* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Lineage Commitment During T cell Development
Survey
Document related concepts
Transcript
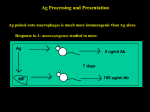
Summary of Thymic Development Zuniga-Pflucker, NRI, 2004 What do we mean by the term “Lineage Commitment”? What do we mean by the term “Lineage Commitment”? Commitment represents the loss in the ability of a cell to make alternative lineage choices under permissive conditions. Name the Lineage Commitment Choices that are made during T cell Development Lineage Commitment Decisions During T Cell Development • • • • To become a T cell To become an TCRab cell vs a TCRgd T cell To become a CD4 or CD8 T cell To become one of the minor T cells subset NK T cell CDaa T cell nTreg First Lineage Commitment Decision in T Cell Development To be, or not to be, a T cell Seeding the Thymus from Precursors in the Blood Bhandoola, et al., Immunity, 26:678-689, 2007 HSC Hematopoietic Stem Cell MPP Multipotent Progenitor ELP Early Lymphoid Progenitor CLP Common Lymphoid Progenitor CMP Common Myeloid Progenitor CTP Circulating T cell Progenitor TSP Thymus Settling Progenitors ETP Early Thymic Progenitor Is the ETP, recently arrived in the thymus, already committed to the T cell lineage? Alternative Fates of the Early Thymic Progenitor Yui and Rothernberg, NRI, 2014 A critical role for Notch in T cell lineage commitment Zuniga-Pflucker, NRI, 2004 How would you demonstrate the importance of Notch in T cell lineage commitment of ETPs? OP9 stromal cells transfected with Deltex-1 can support T Cell Lineage Commitment Zuniga-Pflucker, NRI, 2004 Schmitt, et al, JEM, 2004 ab T Cell Development Yui and Rothernberg, NRI, 2014 Notch signaling is critical for the induction of multiple transcription factors Yui and Rothernberg, NRI, 2014 Early Notch signaling induces TCF-1 (Tcf7) and Gata3 at The DN1 -> DN2 transition Yui and Rothernberg, NRI, 2014 How would you test the importance of one of the transcription factors (for instance, TCF-1) that is upregulated during early Notch signaling for T cell lineage development? Impaired development of TCF-1-deficient BM HSC into CD25+ (i.e.,DN2) thymocytes in OP9 cultures Weber, et al., Nature, 2011 Defects also observed in vivo using competitive repopulation studies of WT vs TCF-1 KO BM Evolving Transcriptional Networks as Notch Influences Early T Cell Development Yui and Rothernberg, NRI, 2014 DN Cells (CD4-/CD8-) DN1 DN2 DN3 DN4 Sequential Rearrangement of TCR ab Genes DN DP Abbas & Lichtman. Cellular and Molecular Immunology, 5th ed. W. B. Saunders 2003 The pre-TCR is Expressed in DN cells g Pre-Ta functions as a surrogate for the a chain during thymic development Expressed in DN cell and heterodimerizes with a functional b chain - assists in quality control for b chain rearrangement The pre-Ta/b chain dimer promotes increased CD3 expression and induces a ligandindependent signal, perhaps because of constitutive localization to lipid rafts or constitutive dimerization (unusual preTalpha structure), that is responsible for maturation and probably shut off RAG expression and further rearrangement, resulting in b chain allelic exclusion How would you test the ligand-independency of signaling by the pre-TCR? Reconstitution of rag1-/- mice with various forms of the pre-TCR Irving, et al., Science, 1998 Ligand-Independent Dimerization of the pre-TCR Pang, et al, Nature 2010 Extended structure of Pre-Ta compared to TCR Ca Dimer of Heterodimers of pre-Ta and TCR b Two Lineages of Cells Expressing Distinct TCRs Develop in the Thymus: Stages of gd and ab T Cell Development Modified from Ciofani and Zuniga-Pflucker, Nature Rev. Immunol., 2010 (C-Kit) Two Lineages of T cells (cont.) Stochastic Instructive model Recent data suggest that gd receptor expression results in stronger signal that can provides instructional cue for cell to become gd lineage ( Reviewed in Ciofani, et al., Nat. Rev. Immunol. 2010) Why might the pre-TCR and the TCRgd signal strengths be different differently? Strength of signal: a fundamental mechanism for cell fate specification Immunological Reviews Volume 209, Issue 1, pages 170-175, 31 JAN 2006 DOI: 10.1111/j.0105-2896.2006.00356.x http://onlinelibrary.wiley.com/doi/10.1111/j.0105-2896.2006.00356.x/full#f3 Reciprocal regulation of Syk and ZAP-70 expression during thymocyte development Palacios & Weiss, JEM, 2007 WT DN3 Development b-selection TCR selection CD44CD25+ DN3 Strength of signal: a fundamental mechanism for cell fate specification Immunological Reviews Volume 209, Issue 1, pages 170-175, 31 JAN 2006 DOI: 10.1111/j.0105-2896.2006.00356.x http://onlinelibrary.wiley.com/doi/10.1111/j.0105-2896.2006.00356.x/full#f1 How might you test the signaling hypothesis? Starting with a TCRgd Transgene: Increasing signaling strength by elimination of a negative regulator, CD5, favors gd lineage commitment Hayes, et al., Immunity, 2005 Instructing ab vs gd Lineage Commitment via Strength of Signal Ciofani and Zuniga-Pflucker, Nat. Rev. Immunol. 2010 Some caveats to the strength of signaling Stochastic Instructive model SOX13, an SRY-related HMG-box transcription factor, is preferentially expressed in gd T cells. Its KO, decreases gd development. Its over-expression in DN thymocytes impairs DN -> DP transition and ab T cell development. Bcl11b, a zinc finger transcription factor, is preferentially expressed in ab lineage T cells and is induced in DN2a-DN2b. It is low in the gd lineage. The KO of Bcl11b has little effect on gd T cell development but completely impairs ab lineage T cell development. There’s more to come. Checkpoints in Thymocyte Development Modified from Carpenter and Bosselut, Nature Immunology 2010 Notch Pre-TCR TCRab Linking CD4 (helper) to CD8 (killer) T cell lineage commitment to the recognition of class I versus class II MHC ensures that T cell effector functions are directed appropriately. MHC class I pathway samples intracellular antigens (e.g. viruses, intracellular bacteria). MHC class II pathway samples extracellular antigens Cytotoxic CD8 T cells can kill invaded host cells before pathogens can replicate and spread. Helper T cells regulate the activity of other cells of the immune system that have endocytosed pathogens. Recognition of MHC-1 or MHC-2 during positive selection in the thymus determines the CD4 versus CD8 T cell lineage choice. Recognition of MHC-1 or MHC-2 during positive selection in the thymus determines the CD4 versus CD8 T cell lineage choice. CD4 CD8 MHC-1 CD8 MHC-2 CD4 Early evidence of the link between TCR specificity for MHC1 vs. MHC-2 and the CD4 vs. CD8 lineage choice: TCR transgenic mice and MHC gene ko mice. Generation of transgenic mice bearing rearranged TCR genes with defined specificity provided important tools for study of positive selection. T cell receptor transgenic mice: Antigen (e.g., ovalbumin) Rest and restimulate with Ag and cytokines, rest, repeat 9 days T cells + DCs + ovalbumin plate at 1 cell/well Test for antigen specificity grow and expand clone the TCR alpha and beta chain genes from the T cell clone T cell receptor alpha- & beta-chain genes specific for MHC class I or MHC class II OT-1 - CD8 T cells AND - CD4 T cells All T cells will express the same TCRab receptor Increased development of CD4 SP thymocytes and T cells in mice expressing a rearranged class II MHC specific transgene 57.4 30.6 9.6 CD4 CD8 0.69 Generating H-Y TCR transgenic mice (anti-male peptide TCR) Male H-2b cells 9 days Female H-2b mouse Expand anti-male CD8 T cells Rest and restimulate with Ag and cytokines, rest, repeat plate at 1 cell/well Test for antigen specificity grow and expand clone the TCR alpha and beta chain genes from the T cell clone Increased development of CD8 SP thymocytes and T cells in mice expressing a rearranged class I MHC specific transgene 5% 37% 10% 46% TCR specificity for MHC I or II determines CD4 versus CD8 lineage commitment OT1, HY, F5, etc OT2, DO11.10, AND, etc MHC deficient mice provide evidence for positive selection. Lack of MHC class II expression prevents development of CD4 cells WT MHC Class II o CD4 CD4 . . . .. .. . . . ……. .. . . .. .. ... .. . . CD8 CD8 And lack of MHC class I expression (b2-microglobulin deficient mice) prevents development of CD8 T cells. MHC class I and II double deficient mice lack both CD4 and CD8 mature T cells, but have normal numbers of DP thymocytes. CD4 CD8 MHC-1 CD8 MHC-2 CD4 What are the gene expression programs that determine the CD4 or CD8 T cell fate? How are distinct gene expression programs linked to TCR recognition of MHC class I or class II? CD4 CD8 MHC-1 CD8 MHC-2 CD4 What are the gene expression programs that determine the CD4 or CD8 T cell fate? How are distinct gene expression programs linked to TCR recognition of MHC class I or class II? How would you identify transcription factors involved in CD4 versus CD8 lineage commitment? How would you identify transcription factors involved in CD4 versus CD8 lineage commitment? Gene profiling mature CD4 vs CD8 T cells to identify differentially expressed TF. Identify TF that regulate key CD4 vs CD8 specific target genes (CD4, CD8, perforin, CD40L, etc) Gene KO of candidate TF and assess impact on CD4, CD8 T cell development. (embryonic lethality, blocks at earlier stages of development (T commitment, TCRb checkpoint, etc) Serendipity Th-POK (c-KROX)--a “master regulator” of CD4 T cell lineage commitment 1997: “HD” mouse strain: lacks CD4 T cells spontaneous mutation in D. Kappes’ animal colony Autosomal recessive Not due to defect in MHC Class II Genetic mapping and BAC complementation used to discover the mutant gene: Th-POK The “HD” allele has a single point mutation in a zinc-finger domain CD4 Block in CD4 T cell development, or “lineage swap”? How could you tell? CD8 He et al. Nature 2005 Mutation in ThPOK leads to the development of “mismatched” class II specific, CD8 T cells CD4 Uncoupling between positive selection and lineage commitment. CD8 Over-expression of ThPOK in thymocytes lead to reciprocal phenotype: CD4 T cells with TCR specific for MHC-1. OT1 TCR tg OT1 x ThPOK tg MHC-2 ko MHC-2 ko x ThPOK tg ThPOK acts together with other transcription factors in a network that specifies the CD4 fate. GATA-3 is required for ThPOK to induce the CD4 fate (but not to repress CD8 fate). GATA-3 promotes ThPOK expression. Loss of GATA-3 can (sometimes) divert thymocytes with TCR specific for MHC-2 into the CD8 lineage. Wang et al Nat Immunol. 2008 Oct;9(10):1122 ThPOK reinforces its own expression and acts together with other transcription factors in a network that specifies the CD4 fate. ThPOK opposes Runx repression of CD4 and ThPOK expression. Mutation of both Runx and ThPOK rescues CD4 development. Mutant ThPOK allele than cannot undergo positive autoregulation leads to “trans-differentiation” of CD4 cells to the CD8 lineage. Egawa and Littman and Miroi et al Nat Immunol. 2008 TRANSCRIPTION FACTOR NETWORK CONTROLING CD4 VS CD8 FATE: THE SIMPLE VERSION MHC-2 positive selection Autoregulation ThPOK CD4 MHC-1 positive selection Mutual antagonism RUNX3 CD8 TRANSCRIPTION FACTOR NETWORK CONTROLING CD4 VS CD8 FATE: MORE PLAYERS MHC-2 positive selection MHC-1 positive selection cMyb MAZR GATA3 ThPOK Tox RUNX3 MAZR ThPOK CD8 CD4 RUNX3 CD4 CD8 Would you consider ThPOK to be a “master regulator” of the CD4 fate. Why or why not? CD4 CD8 MHC-1 CD8 MHC-2 CD4 What are the gene expression programs that determine the CD4 or CD8 T cell fate? How are distinct gene expression programs linked to TCR recognition of MHC class I or class II? Relating the transcription factor networks that control CD4 versus CD8 to recognition of MHC-1 or MHC-2 during positive selection. An “instructive” model? More prolonged signal More transient signal A kinetic model for CD4/CD8 development: duration of TCR signals influence lineage choice. But clearly not the whole story. Quantitative model cannot adequately account for the stringent relationship between MHC-1 vs MHC-2 specificity and positive selection. Another mechanism to ensure that thymocytes adopt the CD4/CD8 fate appropriate for their TCR specificity: Some thymocytes make the “wrong” lineage choice, but a late check for coreceptor expression eliminate cells that turned down the wrong co-receptor. Thymocytes expressing TCR specific for MHC-1 A stochastic/selection model? Constitutive expression of CD8 (or CD4) leads to the (inefficient) development of “mismatched” T cells Davis et al ‘93, Itano et al ‘94, Robey et al ‘94, Baron et al ‘94, Corbella et al ‘94 Paterson et al ‘94, Chan et al ‘94, Rathemtulla et al, ‘02 How long does a thymocyte need to engage MHC and receive TCR signals in order to complete positive selection? How long does it take for a thymocyte to commit to the CD4 or CD8 lineage? Typical time required from initial encounter with extracellular ligand to changes in gene expression? Multiple changes accompany positive selection Pre selection thymocytes are: In cortex CD4+CD8+ TCRneg-low rag+ low expression of activation markers (CD69-) Short-lived, dep on TCR signal post selection thymocytes are: In medulla CD4+CD8- or CD4-CD8+ TCRhigh ragCD69+ Long-lived, indep of TCR signal These changes occur asynchronously over a period of 1-3 days. Can identify thymocytes exhibiting some, but not all signs of positive selection. Would you say that a cortical CD4+CD8+ CD69+ rag- thymocyte has been positively selected? How long does a thymocyte need to engage MHC and receive TCR signals in order to complete positive selection? “Antigen-experienced” CD4+CD8+ thymocytes isolated from TCR trangenic positive selecting mice (HY TCR, H2b) and intrathymically injected into mice with or without the positive selecting ligand. A single hit is not sufficient for positive selection of CD8 T cells: Recent data from Art and Ellen’s lab showing that 2-3 days of continuous TCR signaling needed for efficient positive selection of CD8 T cells (Au-Yeong et al, Ross et al 2014) How long does it take for a thymocyte to commit to the CD4 or CD8 lineage? Lingeage commitment represents the loss in the ability of a cell to make alternative lineage choices under permissive conditions. Impact of removing ThPOK after CD4 lineage commitment: What does TCR “signal strength” really mean? TCR affinity (signal strength) Death by neglect Positive selection CD4 lineage CD8 lineage commitment commitment TCR affinity for peptide-MHC complex? Duration, frequency, or intensity of TCR signaling? Presence/absence of co-stimulatory signals? Clonal deletion T reg development (agonist selection) Temporal requirement for TCR signaling during CD8 T cell positive versus negative selection? days hours Temporal pattern of TCR signaling during positive versus negative selection? intermittent Why does positive selection take 2-3 days? continuous How do distinct temporal signaling patterns encode distinct T cell fates? Positive selection and CD4 versus CD8 lineage development may be lengthy, reversible processes. Interpretation of TCR signaling difference and establishment of stable gene regulatory network for CD4 or CD8 fate likely occur at the same time and may be mechanistically linked. CD4 loop CD8 loop MHC-2 recognition: stronger, continuous signal MHC-1 recognition: weaker, intermittent signal GATA3 RUNX3 RUNX3 ThPOK ThPOK CD8 CD8 CD4 Maintain CD4 expression allowing for late MHC-2 recognition Maintain CD8 expression allowing for late MHC-1 recognition Loops can be initiated by biasing signals, and/ or stochastic fluctuations. Positive feedback makes lineage choices more robust and allows for re-verification.