* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download proteinstructure
Point mutation wikipedia , lookup
Gene expression wikipedia , lookup
Magnesium transporter wikipedia , lookup
Expression vector wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Interactome wikipedia , lookup
Protein purification wikipedia , lookup
Western blot wikipedia , lookup
Metalloprotein wikipedia , lookup
Structural alignment wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Homology modeling wikipedia , lookup
Protein–protein interaction wikipedia , lookup
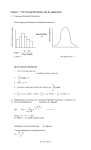
Protein Structure Helps us Understand Protein Function If we do know what a protein does, its structure will tell us how it does it. If we don’t know what a protein does, its structure might give us what we need to know to figure out its function. IIT Biochemistry: 17 Sep 2007 Slide 1 of 56 Plans for Today Methods of Determining Protein Structure Crystallography NMR CryoEM Specialty techniques IIT Biochemistry: 17 Sep 2007 Levels of Protein Structure Hydrogen Bonds Secondary structure in globular proteins Tertiary Structure Domains Slide 2 of 56 Warning: Specialty Content! I determine protein structures (and develop methods for determining protein structures) as my own research focus So it’s hard for me to avoid putting a lot of emphasis on this material But today I’m allowed to do that, because it’s the stated topic of the day. IIT Biochemistry: 17 Sep 2007 Slide 3 of 56 Structures: Fourier transforms of diffraction results Position of spots tells you how big the unit cell is Intensity tells you what the contents are We’re using electromagnetic radiation, which behaves like a wave, exp(2ik•x) Therefore intensity Ihkl = C*|Fhkl|2 Fhkl is a complex coefficient in the Fourier transform of the electron density in the unit cell: (r) = (1/V) hkl Fhkl exp(-2ih•r) IIT Biochemistry: 17 Sep 2007 Slide 4 of 56 F The phase problem a Note that we said Ihkl = C*|Fhkl|2 That means we can figure out |Fhkl| = (1/C)√Ihkl But we can’t figure out the direction of F: Fhkl = ahkl + ibhkl = |Fhkl|exp(ihkl) This direction angle is called a phase angle Because we can’t get it from Ihkl, we have a problem: it’s the phase problem! IIT Biochemistry: 17 Sep 2007 Slide 5 of 56 b What can we learn Electron density map + sequence we can determine the positions of all the non-H atoms in the protein—maybe! Best resolution possible: Dmin = / 2 Often the crystal doesn’t diffract that well, so Dmin is larger—1.5Å, 2.5Å, worse Dmin ~ 2.5Å tells us where backbone and most side-chain atoms are Dmin ~ 1.2Å: all protein atoms, most solvent, some disordered atoms IIT Biochemistry: 17 Sep 2007 Slide 6 of 56 What does this look like? Takes some experience to interpret Automated fitting programs work pretty well with Dmin < 2.1Å ATP binding to a protein of unknown function: S.H.Kim IIT Biochemistry: 17 Sep 2007 Slide 7 of 56 How’s the field changing? 1990: all structures done by professionals Now: many biochemists and molecular biologists are launching their own structure projects as part of broader functional studies Fearless prediction: by 2020, crystallographers will be either technicians or methods developers IIT Biochemistry: 17 Sep 2007 Slide 8 of 56 Macromolecular NMR NMR is a mature field Depends on resonant interaction between EM fields and unpaired nucleons (1H, 15N, 31S) Raw data yield interatomic distances Conventional spectra of proteins are too muddy to interpret Multi-dimensional (2-4D) techniques: initial resonances coupled with additional ones IIT Biochemistry: 17 Sep 2007 Slide 9 of 56 Typical protein 2-D spectrum Challenge: identify which H-H distance is responsible for a particular peak Enormous amount of hypothesis testing required IIT Biochemistry: 17 Sep 2007 Prof. Mark Searle, University of Nottingham Slide 10 of 56 Results Often there’s a family of structures that satisfy the NMR data equally well Can be portrayed as a series of threads tied down at unambiguous assignments They portray the protein’s structure in solution IIT Biochemistry: 17 Sep 2007 Slide 11 of 56 Comparing NMR to X-ray NMR family of structures often reflects real conformational heterogeneity Nonetheless, it’s hard to visualize what’s happening at the active site at any instant Hydrogens sometimes well-located; they’re often the least defined atoms in an Xray structure The NMR structure is obtained in solution! Hard to make NMR work if MW > 25 kDa IIT Biochemistry: 17 Sep 2007 Slide 12 of 56 What does it mean when NMR and X-ray structures differ? Lattice forces may have tied down or moved surface amino acids in X-ray structure NMR may have errors in it X-ray may have errors in it (measurable) X-ray structure often closer to true atomic resolution X-ray structure has built-in reliability checks IIT Biochemistry: 17 Sep 2007 Slide 13 of 56 Cryoelectron microscopy Like X-ray crystallography, EM damages the samples Samples analyzed < 100K survive better 2-D arrays of molecules Spatial averaging to improve resolution Discerning details ~ 4Å resolution Can be used with crystallography IIT Biochemistry: 17 Sep 2007 Slide 14 of 56 Circular dichroism Proteins in solution can rotate polarized light Amount of rotation varies with Effect depends on interaction with secondary structure elements, esp. Presence of characteristic patterns in presence of other stuff enables estimate of helical content IIT Biochemistry: 17 Sep 2007 Slide 15 of 56 Poll question: discuss! Which protein would yield a more interpretable CD spectrum? (a) myoglobin (b) Fab fragment of immunoglobulin G (c) both would be fully interpretable (d) CD wouldn’t tell us anything about either protein IIT Biochemistry: 17 Sep 2007 Slide 16 of 56 Ultraviolet spectroscopy Tyr, trp absorb and fluoresce: abs ~ 280-274 nm; f = 348 (trp), 303nm (tyr) Reliable enough to use for estimating protein concentration via Beer’s law UV absorption peaks for cofactors in various states are well-understood More relevant for identification of moieties than for structure determination Quenching of fluorescence sometimes provides structural information IIT Biochemistry: 17 Sep 2007 Slide 17 of 56 Solution scattering Proteins in solution scatter X-rays in characteristic, sphericallyaveraged ways Low-resolution structural information available Does not require crystals Until ~ 2000 you needed high [protein] Thanks to BioCAT, SAXS on dilute proteins is becoming more feasible Hypothesis-based analysis IIT Biochemistry: 17 Sep 2007 Slide 18 of 56 Fiber Diffraction Some proteins, like many DNA molecules, possess approximate fibrous order (2-D ordering) Produce characteristic fiber diffraction patterns Collagen, muscle proteins, filamentous viruses IIT Biochemistry: 17 Sep 2007 Slide 19 of 56 X-ray spectroscopy All atoms absorb UV or X-rays at characteristic wavelengths Higher Z means higher energy, lower for a particular edge Perturbation of absorption spectra at E = Epeak + yields neighbor information Changes just below the peak yield oxidation-state information X-ray relevant for metals, Se, I IIT Biochemistry: 17 Sep 2007 Slide 20 of 56 Levels of Protein Structure We conventionally describe proteins at four levels of structure, from most local to most global: Primary: linear sequence of peptide units and covalent disulfide bonds Secondary: main-chain H-bonds that define short-range order in structure Tertiary: three-dimensional fold of a polypeptide Quaternary: Folds of multiple polypeptide chains to form a complete oligomeric unit IIT Biochemistry: 17 Sep 2007 Slide 21 of 56 What does the primary structure look like? -ala-glu-val-thr-asp-pro-gly- … Can be determined by amino acid sequencing of the protein Can also be determined by sequencing the gene and then using the codon information to define the protein sequence Amino acid analysis means percentages; that’s less informative than the sequence IIT Biochemistry: 17 Sep 2007 Slide 22 of 56 Components of secondary structure , 310, helices pleated sheets and the strands that comprise them Beta turns More specialized structures like collagen helices IIT Biochemistry: 17 Sep 2007 Slide 23 of 56 An accounting for secondary structure: phospholipase A2 IIT Biochemistry: 17 Sep 2007 Slide 24 of 56 Alpha helix IIT Biochemistry: 17 Sep 2007 Slide 25 of 56 Characteristics of helices Hydrogen bonding from amino nitrogen to carbonyl oxygen in the residue 4 earlier in the chain 3.6 residues per turn Amino acid side chains face outward ~ 10 residues long in globular proteins IIT Biochemistry: 17 Sep 2007 Slide 26 of 56 What would disrupt this? Not much: the side chains don’t bump into one another Proline residue will disrupt it: Main-chain N can’t H-bond The ring forces a kink Glycines sometimes disrupt because they tend to be flexible IIT Biochemistry: 17 Sep 2007 Slide 27 of 56 Other helices NH to C=O four residues earlier is not the only pattern found in proteins 310 helix is NH to C=O three residues earlier More kinked; 3 residues per turn Often one H-bond of this kind at Nterminal end of an otherwise -helix helix: even rarer: NH to C=O five residues earlier IIT Biochemistry: 17 Sep 2007 Slide 28 of 56 Beta strands Structures containing roughly extended polypeptide strands Extended conformation stabilized by inter-strand main-chain hydrogen bonds No defined interval in sequence number between amino acids involved in H-bond IIT Biochemistry: 17 Sep 2007 Slide 29 of 56 Sheets: roughly planar Folds straighten H-bonds Side-chains roughly perpendicular from sheet plane Consecutive side chains up, then down Minimizes intra-chain collisions between bulky side chains IIT Biochemistry: 17 Sep 2007 Slide 30 of 56 Anti-parallel beta sheet Neighboring strands extend in opposite directions Complementary C=O…N bonds from top to bottom and bottom to top strand Slightly pleated for optimal H-bond strength IIT Biochemistry: 17 Sep 2007 Slide 31 of 56 Parallel Beta Sheet N-to-C directions are the same for both strands You need to get from the C-end of one strand to the N-end of the other strand somehow H-bonds at more of an angle relative to the approximate strand directions Therefore: more pleated than anti-parallel sheet IIT Biochemistry: 17 Sep 2007 Slide 32 of 56 Beta turns Abrupt change in direction , angles are characteristic of beta Main-chain H-bonds maintained almost all the way through the turn Jane Richardson and others have characterized several types IIT Biochemistry: 17 Sep 2007 Slide 33 of 56 Collagen triple helix Three left-handed helical strands interwoven with a specific hydrogen-bonding interaction Every 3rd residue approaches other strands closely: so they’re glycines IIT Biochemistry: 17 Sep 2007 Slide 34 of 56 Poll question Remember that there are about 3.6 residues per turn in an alpha helix. Suppose you had a helical protein that was sitting on, not in, a phospholipid bilayer so that the side chains point inward and outward along the surface. Which of the following sequences would be the most stable in this environment? IIT Biochemistry: 17 Sep 2007 Slide 35 of 56 Options Assume side chain of residue 2 points DOWN into the bilayer: (a) GADHKYEKLRG (b) GLDGIVESVGG (c) AKRTTVWKDKD (d) YRNNADRRKLG IIT Biochemistry: 17 Sep 2007 Slide 36 of 56 Tertiary Structure The overall 3-D arrangement of atoms in a single polypeptide chain Made up of secondary-structure elements & locally unstructured strands Described in terms of sequence, topology, overall fold, domains Stabilized by van der Waals interactions, hydrogen bonds, disulfides, . . . IIT Biochemistry: 17 Sep 2007 Slide 37 of 56 Quaternary structure Arrangement of individual polypeptide chains to form a complete oligomeric, functional protein Individual chains can be identical or different If they’re the same, they can be coded for by the same gene If they’re different, you need more than one gene IIT Biochemistry: 17 Sep 2007 Slide 38 of 56 Not all proteins have all four levels of structure Monomeric proteins don’t have quaternary structure Tertiary structure: subsumed into 2ndry structure for many structural proteins (keratin, silk fibroin, …) Some proteins (usually small ones) have no definite secondary or tertiary structure; they flop around! IIT Biochemistry: 17 Sep 2007 Slide 39 of 56 Note about disulfides H Cysteine residues brought S into proximity under H oxidizing conditions can C form a disulfide Forms a “cystine” residue Oxygen isn’t always the oxidizing agent Can bring sequence-distant residues close together and stabilize the protein IIT Biochemistry: 17 Sep 2007 H S H H + (1/2)O 2 H2O H H C C S S H Slide 40 of 56 H H C Hydrogen bonds, revisited Biological settings, H-bonds are almost always: Between carbonyl oxygen and hydroxyl: (C=O ••• H-O-) between carbonyl oxygen and amine: (C=O ••• H-N-) These are stabilizing structures Any stabilization is (on its own) entropically disfavored; Sufficient enthalpic optimization overcomes that! In general the optimization is ~ 1- 4 kcal/mol IIT Biochemistry: 17 Sep 2007 Slide 41 of 56 Secondary structures in structural proteins Structural proteins often have uniform secondary structures Seeing instances of secondary structure provides a path toward understanding them in globular proteins Examples: Alpha-keratin (hair, wool, nails, …): -helical Silk fibroin (guess) is -sheet IIT Biochemistry: 17 Sep 2007 Slide 42 of 56 Alpha-keratin Actual -keratins sometimes contain helical globular domains surrounding a fibrous domain Fibrous domain: long segments of regular helical bonding patterns Side chains stick out from the axis of the helix IIT Biochemistry: 17 Sep 2007 Slide 43 of 56 Silk fibroin Antiparallel beta sheets running parallel to the silk fiber axis Multiple repeats of (Gly-Ser-GlyAla-Gly-Ala)n IIT Biochemistry: 17 Sep 2007 Slide 44 of 56