individual and to its environment: P = G + E The genotypic value (G) reflects the influence of every gene carried by the individual on the phenotypic value. The environmental deviation (E) is a measure of the influence of the environment of the phenotypic value of an individual. We can see how these components are estimated in an example from crop yield in wheat. 11 336-9 12 Average Yield of three wheat strains over a ten year period (bushels/acre) 336-9 Average Yield of three wheat strains over a ten year period (bushels/acre) Year Roughrider Seward Agassiz Year Roughrider Seward Agassiz 1986 47.9 55.9 47.5 1986 47.9 55.9 47.5 1987 63.8 72.5 59.5 1987 63.8 72.5 59.5 1988 23.1 25.7 28.4 1988 23.1 25.7 28.4 1989 61.6 66.5 60.5 1989 61.6 66.5 60.5 1990 0.0 0.0 0.0 1990 0.0 0.0 0.0 1991 60.3 71.0 55.4 1991 60.3 71.0 55.4 1992 46.6 49.0 41.5 1992 46.6 49.0 41.5 1993 58.2 62.9 48.8 1993 58.2 62.9 48.8 1994 41.7 53.2 39.8 1994 41.7 53.2 39.8 1995 53.1 65.1 53.5 1995 53.1 65.1 53.5 Mean 45.63 52.18 43.49 Mean 45.63 52.18 43.49 13 Using Genetic values in breeding: The Breeding Value Mean yield of population Genetic value of a parent (60 bushels/acre) (80 bushels/acre) Expected genetic value of offspring (70 bushels/acre) The genetic value of a genotype reflects the sum total effect of all alleles at the loci that affect the trait of interest. Given that a parent in a sexual species passes half of its alleles to the offspring, what is the expected genetic value of the offspring? (assume a randomly 336-9 15 chosen mate) 336-9 Environmental values (E) 63.8 - 45.63 = 18.17 49.0 - 52.18 = -3.18 Genetic values (G) 14 Using Genetic values in breeding: The Breeding Value Mean yield of population (60 bushels/acre) Actual genetic value of offspring (67 bushels/acre) Genetic value of a parent (80 bushels/acre) In reality, the yield of the offspring may differ from that predicted on the basis of the genetic value of the parent. Why? - Dominance (interactions among alleles at a locus) - Epistasis (interactions among alleles at different loci) 336-9 16 Using Genetic values in breeding: The Breeding Value d d Mean yield of population (60 bushels/acre) Actual genetic value of offspring (67 bushels/acre) Breeding Value (A) of the parent genotype (74 bushels/acre) The breeding value of a genotype (A) is obtained by adding twice the deviation of the mean (d) of the offspring from the population mean. 336-9 17 Breeding Value Example 1 To increase milk yield, dairy farmers estimate the breeding value of bulls from the average dairy production of each bull’s daughters. When a particular bull is mated to several cows, his daughters produce an average of 100 liters of milk per day, in a herd with an average production of 75 liters. In terms of dairy production, ...what is the breeding value (A) of the bull?(125) ...what is the phenotypic value of the bull? (!!) 336-9 18 Effects of Dominance Breeding Value Example 2 Now say that a particular cow produces 100 liters of milk per day, compared to a herd average of 75 liters per day. When mated to different bulls, this cow’s daughters produce an average of 80 liters of milk per day. In terms of dairy production, ...what is the breeding value (A) of the cow? (85) ...what is the phenotypic value of the cow? (100) What contributes to this difference (assuming no environmental effects)? If alleles at some loci affect traits differently depending on the rest of the genotype (Interactions) Dominance (D) (interactions at the same locus) Epistasis (I) (interactions at different loci) 336-9 19 Dominance relationships among alleles at a locus affect the way in which a trait is transmitted to the offspring. A parent that is homozygous (e.g. BB) at a locus that affects a trait cannot transmit this condition to its offspring. If B is recessive to b, a high fitness BB parent mated to a low fitness bb parent produces only Bb (low fitness) offspring. Such dominance effects have an impact on trait expression of the offspring from any cross. 336-9 20 Effects of Gene Interactions /Epistasis Average Allele Effect Similarly, good interaction among the alleles at different loci are not faithfully transmitted, as illustrated in these card hands. Even though Mom and Dad have good combinations, they may not combine well in the offspring. Because of dominance and epistasis, a given allele may not always have the same effect of the phenotype. 5 6 7 8 9 6 4 " " " " " ! " Mom 336-9 A A # $ A " Dad 6 4 7 A 9 ! " " $ " Offspring The average effect of an allele accounts for the difference in the effect of an allele paired with any other alleles /genes currently found within the population (e.g., accounting for the chance that it is found in a heterozygote or homozygote, in any particular genetic background). The breeding value of an individual (A) represents the average effects of all of his/her alleles. 21 Expanding the basic quantitative genetics equation We earlier described the relationship, 336-9 From individuals to populations: patterns of phenotypic variation With an understanding of factors that determine the phenotype of an individual, we can move back up to the level of the population to develop our understanding of how to estimate the genetic component of quantitative trait variation. P = G + E, Which describe the factors that determine an individual’s phenotype, but we now understand that the component G can be further broken down into: G= A + D + I, to describe the components of Genetic effects on the phenotype attributed to Additive genetic effects (as measured by he Breeding value), Dominance effects and Interaction effects (Epistasis). Our description of the Breeding value (A) showed that the phenotype of an individual’s offspring is mainly determined by the 336-9 23 breeding value of its parents. 22 Q: How much of the phenotypic variation that we observe is due to genetic variation? How much of this genetic variation contributes to the response to selection? 336-9 24 From individuals to populations: patterns of phenotypic variation From individuals to populations: patterns of phenotypic variation VP = VG + VE The phenotypic variance (VP) measures the extent to which individuals vary in phenotype for a particular trait. The genetic variance (VG) can be further broken down into additive, dominance and interaction components, analogous to those used to describe an individual phenotype: The phenotypic variance within a population may be due to genetic and/or environmental differences among individuals: VP = VG + VE VG = VA + VD + VI The additive genetic variance (VA) equals the variance in breeding values within a population and measures the degree to which offspring resemble their parents.26 (Ignoring interactions between genes & environment) 336-9 25 Calculating phenotypic and additive genetic variances Calculating phenotypic and additive genetic variances Example: Milk yield in cows Example: Milk yield in cows Variance: _ Vx= !i (Xi – X)2 --------------(n – 1) 336-9 336-9 Cow yield 1 75 2 88 3 52 4 83 5 82 6 43 7 100 8 48 9 79 10 100 mean 75 Variance: VP = (75-75)2 + (88-75)2 +… n-1 = 425.6 27 _ Vx= !i (Xi – X)2 --------------(n – 1) 336-9 Cow yield Offspring Yield 1 75 2 88 81.5 3 52 65.5 4 83 79 5 82 78.5 6 43 66 7 100 84 8 48 64.5 9 79 79.5 10 100 78 mean 75 A = (offspringmean) X 2 + mean 74.5 A = 83 A = 93 28 Calculating Phenotypic and Additive Genetic Variance Calculating Phenotypic and Additive Genetic Variance Example: Milk yield in cows Variance: _ Vx= !i (Xi – X)2 --------------(n – 1) 336-9 Cow yield Offspring Yield A 1 75 74.5 74 2 3 88 81.5 88 52 65.5 56 4 5 83 79 83 82 78.5 82 6 43 66 57 7 100 84 93 8 48 64.5 54 9 79 79.5 84 10 100 78 mean 75 VP = 425.6 VA = 205.5 81 75.2 29 Knowledge of the phenotypic variance and additive genetic variance allows us to predict how similar we expect the phenotypes of parents and offspring to be. It will also allow us to predict the magnitude of the response to selection when individuals with different phenotypic means have 336-9 different probabilities of survival or reproduction. Narrow-Sense Heritability h2 = VA / VP In the previous example, nearly half of the phenotypic variance was the result of additive genetic variance. 30 Broad sense heritability describes the proportion of phenotypic variance due to total genetic variance among individuals. H2 = VG / VP Broad-sense heritability will be 1 if all of the phenotypic variation within a population is due to genotypic differences among individuals (VG = VP). Broad-sense heritability will be 0 if all of the phenotypic variation is caused by environmental differences. h2 = VA / (VA + VD + VI + VE) Heritability can be low due to: 336-9 VA = 205.5 Broad-Sense Heritability H2 = VG / VP Narrow sense heritability describes the proportion of phenotypic variance due to additive genetic variance among individuals, or the extent to which variation in phenotype is caused by genes transmitted from parents. Conversely, h2 will be 1 only if there is no variation due to dominance, epistasis, or environmental effects. When h2 = 1, P = G = A. VP = 425.6 31 336-9 32 Important points about heritability 1. When we use the term ‘heritability’, we are almost always referring to narrow sense heritability. 2. Estimates of heritability are not transferable. They are specific to the population and the environment in which they are estimated. 3. Heritability estimates are for populations, not individuals 4. Heritability does not indicate the degree to which a trait is genetically based. Rather, it measures the proportion of the phenotypic variance that is the result of genetic factors. 336-9 33