* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Manual
SNARE (protein) wikipedia , lookup
Cyclic nucleotide–gated ion channel wikipedia , lookup
Membrane potential wikipedia , lookup
Implicit solvation wikipedia , lookup
List of types of proteins wikipedia , lookup
Endomembrane system wikipedia , lookup
Cell membrane wikipedia , lookup
Theories of general anaesthetic action wikipedia , lookup
Mechanosensitive channels wikipedia , lookup
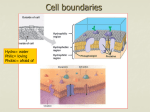
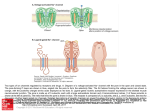
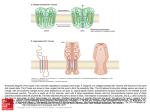
Investigation of lipid bilayers with naturally occuring protein pores and biomimmetic DNA channels Swati Krishnan and Vera Arnaut Venue: ZNN, 2nd oor October 19, 2016 Contents 0.1 Introduction . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 0.1.1 Lipid bilayer . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 0.1.2 Elemental properties of pores . . . . . . . . . . . . . . . . . . . . . . . . . . 0.1.3 Protein channels and functions . . . . . . . . . . . . . . . . . . . . . . . . . 0.1.4 Biomimetic DNA channels . . . . . . . . . . . . . . . . . . . . . . . . . . . 0.2 Experimental protocol . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 0.2.1 Over view . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 0.2.2 Painted bilayer setup and electrical characterization of channels . . . . . . . . 0.2.3 Giant unilamellar vesicles - Inverted emulsion technique . . . . . . . . . . . . 0.2.4 Modication of the DNA pores with Fluorescentlylabeled staples for imaging . 0.2.5 Fluorescence based detection of hydrophobic interactions between DNA and lipid membranes . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 0.3 Investigation . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 2 2 2 4 5 7 7 7 8 8 9 9 1 0.1 Introduction 0.1.1 Lipid bilayer Cell membranes are essential boundaries encapsulating the cellular machinery with a lipid bilayer and several embedded proteins to allow controlled communication with the external environment. A lipid molecule is an amphiphile with a hydrophillic head and hydrophobic fatty acid tail. In a bilayer the lipids are arranged in such a way that the hydrophobic tails of both the layers interact with each other. The edges of the bilayer are not in a favourable state as at the edges the tails are exposed to the surrounding hydrophillic environment. Based on the geometry of the lipid, they tend to form vesicles or micelles. Model membrane systems are made invitro to study membrane processes and for biotechnological Figure 0.1: Dierent lipid structures applications. One of the earliest methods to make model lipid bilayers invitro is the production of painted bilayers. The lipids are dissolved in an organic solvent like decane and painted across a teon aperture separating two buer reservoirs (Figure 0.2). The orbit mini system used in our studies is a modication of the painted bilayer technique. A 2x2 microarray cavity chip on an inert polymer is used to paint the bilayers with a small brush. Each cavity has a set of electrodes which enables current recordings. The cavities act as the teon aperture in the classical painted bilayer system. 0.1.2 Elemental properties of pores Channels or pores allow the movement of molecules across an impermeable membrane. The dimensions of the pore dictate the number of molecules it will transport across the membrane in a given time. Nature of the channel like its charge, presence of receptors etc. dictate selectivity of the ions passing through the pore. The molecules which are smaller than the pore sheds its hydration shell which is usually compensated by interactions between the pore walls and the molecule. The conductance of a pore is given by : γ = A/ρL (0.1) 2 Figure 0.2: Left:MECA chip with 4 cavities; Right: Painted bilayer technique. where ρ is the resistivity of the solution, L is the length of the pore and A is the area of the pore. For a pore with a dened length and area in a certain solution and small test potential equation (0.1) provides the (maximum) limiting conditions. It can be deduced from the equation that longer channels are less conductive than shorter counterparts. Once a potential eld is applied, the conductance of the ion channel can be measured using : γ = I/V (0.2) Consider the following trace The trace is obtained by measuring passage of ions or current across the Figure 0.3: Example current trace bilayer when protein pores are added to it. The otherwise impermeable membrane now shows a small step wise increases in its conductance. Each step increase is viewed as an addition of a pore to the membrane and each decrease is the removal of a pore. The length of the step is the mean life time of the pore in the membrane. Note that at time zero the bilayer already shows conductance which indicates?. The smaller peaks which are more transient are due to uctuations in the membrane or pore structure, often called gating events. How can we nd the conductance of a single pore from this trace? We know from equation (0.2) that γ is I/V. The smallest sustainable jump or the unitary 3 conductance step in the trace will give us an indication of I or current across a single pore. Since we apply the potential across the bilayer we also have the V value. Note the subsequent higher jumps in the current trace would be approximately a multiple of the smallest step indicating simultaneous insertion of 2,3 or more pores. 0.1.3 Protein channels and functions Gramicidin An ideal model pore would have a precisely dened structure, functionally similar to wide range of natural pores and should be able to withstand dierent experimental conditions. Gramicidin was the rst ion selective pore to be crystallized. The availability of structural details as well as functional robustness sealed the place for Gramicidin as the model pore. Gramicidin is a highly hydrophobic peptide containing alternating L and D amino acids. Both the N and C terminals of the peptide are "blocked" meaning having no free charges, making it poorly soluble in water. The chirality of the amino acids of the linear peptide renders Gramicidin sensitive to its environment. There are two main folding conformations identied for the peptide : 1 ) Double helical intertwined helix (non channel conformation) found when dissolved in organic solvents like chloroform or methanol. 2) Single stranded helical dimer or channel form. Why does the channel conformation have amphillic tryptophan at channel interface? Figure 0.4: Channel and Non channel conformations of Gramicidin. Gramicidin channel formation occurs as seen in Figure (0.4) by two peptides linked head to head by hydrogen bonds, each forming a half channel. The tryptophan residues along the channel are both hydrophobic as well as capable of long range electrostatic interactions. The activity of the channel measured in planar lipid bilayers requires the channel form of the peptide . Gramicidin exhibits a sort of "memory", based on its storage before addition to the bilayer. If the peptide is stored in methanol or chloroform the peptides have to establish the pore by rearranging from their non channel form to channel conformation which is aided by heating or sonication. However, if stored in solvents like triuroethanol the peptide adopts its channel form instantaneously. Fully formed Gramicidin channel has an aqueous pore of 4 Åand about 25 Ålong. The pore is 4 Figure 0.5: Left:Gramicidin pore; Right: Monomer arrangement during closed and open conformations. lined with polar residues and is cation selective. Gating of the pore is seen due to dissociation of the monomers by breaking of the hydrogen bonds between them. Since the pore is shorter than the bilayer the pore insertion would cause membrane deformation which is shown in gure (0.5). Alpha Hemolysin Alpha Hemolysin is considered an archetype pore for all beta barreled shaped channels. It is a toxin secreted by Staphylococcus Aureus which causes cell death by perforating the membrane. Figure 0.6: Alpha hemolysin : Three parts of the fully formed pore and the water lled transmembrane channel. The fully formed pore contains seven subunits which rearrange in the membrane to form a mushroom shaped pore 100 Åin length and ranging from ≈ 16 - 46 Åin diameter. The peptide sequence of each monomer reveals a glycine rich hydrophobic region between the C and N terminals. The structure can be divided into : The cap, The rim and the stem. The cap and the rim contain charged and polar amino acids where as the bulk of the hydrophobic patch folds into the stem region. The rim houses 7 important aromatic amino acids capable of membrane interactions. 0.1.4 Biomimetic DNA channels DNA as a building material is based on its unique base pairing abilities. DNA origami technique exploits this property to "tie up" a long circular DNA, scaold, into a previously conceived structure 5 using 200-250 shorter DNA sequences or staples. The shorter DNA sequences are designed to be complementary to certain parts of the scaold based on which dierent parts are brought together using Mg to reduce or eliminate the repulsion between negatively charged DNA segments. +2 Figure 0.7: DNA origami technique; a) Depicts a linear single stranded DNA scaold in black. Typically M13 phage of length 7249 base pairs is used. The tiny colored strands represent the staple strands which are designed to be complementary to specic parts of the scaold using caDNAno software. b-d) Based on the scaold routing designed as well as staple sequences DNA origami structure is formed using a one pot reaction technique where all the components are added to a tube and a heating ramp is applied to allow annealing of the staples to the scaold Taking inspiration from the biological protein pore structures, DNA origami channels with nanometer pore dimensions are designed. In order to allow interaction and further incorporation of a negatively charged hydrophillic structure into the lipid bilayer, the structure is decorated with cholesterol or tocopherol molecules at its base (much like the "rim" in Alpha Hemolysin). The hydrophobic molecules are added by using modied staple strands having a 5 or 3 end replaced with the desired molecule. The versatility of the technique allows to form these channels with dierent shapes and dierent central pore size. The membrane interaction ability is checked using small unilamellar vesicles which is a curved lipid bilayer. Transmission electron microscopy is used to indicate interaction tendency of the modied DNA structures. The energy required for a channel to penetrate through a membrane would be roughly speaking the amount of energy required to disrupt the lipid interactions in an area which is the same as that of the pore. This follows that larger the pore greater will be the energy required to insert it into the membrane as a larger area of the lipids are cleared away or rearranged to accommodate for the pore. This energy in DNA channels is provided by favorable interactions of the hydrophobic molecules on the channel with the lipid tails. Can you calculate the energy required to insert a 4 nm pore into the ' ' lipid bilayer and how many cholesterols are required to do the same? Applications of the bioinspired DNA channels are aimed at designing a sensor using resistive pulse technique (previously introduced) to detect single molecules and distinguish mixtures of molecules via their electrical signatures. Another exciting eld with potential application for DNA channels is as a component of an articial cell to allow selective passage of molecules in and out of a compartment. The ease of modication makes investing in DNA channels lucrative which additionally also sheds light to basic membrane interaction parameters. 6 Figure 0.8: a)The base of the channel is decorated with hydrophobic molecules which help in providing energy required for the "stem" of the channel to pierce through the lipid bilayer.b) First two columns show the various shapes of bio inspired DNA channels that have been realized. The third and fourth column show the TEM images conrming structure formation and membrane interaction respectively 0.2 Experimental protocol 0.2.1 Over view The practical course will provide basic introduction to lipid bilayers and protein channels. The rst half of the course entails using puried protein pores on painted lipid bilayers. This is done using Orbit mini setup. The second half of the practical course involves liposome production and imaging. This part also introduces DNA origami pores, purication methods and nally interaction with liposomes imaged using uorescence imaging. 0.2.2 Painted bilayer setup and electrical characterization of channels 1) The Orbit mini setup is used for demonstrating electrical activity of ion channels in a lipid bilayer. The recording is done on a MECA chip containing a 2x2 array of microcavities in a polymer. Each hole or cavity is associated with an individual integrated Ag/AgCl microelectrode. 2) The chip is inserted to the orbit mini setup and 150 µl of 100 mM KCl is added to wet the chip. This is indicated by an electrical "open channel" signal. 3) Lipid bilayers are formed on the MECA chip with four channels. DPhPC lipids dissolved in octane are used at a concentration of 1 mg/ml and painted across the four cavities using a brush. Formation of the bilayer is observed by reduced or no current passing through the channels otherwise called a "seal" signal (Indicated by a large resistance in the order of gigaohms. 4) The protein or DNA pores are added to the chip and a V or a constant voltage is applied to assist the pore insertion to the bilayer. Once steps of insertions are observed as in (g.1), voltage is changed to obtain an I-V curve. hold 7 0.2.3 Giant unilamellar vesicles - Inverted emulsion technique There are several methods for Giant unilamellar vesicle formation. The methodology used here is called inverted emulsion technique. 1) Lipids dissolved in an organic solvent are processed in a rotary evaporator to produce a uniform lipid lm under vacuum. This produces a thin lipid lm or cake which is hydrated in subsequent steps to yeild the GUV formation. 2) The lipid lm is dissolved in mineral oil by repeated heating and vortexing. The solution is sonicated and allowed to stand at room temperature overnight for maximum dissolution of the lipid into the mineral oil. 3)The lipid oil mixture is now ready to be processed for further. The 200 µl of the solution is placed in an eppendorf on ice. 30 µl of the solution designed to be inside the vesicle (usually 200 mM sucrose solution) is added to the cooled down emulsion and vortexed for 30 s. This step helps in formation of aqueous droplets surrounded by lipid molecules.The solution is allowed to stand for 30-45 minutes to stablise the droplets 4) The droplet emulsion is now gently placed on top of the solution intended to be outside the vesicles (usually 200mM Glucose solution). Care must be taken to adjust the osmolarity of the internal and external solutions (should be almost equal, outside solution having ≈ 20 mosm more osmolarity helps in dragging the vesicles to the bottom of the imaging chamber). This allows all the free lipids to line up at the emulsion external solution interface, creating a layer of free lipids. This is again allowed to stand for 45 minutes. 5) The column of solution is now gently centrifuged at 12,000 rcf for half hour. This transfers the lipid surrounded droplets through another layer of the free lipids into the aqueous external solution. As the droplets move through the interface it zips o a part of the free lipid layer forming a bilayer along the droplet. This "hole" in the lipid layer at the interface is then lled up by more free lipids present in the oil solution. 6) The GUVs are present at the bottom of the centrifuge. The oil layer is removed using a pipette and the vesicles are added to the imaging chamber. 0.2.4 Modication of the DNA pores with Fluorescentlylabeled staples for imaging 1) DNA origami pores are previously folded in two eppendorfs with a scaold concentration of 50 nM and staple concentration of 150 nM. To one of the tubes 3 fold excess of tocopherol modied staples (57 possible positions) as well as 3 fold excess of Fluorescentlylabeled staples are added and incubated for 45 minutes. The second tube is incubated with only Fluorescentlylabeled staples. 2) Both the hydrophobically tagged as well as the plain pores are puried using lter purication. Millipore lters with a 100 kDa cutos are used for removing the excess staples.The lters are rst equilibrated with the buer (500 mM KCl) by adding 500 µl of buer to the lter, placing it in the spin column and centrifuging at 5000 rcf for 5 minutes.The solute is removed from the spin column after the centrifugation. 8 3)450 µl of fresh buer is added to the lter and covered at the top with the sample to be puried. The lter is centrifuged at 5000 rcf for 5 minutes. The solute is then removed again from the column and fresh buer is added. The sample is washed with 5 rounds of such centrifugation steps. 4) In order to retrieve the sample the lter is inverted into an empty spin column and spun down at 5000 rcf for 5 minutes. The solution expelled in the spin column now contains the origami sample in the required buer. 0.2.5 Fluorescence based detection of hydrophobic interactions between DNA and lipid membranes 1) The imaging chamber is prepared using IBID chambers. 60 µl of the outside solution is added to the chamber and 5 µl of the GUV solution is added and mixed well. The chamber is kept undisturbed to allow the GUVs to sink for easier imaging. 2) The GUVs are seen using IX-71 microscope and bright eld images of the vesicles are made. 3) Fluorescentlylabelled DNA pores with and without hydrophobic tags are added to dierent lanes of the chamber. Colocalization of the pores modied with hydrophobic tags is observed and imaged in contrast to lack of attachment of the pores to the vesicles in absence of the hydrophobic tags. 0.3 Investigation The main objective of the practical course is to provide an understanding of how channels and membranes interact. The following questions should be answered after the experimental part is completed. 1) What is the conductance of a Gramicidin pore and Alpha Hemolysin in 100 mM KCl? Illustrate with trace and steps to deduce. 2) If the energy per lipid due to tail-tail contact is 20 kT, how much energy is required to form a pore of radius 2.5 nm? Hint : Consider the area in the bilayer reshued due to insertion of the pore. 3) How would the IV curve of a pore change with salt for eg: 100 mM KCl and 100 mM NaCl? 4) Illustrate the inverted emulsion method of vesicle formation diagramatically. 5) What should be the minimum length of an origami channel? Comment on the number of hydrophobic tags requried to allow the pore to insert a bilayer. 9