* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download A Novel Temporal Expression Pattern of Three C
Extracellular matrix wikipedia , lookup
Signal transduction wikipedia , lookup
Tissue engineering wikipedia , lookup
Cell culture wikipedia , lookup
Organ-on-a-chip wikipedia , lookup
Cell encapsulation wikipedia , lookup
Cellular differentiation wikipedia , lookup
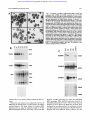
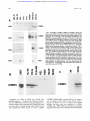
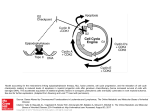
From www.bloodjournal.org by guest on August 11, 2017. For personal use only. A Novel Temporal Expression Pattern of Three C/EBP Family Members in Differ entia t i ng My el om onocyt i c Cell s By Linda M. Scott, Curt I. Civin, Pernille Rorth, and Alan D. Friedman Members of the CCAAT/enhancer binding protein (C/ EBP] family have been shown to regulate the terminal differentiation of adipocytes and hepatocytes. In these cell lineages, high levels of C/EBPa are found only in mature, nondividing cells. UsingWestern blotting and immunohistochemicalstaining, we have determinedthe temporal order of expression for C/EBPa, C/EBPp, and C/ EBPG in differentiatingmyelomonocytic marrow cells. These studies show a unique temporal pattern of C/EBP isoform expression in the myeloid lineage. In particular, C/EBPa expression is very high in proliferative myelomonocytic cells, and diminishes during phenotypic maturation. While we have detected C/EBPa, C/EBPp, and C/EBPG in multiple myeloid leukemia cell lines, and C/EBPa in normal myeloid cells and in de novo human myeloid leukemias, we have not detected these C/EBP isoforms in either erythroid or lymphoid cells. Finally, we show that C/EBPa. C/EBPp, and C/EBPS protein and messenger RNA levels correlate in maturing granulocytic cells. The formation of tissue-specific combinations of C/EBP homodimers and heterodimersmay allow this family of transcriptionfactors to regulate different sets of genes in adipocytes, hepatocytes, and myelomonocytes. o 1992by The American Society of Hematology. T function of granulocytic differentiation. Although variability of this cell line in culture resulted in some quantitative differences between experiments, a surprisingly different pattern from that described for adipocyte development was evident. Most notably, C/EBPa was found to be expressed at a high level in dividing myeloblasts and to diminish to low levels during their terminal differentiation into polymorphonuclear leukocytes (PMNs). We will discuss the implications of this novel temporal pattern of C/EBP isoform expression. HE CCAAT/enhancer-binding protein (C/EBP) family of transcription factors consists of several proteins with highly homologous dimerization and DNA contact domains. The founding member of this family, C/EBPa, was originally purified from rat This protein binds DNA as an obligate homodimer via a basic region-leucine zipper (bZIP) domain.'-6 The C/EBP family also includes C/EBPP (also known as NF-IL6, LAP, IM-DBP, AGP/ EBP, and CRP2),7-12CIEBPG (also known as CRP3),"J2 and C/EBPy (also known as Ig/EBP-1).I3 The various C/EBP family members can homodimerize, but can also heterodimerize and maintain DNA-binding activity.11-13 Members of the C/EBP family, especially C/EBPu, have been implicated in regulating the terminal differentiation of several mammalian cells. Although present at low levels in hepatoma cell lines, C/EBPa is expressed at high levels in mitotically quiescent hepatocytes, in which it is believed to regulate a variety of hepatocyte-specific genes.14J5 Similarly, although absent from 3T3-Ll preadipocytes, C/EBPa is expressed at high levels when these cells differentiate into nondividing adipocytes.11J6C/EBPa is capable of activating several "fat-specific'' genes in 3T3-Ll cells, including steroylCoA desaturase and the insulin-responsive glucose transp ~ r t e r . ' ~ JSelective * inhibition of C/EBPa expression in differentiating adipocytes by expression of antisense C/EBPa RNA reduced the ability of these cells to form lipid droplets, a phenotype of their terminally differentiated ~ t a t e . ' In ~ ,contrast ~~ to C/EBPa, C/EBPP and C/EBPG are expressed highest in the early stages of adipocyte formation." The ability of C/EBP family members to heterodimerize may allow complex regulation of hepatocyte and adipocyte development.21 In both hepatocytes and adipocytes, high levels of C/EBPa expression have been found only in nondividing cells. Moreover, when C/EBPa is ectopically expressed in dividing preadipocytes, mitotic proliferation ceases.22Therefore, it has been proposed that C/EBPu may regulate a genetic program that blocks cell divi~ion.~~J2 We have now discovered that members of the C/EBP family are also expressed in myelomonocytic cells of human and rodent bone marrow. Using the myelomonoblastic murine cell line 32D C13, a valuable model of granulocytic myeloid differentiati0n,2~,~~ we have examined the expression profile of C/EBPa, C/EBPP, and C/EBPS as a Blood, Vol 80, No 7 (October 1). 1992: pp 1725-1735 MATERIALS AND METHODS Cell culture, marrow cells, and human leukemia cells. 32D C13 cells23 were maintained at 37"C, 5% C o t in Iscove's modified Dulbecco's medium (IMDM) supplemented with 10% heatinactivated fetal bovine serum (HI-FBS) and 5% WEHI-3B supernatent as a source of interleukin-3 (IL-3).25 For induction of granulocytic differentiation, cells were washed twice with phosphatebuffered saline (PBS) and placed in IMDM/lO% HI-FBS supplemented with 500 U/mL granulocyte colony-stimulating factor (G-CSF; Amgen, Thousand Oaks, CA). Cell morphology was assessed by Wright-Giemsa ~taining.2~ HL-60, SP2, U937, PLB985, KGla, HEL, K.562, Molt-3, Daudi, and WEHI 274.1 cells were maintained in RPMII10% HI-FBS. P388D1 and IC-21 cells were grown in Dulbecco's minimum essential medium (DMEM)/ 10% HI-FBS. Cytotoxic T-cell lymphocytic leukemia (CTLL) cells were provided by A. Hess (John Hopkins Oncology Center), and From the Division of Pediatric Oncology, Zhe Johns Hopkins Oncology Center, Baltimore, MD; and the Department of Embyology, Camegze Institution of Washington, Baltimore, MD. SubmittedApnl IO, 1992; accepted June 15,1992. Supported by grants (to A.D.F.) from the W.W. Smith Charitable Trust, the Searle Scholars ProgramlChicago Community Trust, and the National Institute of Health (CA01326), by Grant No. CH-480 from the American Cancer Society (to C.I.C.), and by a grant from the Danish Medical Research Council (toP.R.). Address reprint requests to Alan D. Friedman, MD, Division of Pediatnc Oncology, Johns Hopkins Oncology Center, Room 3-109, 600 N Wove St, Baltimore, MD 21287. The publication costs of this article were defrayed in part by page charge payment. Thul article must therefore be hereby marked "advertisement" in accordance with 18 U.S.C.sectton 1734 solely to indicate thul fact. 0 I992 by The American Society of Hematology. 0OO4-4971/92l8007-0020$3.00l0 1725 From www.bloodjournal.org by guest on August 11, 2017. For personal use only. SCOTT ET AL 1726 induced 3T3-Ll cells were provided by Z. Cao (Carnegie Institution). Normal human marrow cells were obtained from the posterior iliac crest of volunteer donors. Approval was obtained from the Institutional Review Board for these studies. Informed consent was provided according to the Declaration of Helsinki. CD15+ cells were isolated as described.2628 In brief, marrow was first layered onto a cushion of Ficoll-Hypaque (Pharmacia, Uppsala, Sweden), and mononuclear cells were collected from the interface after centrifugation at 5OOg for 30 minutes at 22°C. The cells were then washed twice and incubated for 30 minutes at 4°C in RPMI/10% HI-FBS, with immunomagnetic Dynabeads M-450 (Dynal, Great Neck, NY), 5 beads per cell, to which had been adsorbed CD15 monoclonal antibody MOA^).^^ Beads were collected with a magnet and washed, and protein extraction buffer (see below) was added. The morphology and number of cells in the initial and unbound samples were determined by Wright-Giemsa staining of cytospin slides. Human leukemia cells, obtained from bone marrow or peripheral blood at diagnosis or relapse and stored in liquid nitrogen, were thawed, washed three times in the presence of DNase1 (100 UlmL), subjected to Ficoll-Hypaque centrifugation as above to remove dead cells and debris, washed, counted, and extracted. Each sample contained greater than 95% leukemic blasts, as verified by morphologic examination. Protein extraction and Western blotting. For protein extraction, cells were pelleted, washed with PBS, and extracted with 3 mL of 10 mmol/L Tris, pH 7.5, 2 mmol/L EDTA, 1% sodium dodecyl sulfate (SDS). The extract was then sonicated and concentrated by precipitation with four volumes of acetone at -20°C. The precipitates were resuspended in sample buffer by boiling for 5 minutes and applied to 10% polyacrylamide gels,)O 4 x lo6 cell equivalents per lane. After electrophoresis. the proteins were transferred to nitrocellulose?1 The integrity and levels of transferred proteins were determined by staining the blots with Ponceau S (0.2% in 2% trichloroacetic acid [TCA]) followed by destaining with 5% glacial acetic acid. Filters were then washed extensively with Tris-buffered salineTween (TBS-T, 150 mmol/L NaCI, SO mmol/L Tris, pH 7.5, 0.3% Tween-20), blocked for 20 minutes with TBS-T/5% nonfat dried milk (NFDM), incubated at room temperature for 1 hour with antisera diluted 1:1,000 in TBS-T/0.25% NFDM, washed three times with the same solution, incubated with horseradish peroxidase (HRP)-conjugated donkey antirabbit Ig (1:15,000), washed with TBS-T three times, and developed with ECL reagents (Amersham, Arlington Heights, IL) as described by the manufacturer. Developed filters were then exposed to Kodak XAR film (Eastman Kodak, Rochester, NY). C/EBPa peptide antisera3-16and the C/EBPP and C/EBPG antisera” were kindly provided by 2. Cao and S. McKnight (Carnegie Institution). The C/EBPa protein antiserum was prepared by expression of CIEBPAl-2 protein3z in Escherichia coli using the pT5 expression system33 and purified to apparent homogeneity by heat treatment and clearing of bacterial lysate, followed by chromatography on S-Sepharose. The protein was then concentrated and used to immunize rabbits (Spring Valley Laboratories, Woodbine, MD). The myeloperoxidase (MPO) antiserum34 was kindly provided by W. Nauseef (University of Iowa). Immunohistochemical staining. Immunohistochemical staining was performed as described.35In brief, lo5cells were cytospun onto glass slides and fixed for 30 minutes on ice in 4% paraformaldehyde/ PBS. Slides were then rinsed (and stored at 4°C) with PBT (PBS, 0.2% bovine serum albumin [BSA], 0.05% Tween-20), blocked with PBT containing 2% goat and 2% human AB+ sera, aspirated, and antisera diluted 1:500 in the same solution were then added for 1 hour. Slides were then washed and incubated in PBS/60% methanol/3% H z O for ~ 5 minutes to inactivate endogenous peroxidase. After washing, the slides were then developed using biotinylated antirabbit Ig and avidin-biotin/HRP (Vector Labs, Burlingame, CA) as described by the manufacturer. Slides were then incubated with diaminobenzene (1 mg/mL) and H202 (0.1%) for 30 minutes, washed, incubated with 20 mmol/L CuSO4/86 mmol/L NaCl for 5 minutes, washed, and counterstained with 0.001% Fast Green FCF (Sigma, St Louis, MO) in 1% acetic acid. RNA extraction and Northern blotting. Total cellular RNA was prepared using the acid guanidinium thiocyanate p r ~ c e d u r eFor .~~ Northern blotting, RNA samples ( lo7 cell equivalents) were separated on 1% agaroseiformaldehyde gels and transferred3’ to a nylon membrane (Genescreen; New England Nuclear, Boston, MA). Filters were prehybridized at 68°C in a solution3s containing 0.5 mol/L NaP04, pH 7.0, 1 mmol/L EDTA, 7% SDS, 10% dextran sulfate, and hybridized similarly after the addition of probes labeled with 32Pby random priming.39 CIEBPa, CIEBPP, C/EBPG, MPO, and mouse P5-tubulin cDNAs used for labeling have been d e s ~ r i b e d . ”Filters , ~ ~ were then washed to a stringency of 0.1 x SSC/O.5% SDS at 68°C and exposed to Kodak XAR film at -70°C. RESULTS CIEBPa CIEBPP, and CIEBPS expression during granulopoiesis in 320 C13 cells. 32D C13 cells divide with a generation time of approximately 18 hours in the presence of IL-3. Upon removal of IL-3 and exposure to G-CSF, they undergo, asynchronously, a 6- to 14-day program of differentiation, resulting in terminally differentiated granulocytes. Cell division continues, with a generation time of 24 hours, during the first half of this program, and then cell division ceases. 32D C13 cells growing in IL-3 were washed and placed in G-CSF-containing media. Differentiation was monitored by morphologic examination of the cells (Fig 1A). Uninduced cells were large myeloblasts. Several of the cells have phagocytosed dust particles. After 1 day in G-CSF, the cells diminished in size and had begun chromatin condensation. By day 4, promyelocytes predominated, with prominent primary granules. By day 6, myelocytes, bands, and PMNs were evident. Protein extracts from this culture were analyzed for the presence of CIEBPa (Fig lB), using two different antisera. One antiserum was raised against the entire CIEBPa (top panel), and the other was raised against an internal C/EBPa peptide unique to that protein (middle panel). C/EBPa, a 42-Kd protein, was present at high levels in the uninduced cells, increased approximately twofold during the first day of differentiation, was maintained for several days, and then diminished to low levels by day 6 of differentiation. Occasionally, C/EBPa resolved into closely spaced bands, perhaps alternatively modified forms (eg, Fig lB, top panel; and also see Figs 2, 3, and 5). On the other hand, these multiple bands were not evident in some blots (Fig lB, middle panel; and Fig lC, top panel). This discrepancy may result from variability between cultures and/or differences in gel conditions. The integrity of the cellular proteins in these extracts was confirmed by staining the blots with Ponceau S (Fig lB, bottom panel). Protein From www.bloodjournal.org by guest on August 11, 2017. For personal use only. ClEBP EXPRESSION IN MYELOID CELLS 1727 A 4- I "d. I Fig 1. Expression of ClEBPa in differentiating 320 C13 cells and comparison with ClEBPp and ClEBPS expression and with ClEBP expression in 3T3-Ll cells. 320 C13 cells growing in IL-3 were induced for granulocytic differentiation by exposure t o G-CSF. (A) Cell morphology was determined, by Wright-Giemsa staining of cytospins, for uninduced cells (IL-3) and for cells exposed t o G-CSF for 1,4, or 6 days (Gl, G4, or G6, respectively). (e)Total cellular protein extracts were obtained daily, and C/EBPa levels were determined by Western blotting using an antisera raised against bacterially expressed C/EBPa (top panel), or against an internal C/EBPa peptide (middle panel). MPO levels were determined using an antisera raised against human MPO (bottom panel). Protein integrity and levels in these extracts were assessed by staining one of these blots with Ponceau S before blocking and incubation with antiserum (bottom panel). (C) A set of protein extracts obtained from a second culture of induced 320 C13 cells was probed with an antiserum raised against a C-terminal ClEBPa peptide (top panel) or with antisera specific for ClEBPB or ClEBPG (middle panel). Extracts from an equivalent number of 3T3.Ll cells were also analyzed; these extracts were taken from cells induced t o form adipocytes for either 7 days (top panel) or 2 days (middle panel). Total cellular proteins were visualized by Ponceau S staining (bottom panel). Extracts from 4 x lo" cells were loaded in each lane. The positions of the molecular weight markers are indicated. 68 kd 42kd CIEBPt CEBPa 42kd 42kd CIEBPG 1 -26kd -68kd 42kd concentrations were similar (within twofold) in these extracts. Finally, the same extracts were analyzed for the level of MPO (Fig lB, bottom panel), a protein present in primary myeloid granules. The large subunit of murine MPO, migrating with an apparent molecular weight of 67 Kd, achievedits highest l&el per cell on days 3 and 4. The late decrease in MPO expression may reflect the diminished MPO messenger RNA (mRNA) expression observed in fully differentiated 32D C13 cells." The two bands detected between 25 and 27 Kd on the MPO blot represent direct binding of the secondary antibody to endogenous proteins (as will be shown in the bottom panel of Fig 3B). They were detected with seven different primary antisera and they From www.bloodjournal.org by guest on August 11, 2017. For personal use only. SCOlT ET AL serve, to some extent, as another control for protein integrity. Granulocytic differentiation was also induced in a second culture of 32D C13 cells. Aliquots were removed periodically and used to prepare total cellular protein. These extracts were analyzed for C/EBPa using a third specific antiserum, raised against a C-terminal peptide also unique to that protein (Fig lC, top panel). These extracts were also analyzed for C/EBPP and C/EBPG using antisera raised against peptides specific for those proteins (Fig lC, middle panels). For comparison, extracts from an equivalent number of induced 3T3-Ll preadipocytes were also analyzed. Again, C/EBPa was expressed at high levels early, but not late, in the granulocytic differentiation program; these high levels were similar to the maximum obtained by C/EBPa, on day 7, in induced 3T3-Ll cells. Thus, C/EBPa was detected in 32D C13 cells using three different antisera. In contrast, C/EBPP, a 31-Kd protein, increased during granulocytic differentiation of 32D C13 cells. C/EBPP achieved levels similar to the maximum obtained by this C/EBP isoform, on day 2, in induced 3T3-Ll cells. Crossreactive material located between 25 and 27 Kd again represents direct binding of the secondary antibody. Finally, C/EBPG, a 29-Kd protein, was expressed at a very low level in 32D C13 cells maintained in IL-3. When these cells were transferred to media containing G-CSF, the level of this C/EBP isoform first increased markedly and then diminished somewhat as the cells progressed to terminally differentiated granulocytes. Induced 32D C13 cells expressed C/EBPG at levels higher than the maximum found in 3T3-Ll cells. Of note, these data do not allow determination of the absolute levels of C/EBPa, C/EBPP, and C/EBPG, nor the ratios of these levels, as each antiserum has a different affinity for its corresponding epitope. In A NI P1 P2 D1 D2 NI addition, the radiographic exposure times of the various Western blots in this report have not been normalized to an internal standard. However, as extract corresponding to equal cell numbers were loaded in all lanes, comparisons are valid between lanes on any given blot. The integrity and level of intracellular proteins recovered in these extracts was again monitored by Ponceau S staining (Fig lC, bottom panel). Total cellular protein diminished approximately twofold during granulocytic differentiation. Given the modest decrease in total cellular protein in this experiment, and given the large decrease in nuclear volume during granulocytic differentiation, it is difficult to be certain that C/EBPa nuclear concentration actually diminishes in 32D C13 cells as they differentiate. This uncertainty will be addressed directly using immunohistochemicalstaining (see below). On the other hand, the nuclear concentrations of C/EBPP and C/EBPG evidently increase during granulocytic differentiation, as will be confirmed for C/EBPP immunohistochemically. CIEBPa expression during differentiation of HL-60 cells. The expression of C/EBPa was also determined in a human model of myeloid differentiation, HL-60 cells.40HL-60 cells were induced to differentiate along either the monocytic lineage with phorbol ester (phorbol myristate acetate [PMA]) or along the granulocytic lineage with dimethylsulfoxide (DMSO). Extracts were prepared on days 0,1, and 2 and analyzed for C/EBPa expression (Fig 2A, left). Total protein integrity and levels in these extracts were also assessed (Fig 2A, right). C/EBPa levels were high in uninduced, rapidly dividing HL-60 cells, and diminished markedly during both monocytic and granulocytic diffcrentiation of these cells. The diminution is more rapid than in 32D C13 cells; this difference could be accounted for if uninduced HL-60 cells are more mature than uninduced P1 ClEBPtr B ClEBPa NI Dlh 2h 4h 8h 24h Pfh 2h P2 D1 D2 I 4h 8h 24h Fig 2. Expression of ClEBPa in differentiating HL-80 cells. (A) Uninduced HL-60 cells (NI) or HL-60 cells induced for monocytic differentiation with 50 nmollL phorbol ester (PMA) for 1 (Pl) or 2 (P2) days or for granulocytic differentiation with 1.2% DMSO for 1 ( D l ) or 2 (D2) days were analyzed for ClEBPa expression (left) as in Fig 1B (top panel). By Wright's staining, NI cells were promyelocytic, P1 and P2 cells were monocytic, and D1 and D2 cells were myelocytic and metamyelocytic. Total cellular proteinswere visualized by Ponceau S staining (right). (6) A second culture was induced similarly, and C/EBPa expression determined for uninduced cells (NI)and for cells induced for 1,2, 4,s. or 24 hourswith DMSO (Dlh through 24h) or with phorbol ester (Plh through 24h). From www.bloodjournal.org by guest on August 11, 2017. For personal use only. C/EBP EXPRESSION IN MYELOID CELLS 32D C13 cells. The small increases noted on day 2 may result from the growth of PMA- and DMSO-resistant subpopulations of cells. To determine whether the observed marked decrease in CIEBPa expression is an immediate-early (within several hours) response to induction, additional HL-60 cultures were treated with either DMSO or PMA and protein extracts from hours 0, 1, 2, 4, 8, and 24 were prepared and analyzed for C/EBPa expression (Fig 2B). No decrease in C/EBPa was observed during the first 4 hours of induction, though marked reduction was again observed by 24 hours. Of note, our CIEBPP and C/EBP8 antisera detected C/EBPP or C/EBPS in murine (see below) but not in human (data not shown, A.D.F., June 1991) cell lines. This observation suggests that these antisera, raised against peptides from the murine forms of these proteins, react in a species-specific manner. Indeed, the CIEBPP epitope is not conserved between humans and CIEBPcu, CiEBPfi and CIEBPS are expressed in myelomonocytic, but not in lymphoid or elythroid, leukemia cells. To determine whether CIEBPa, C/EBPP, and CIEBPS are present in other hematopoietic cell lines, extracts from several murine leukemic cell lines were prepared and analyzed (Fig 3A). None of these C/EBP isoforms were detected in uninduced murine erythroleukemia (MEL) (erythroid lineage) cells, SP2 (B-lineage) cells, or CTLL (T-lineage) cells. A very low level of C/EBPa was evident in 8-day induced MEL cells. Two additional bands located just below C/EBPP and CIEBPs again represent nonspecific binding of the secondary antibody to endogenous proteins, as shown in the bottom panel of Fig 3A, wherein these extracts have been probed with secondary antibody alone. The degree of nonspecific binding to these proteins varied between extracts and between experiments. C/EBPa was detected in the immature myelomonocytic cell line, WEHI 274.1 (and in induced 32D C13 cells), but not in the more mature macrophage cell lines P388D1 and IC-21. This pattern is reminiscent of the diminished expression of CIEBPa observed during differentiation of 32D C13 and HL-60 cells (Figs 1 and 2). C/EBPP was expressed most highly in the more mature P388D1cells, again reminiscent of the increased expression of C/EBPP observed as 32D C13 cells differentiated into mature granulocytes (Fig 1C). Finally, C/EBPS was present in WEHI 274.1 cells, as well as in 32D C13 cells. We also examined the expression of C/EBPa in a variety of human leukemia cell lines (Fig 3B). This protein was present in all four human myelomonocytic cell lines examined (U937, HL-60, PLB-985, and KGla). However, CIEBPa was absent from both human erythroid (HEL and K562) cell lines, from the B-lineage cell line (Daudi), and from the T-lineage cell line (Molt-3). In addition, we determined the expression of C/EBPa in human myeloid and lymphoid leukemias, using cryopreserved marrow or peripheral blood samples obtained from patients at diagnosis or relapse (Table 1). Nine of 10 myeloid leukemias expressed this transcription factor, whereas C/EBPa was not detected in any of the seven lymphoid leukemias examined. 1729 C/EBPa is expressed in normal myeloid cells. To determine whether C/EBPa is present in normal myelomonocytic cells, a virtually pure population of developing myeloid cells was isolated from human marrow, extracted, and analyzed for C/EBPa by Western blotting (Fig 3C). CIEBPa was detected in these cells. To further examine the lineage-specific expression of CIEBPa, cytospins of human marrow were incubated with either normal rabbit serum or C/EBPa antiserum, and bound antibodies were detected immunocytochemically (Fig 4a and b). The predominant CIEBPa staining was in cells with eccentric, kidney-shaped nuclei, representing fairly mature granulocytic cells. Very few of the cells with round nuclei (the erythroid and lymphoid cells) showed specific staining. Interestingly, all of the fully mature PMNs (arrow, lower right) and some of the small band forms stained at background levels. This staining pattern suggests that there is diminished expression of CIEBPa in these most mature granulocytic cells. As will be shown, immunohistochemical staining of induced 32D C13 cells also demonstrated decreased CiEBPa levels in the nuclei of these mature cells. CIEBPa and C/EBPpare present in the nuclei of 3.20 C13 cells. One mechanism that might allow dividing myeloblasts to tolerate high levels of CIEBPa would be to sequester this protein in the cytoplasm, as can occur for C/EBPP.41 As the majority of myeloid cells in human marrow (such as in Fig 4b) are more mature, nondividing cells, we addressed this possibility using 32D C13 cells. Cytospins of uninduced, 4-day- and 8-day-induced 32D C13 cells were stained with either normal rabbit serum (Fig 4c and data not shown), C/EBPa antiserum (Fig 4d through f), or C/EBPP antiserum (Fig 4g through i). Bound antibodies were again detected immunocytochemically. Rabbit serum gave only light staining. Both C/EBPa and C/EBPP were detected only in the nucleus of positively staining cells. In this experiment, the 32D C13 cells differentiated at a slower rate than those shown in Fig 1; PMNs first became evident on day 8. The 32D C13 cells manifested significant variability in their rate of differentiation, perhaps accounting for quantitative differences in C/EBP isoform, MPO, and total protein levels at particular points in time between experiments. Nevertheless, a consistent qualitative temporal pattern of C/EBP isoform expression was evident in these cells, as will be discussed. Although quantification of protein expression is imprecise using immunohistochemical staining, large changes in staining intensity are qualitatively meaningful. Interestingly, although uninduced and 4-dayinduced 32D C13 cells stained similarly for CIEBPa (Fig 4d and e), by day 8 all of the PMNs and some of the small band forms stained only at background level. These results are consistent with the diminishing average level of C/EBPa per cell detected by Western blotting as 32D C13 cells differentiate (Fig 1B). The presence of large, immature cells on day 8, which show heavy staining for C/EBPa, is a consequence of the slower maturation of this culture. CIEBPP staining increased steadily as the cells matured (Fig 4g through i), again consistent with the results of Western blotting (Fig 1C). From www.bloodjournal.org by guest on August 11, 2017. For personal use only. SCOTT ET AL 1730 5 A 42kd- C/EBPa Fig 3. Expression of C/EBPa. CIEBPP, and C/EBPS in murine cell lines, and of ClEBPa in human cell lines and in normal human myeloid cells. (A) Protein extracts from the murine cell lines MEL (erythroid), SP2 (6-lineage), CTLL (T-lineage), IC-21 and P338D1 (macrophage), and WEH1274.1 (myelomonoblastic) and 32D C13 induced for 6 days with G-CSF (32D-G6) were analyzed for C/EBPaand, in some cases, C/EBPB and C/EBPS expression by Westem blotting (top three panels). MEL cells were either uninduced (MEL-NI) or induced for erythroid differentiation by 8 days of exposure t o 2% DMSO (MEL-DO). Two bands were detected between 25 and 27 Kd in many lanes due t o nonspecific binding of the secondary antibody t o endogenous proteins. This is shown in the bottom panel wherein several of these extracts were hybridized only with this second antibody. CRM, crossreactive material. (9) Protein extracts from the human leukemia cell lines U937, HL-60, KGla and PLB-985 (myelomonoblastic), HEL and K562 (erythroid), Daudi (6-lineage), and Molt-3 (T-lineage) were analyzed similarly for C/EBPa expression. (C) Myeloid cells were isolated from human marrow by adsorption onto immunomagnetic beads t o which had been bound monoclonal anti-CD15 antibodies. Protein extracts from these cells (CD15) and from beads with bound antibodies alone (Ab) were again analyzed for C/EBPa expression. The CD15- cells were a viltually pure population of myelomonocytic cells, with approximately 4% myeloblasts, 8% promyelocytes, 28% myelocytes, 40% metamyelocytes, 16% bands,and4% PMNs. ClEBPB 26kd- C/EBPG 26kd. 26kd- -- CRM _. v) " Z I 9 B 5 c n a b 0 I C/EBPa Regulation of C / E B P s CIEBPP, and CIEBPG levels duringgranulopoiesis. To address the regulation of expression of C/EBP family members during granulopoiesis, we probed Northern blots containing total cellular RNA from an equal number of uninduced and induced 32D C13 cells with radiolabeled cDNAs specific for either C/EBPa, C/EBPP, C/EBPG, MPO, or tubulin (Fig SA). For comparison, an aliquot of cells from the same cultures used to prepare the RNAs was used to prepare total cellular proteins, and these cells were analyzed for C/EBPa, CIEBPP, and C/EBPG, MPO, and total protein expression (Fig SB). Monitoringof cell morphology showed that PMNs From www.bloodjournal.org by guest on August 11, 2017. For personal use only. 1731 C/EBP EXPRESSION IN MYELOID CELLS Table 1. C/EBPu Expression in Human Leukemias Patient No. 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 - CIEBPu Diagnosis AM L AM L AM L AM L AM L AM L AM L AM L AML AML* BALL B-ALL B-ALL B-ALL B-ALL T-ALL T-ALL ~_____ Surface Phenotype DR, CD11b, 15,33 CD33 DR, CD11b, 15,33 DR, CD15.34 DR, CD15,33,34 DR, CD7. 15,34 DR, CD7 Unknown DR,CDllb, 15 DR, CD7.33,34 DR, CD19,22,34 DR, CD10,19,22 CD10,19,22,34 DR, CD19,22,34 DR, CD19 CD2. 5, 7,34 CD2.5, 7,8 Expression ++++ ++++ ++++ +++ ++ ++ + + + - ~ Marrow or peripheral blood cells were obtained from patients at diagnosis or relapse and stored in liquid nitrogen. Surface phenotypes, which help establish the diagnoses, were determined by FACS analysis26,28before freezing. Cells were then thawed and analyzed for C/EBPa expression by Western blotting. Abbreviations: AML, acute myeloid leukemia; ALL, acute lymphocytic leukemia. 'Relapse sample from a patient initially diagnosed as ALL. first became evident at day 10 in this experiment. Total protein diminished twofold to threefold during granulocytic differentiation, similar to the experiment described in Fig 1C. C/EBPa, CIEBPP, and C/EBPS mRNA and protein levels correlated fairly well throughout 32D C13 cell differentiation, suggesting predominant pretranslational regulation of these three C/EBP isoforms. MPO mRNA was induced by day 4, and tubulin mRNA diminished somewhat as the 32D C13 cell matured. MPO protein levels paralleled MPO mRNA levels and remained high on day 10 of induction. Of note, MPO protein did not show a diminution as it had in Fig lA, perhaps reflecting the fact that the 10-day-induced culture in this experiment was not as mature (10% PMNs) as the 6-day-induced culture in the former experiment (50% PMNs). DISCUSSION Whereas some transcription factors, such as M ~ o and D ~ O ~ t - 2 appear , ~ ~ to be truly restricted to a single tissue, others, such as GATA-1,44,45and HNF-1,46,47have been found in several cell types. CIEBPa, C/EBPP, and C/EBPG fall into this latter group. We have shown herein that these three C/EBP isoforms are highly expressed in myelomonocytic cells. This finding is consistent with the recent description of C/EBP DNA-binding activity in chicken myeloid, but not erythroid or lymphoid, cells.48 C/EBPs are also expressed in hepatocytes and adipocytes, and perhaps in lung and intestinal tissues as well, but have not been detected in kidney, brain, heart, testis, or spleen.11J2J6 Although we also did not detect C/EBPa in lymphoid or erythroid cells, a subset of these cells may well express this protein. Indeed, the mRNA for C/EBPy is expressed most highly in B lymphocyte^.^^ C/EBPa has been implicated in the regulation of both hepatic-~pecific~~J~ and adipocyte-specific gene^.'^,'^ The presence of several DNA-binding proteins capable of binding the C/EBP sites within these genes (eg, see Mueller et aP9) has made it difficult to ascertain which protein(s) actually mediates transcription through these DNA elements. However, recent experiments using antisense RNA expression have directly implicated C/EBPa as a regulator of a d i p o g e n e s i ~ . ~ ~ , ~ ~ The inability of preadipocytes to divide when overexpressing C/EBPLX~~ suggests that this transcription factor might also activate genes that inhibit cell division. Expression of C/EBPa even overcame the proliferative and antidifferentiative effects of overexpressed Myc in preadipocytes, and induced cessation of cell division and differentiati~n.~~ In contrast, C/EBPa's expression pattern in differentiating myeloid cells suggests that it may even activate genes, such as those encoding protein hormones, which stimulate cell division in these cells. For example, in chicken myeloid cells,@a C/EBP-family member has been implicated in the activation of the gene encoding chicken myelomonocytic growth factor (cMGF), a homologue of human G-CSF.51 Moreover, the human G-CSF promoter contains a C/EBP binding site active in macrophage^^^ that binds C/EBPf3.53 C/EBPa, along with C/EBPp and CIEBPG, is one of only a few tissue-restricted transcription factors known to be present in immature myeloid cells (see Hromas et aIs4for a recent review of hematopoietic transcription factors). C-myb is predominantly expressed in hematopoietic cells,55 can induce myeloid-specific and is required for fetal hematopoiesi~.~~ CCAAT-displacement protein is a ubiquitous repressor that participates in the regulation of the myeloid-specific gp91-phox gene.58 MZF-1, a zinc-finger protein that plays a role in granulocyte de~elopment,5~ is only expressed in later-stage granulocytic c e k m The activity of a helix-loop-helix protein(s) has recently been shown to be required for the early stages of 32D C13 cell differentiatione61 Three models of how C/EBPa might participate in the activation of different sets of genes in myeloid, adipocyte, and hepatic cells are: (1) a different set of C/EBPa's several trans-activating domains32might be active in each tissue; (2) ~ CiEBPa might dimerize with a different subset of C/EBP family members in each tissue; or (3) C/EBPa might dimerize with a predominant partner, but interact with additional, tissue-restricted transcription factors in each lineage. The ability of C/EBP family members to heterodimerize readily in ~ i t r o l suggests ~ . ~ ~ that they could do so in vivo as well. The expression pattern of C/EBPa, C/EBPP, and CIEBPS in differentiating preadipocytes suggests that PP, 66, and PS dimers could predominate early and that aa homodimers could predominate late in adip0genesis.l' Determination of the pattern of C/EBP isoform expression in maturing 32D C13 myeloblasts is confounded somewhat From www.bloodjournal.org by guest on August 11, 2017. For personal use only. S C O T ET AL 1732 Fig 4. In situ antibody staining of human manow and of 32D C13 cells. Cytospins of human marrow were stained with either normal rabbit serum (a) or with rabbit anti-C/EBPa protein antiserum (b). Bound antibodieswere detected immunocytochemically and the cells were then counterstained with Fast Green FCF. The arrow at the lower right in (b) indicates a PMN. (c) 32D C13 cells were Induced with GCSF for 4 days and stained with normal rabbit serum. (d through f l 320 C13 cells were induced for 0, 4, or 8 days and stained with a n t i 4 1EBPa protein antiserum. (g through i) 32D C13 cells were induced for 0, 4, or 8 days and stained with anti-C/EBPp antiserum. Photomicrographs are at 40x. by the variability of this cell line. Maturation of an entire culture to PMNs required anywhere from 8 to 14 days. The levels of C/EBPu and C/EBPP in undinduced 32D C13 cells was also variable, as were the quantitative changes in the expression levels of these isoforms during granulocytic differentiation. These quantitative differences could be partly explained if the maturation level of uninduced cells is itself variable. More “mature” uninduced 32D C13 cells B A 113 ClEBPa 04 G7 I13 G10 ’ MPC 04 113 04 07 010 67 GI0 Y,s)rr, CEBPa 113 04 GI G10 MU TUl ClEBPG Fig 5. CIEBPa. CIEBPp, and CIEBP8 mRNA expression In d l f f e r e n t l ~ n g32D C13 wlls and comparison with the levels of the corresponding proteins. 32D C13 cells growing In IL-3 wore Induced for granulocytic differentiation by exposure t o G-CSF. Initially, and after 4,7, or 10 d a y , aliquots were removed and used t o prepare both total cellular RNA and protein. (A) C/EBPa, CIEBPp, C/EBPG, MPO, and tubulin (Tub) mRNA expression was determined by Northern blotting. RNA from lo7 cells were loaded In each lane. Blots were stripped between probes by incubation In boiling water for 5 minutes. (B) C/EBPcr. C/EBPp. C/EBPS, and MPO protein expression was determined by Western blotting. Total cellular proteins were visualized by Ponceau S staining (bottom right). From www.bloodjournal.org by guest on August 11, 2017. For personal use only. 1733 C/EBP EXPRESSION IN MYELOID CELLS 100 CIEBPa 50 2 u > u CIEBPP A <r CIEBPG x 2I- 100 CIEBPP 50 CIEBPG 2 u w n CIEBPa NI TD TIME OF DIFFERENTIATION Fig 6. Temporal pattern of C/EBPu, C/EBPp, and C/EBPG expression in differentiating 3T3-Ll preadipocytes“ and in differentiating 32D C13 myeloblasts. The absolute levels of these three proteins, within each cell type, are unknown. The temporal pattern for each C/EBP isoform in 32D C13 cells can only be considered an approximation given the variability of these cells in culture. NI, noninduced; TD, terminally differentiated.Arrows indicatethe approximatetime when proliferationceases in each pathway. would differentiate into PMNs faster (eg, Figs 1and 2) than would less “mature” uninduced cells (eg, Figs 4 and 5). Perhaps the levels of C/EBPa and C/EBPP are lower in the more “immature,” compared with the more “mature” uninduced 32D C13 cells. Despite these caveats, a consistent qualitative temporal pattern of C/EBP isoform expression in differentiating 32D C13 myeloblasts emerges from the data presented. The nuclear concentration of C/EBPa first increases, being expressed maximally in cells that retain the capacity for cell division, and eventually diminishes, at least in some of the most mature cells; C/EBPP increases steadily throughout; and C/EBPG increases markedly early during induction and then remains fairly constant, or may diminish mildly. This pattern of C/EBP isoform expression in granulocytic differentiation is contrasted with that described for adipocyte developmentll in Fig 6. Our results with differentiating myeloblasts therefore suggest that act, PP, and ctP dimers could predominate in immature myelomonocytic cells; that all possible combinations could exist at intermediate stages of differentiation; and that Pp, 66, and PS dimers could predominate in terminally differentiated cells. In addition, other C/EBP family members, such as C/EBPy, may be present in these cell types, allowing for the formation of additional heterodimers. Because the C/EBP family appears likely to play a role in regulating myelopoiesis, we are also interested in determining how the levels of these transcription factors are themselves regulated. Our data suggest that regulation of mRNA production or turnover could account for the changes in C/EBPa, C/EBPP, and C/EBPG levels observed during granulopoiesis. Finally, our observation of CiEBPa expression in myeloid, but not lymphocytic, human leukemias has potential clinical utility, as assessing C/EBPa expression may aid in diagnosis, and inhibition of CIEBPa expression in myeloid leukemia cells may prove therapeutic. ACKNOWLEDGMENT We thank Steven McKnight and Zhaodan Cao for their encouragement and generous provision of antisera and CIEBP cDNAs. We also thank W. Nauseef for MPO antiserum; J. Suzow, S. Amin, J. Hebb, and T. Trischmann for technical assistance; M. Kastan for guidance on antibody staining; and A. Hess for CTLL cells. REFERE NCES 1. Graves BJ, Johnson PF, McKnight S L Homologous recognition of a promoter domain common to the MSV LTR and the HSV tk gene. Cell 44:565,1986 2. Johnson PF, Landschulz WH, Graves BJ, McKnight S L Identification of a rat liver nuclear protein that binds the enhancer core element of three animal viruses. Genes Dev 1:133,1987 3. Landschulz WH, Johnson PF, Adashi EY, Graves BJ, McKnight S L Isolation of a recombinant copy of the gene encoding CIEBP. Genes Dev 2786,1988 4. Landschulz WH, Johnson PF, McKnight SL: The leucine zipper: A hypothetical structure common to a new class of DNA binding proteins. Science 240:1759,1988 5. Landschulz WH, Johnson PF, McKnight S L The DNA binding domain of the rat liver protein C/EBP is bipartite. Science 243:1681,1989 6. Vinson CR, Sigler PB, McKnight SL: Scissors-grip model for DNA recognition by a family of leucine zipper proteins. Science 246:911, 1989 7. Akira S, Isshiki H, Suguita T, Tanabe 0, Kinoshita S, Nishio Y, Nakajima T, Hirano T, Kishimoto T: A nuclear factor for IL-6 expression (NF-IL6) is a member of a CIEBP family. EMBO J 9:1897, 1990 8. Poli V, Mancini FP, Cortese R: IL-6DBP, a nuclear protein involved in interleukin-6 signal transduction, defines a new family of leucine zipper proteins related to CIEBP. Cell 63:643, 1990 9. Descombes P, Chojkier M, Lichsteiner S, Falvey E, Schibler U: LAP, a novel member of the CIEBP gene family, encodes a liver-enriched transcriptional activator protein. Genes Dev 4: 1541, 1990 10. Chang C-J, Chen T-T, Lei H-Y, Chen D-S, Lee S - C Molecular cloning of a transcription factor, AGPIEBP, that belongs to members of the C/EBP family. Mol Cell Biol 10:6442, 1990 11. Cao Z, Umek RM, McKnight S L Regulated expression of three CIEBP isoforms during adipose conversion of 3T3-Ll cells. Genes Dev 5:1538,1991 12. Williams SC, Cantwell CA, Johnson PF: A family of CIEBP- From www.bloodjournal.org by guest on August 11, 2017. For personal use only. 1734 related proteins capable of forming covalently linked leucine zipper dimers in vitro. Genes Dev 5:1553, 1991 13. Roman C, Platero JS, Shuman JD, Calame K: IgiEBP-1: A ubiquitously expressed immunoglobulin enhancer binding protein that is similar to CiEBP and heterodimerizes with C/EBP. Genes Dev 4:1404,1990 14. Friedman AD, Landschulz WH, McKnight SL: CCAATi enhancer binding protein activates the promoter of the serum albumin gene in cultured hepatoma cells. Genes Dev 3:1314, 1989 15. Park EA, Roesler WJ, Liu J, Klemm DJ, Gurney AL, Thatcher JD, Shuman JD, Friedman AD, Hanson RW: The role of CAAT/enhancer binding protein in the transcriptional regulation of the gene for phosphoenolpyruvate carboxykinase (GTP). Mol Cell Biol10:6264,1990 16. Birkenmeier EH, Gwynn B, Howard S, Jerry J, Gordon JI, Landschulz WH, McKnight S L Tissue-specificexpression, developmental regulation, and genetic mapping of the gene encoding CCAAT/enhancer binding protein. Genes Dev 3:1146,1989 17. Christy RJ, Yang VW, Ntambi JM, Geiman DE, Landshulz WH, Friedman AD, Nakabeppu Y, Kelly TJ, Lane MD: Differentiation induced gene expression in 3T3-Ll preadipocytes: CCAAT/ enhancer binding protein interacts with and activates the promoters of two adipocyte-specific genes. Genes Dev 3:1323,1989 18. Kaestner KH, Christy RJ, Lane MD: Mouse insulinresponsive glucose transporter gene: Characterization of the gene and trans-activation by the CCAATienhancer binding protein. Proc Natl Acad Sci USA 87:251,1990 19. Samuelsson L, Stromberg K, Vikman K, Bjursell G, Enerback S: The CCAATienhancer binding protein and its role in adipocyte differentiation: Evidence for direct involvement in terminal adipocyte development. EMBO J 10:3787, 1991 20. Lin FT, Lane D: Antisense CCAAT/enhancer-binding protein RNA suppresses coordinate gene expression and triglyceride accumulation during differentiation of 3T3-Ll preadipocytes. Genes Dev 6533,1992 21. Lamb P, McKnight SL: Diversity and specificity in transcriptional regulation: The benefits of heterotypic dimerisation. Trends Biochem Sci 16:417,1991 22. Umek RM, Friedman AD, McKnight SL: CCAATienhancer binding protein: A component of a differentiation switch. Science 251:288,1991 23. Valtieri M, Tweardy DJ, Caracciolo D, Johnson K, Mavilio F, Altman S, Snatoli D, Rovera G: Regulation of proliferative and differentiative responses in a murine progenitor cell line. J Immuno1 138:3829,1987 24. Friedman AD, Krieder BL, Venturelli D, Rovera G: Transcriptional regulation of two myeloid-specific genes, myeloperoxidase and lactoferrin, during differentiation of the murine cell line 32D C13. Blood 78:2426,1991 25. Warner NL, Moore MAS, Metcalf D: A transplantable myelomonocytic leukemia in Balb/c mice: Cytology, karyotype and muramidase content. J Natl Cancer Inst 43:963,1969 26. Strauss LC, Stuart RK, Civin CI: Antigenic analysis of hematopoiesis: I. Expression of the My-1 granulocytic surface antigen on human marrow cells and leukemic cell lines. Blood 61:1222, 1983 27. Strauss LC, Trischmann TM, Rowley SD, Wiley JM, Civin CI: Selection of normal human hematopoietic stem cells for bone marrow transplantation using immunomagnetic microspheres and CD34 antibody. Am J Pediatr Hematol Oncol 13:217,1991 28. Gore SD, Kastan MB, Civin CI: Normal human bone marrow precursors that express TdT includes T-cell precursors and possible lymphoid stem cells. Blood 77:1681, 1991 29. Civin CI, Mirro J, Banquerigo ML: A new myeloid specific SCOTT ET AL antigen identified by a mouse monoclonal antibody. Blood 57:842, 1981 30. Laemmli UD: Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680, 1970 31. Towbin H, Staehelin T, Gerdon J: Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: Procedure and some applications. Proc Natl Acad Sci USA 76:4350,1979 32. Friedman AD, McKnight S L Identification of two polypeptide segments of CCAATienhancer-binding protein required for transcriptional activation of the serum albumin gene. Genes Dev 4:1416, 1990 33. Shuman JD, Vinson CR, McKnight S L Evidence of changes in protease sensitivity and subunit exchange rate on DNA binding by C/EBP. Science 249:771,1990 34. Nauseef WM, Metcalf JA, Root RK: Role of myeloperoxidase in the respiratory burst of human neutrophils. Blood 61:483, 1983 35. Kastan MB, Stone KD, Civin CI: Nuclear oncoprotein expression as a function of lineage, differentiation stage, and proliferative status of normal human hematopoietic cells. Blood 74:1517, 1989 36. Chomczynski P, Sacchi N: Single-step method of isolation of RNA by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal Biochem 162:156, 1987 37. Maniatis T, Fritsch EF, Sambrook J: Molecular Cloning: A Laboratory Manual (ed 2). Cold Spring Harbor, NY, Cold Spring Harbor Laboratory, 1989 38. Church GM, Gilbert W: Genomic sequencing. Proc Natl Acad Sci USA 81:1991,1984 39. Feinberg AP, Vogelstein B: A technique for radiolabeling DNA restriction endonuclease fragments to high specific activity. Anal Biochem 132:6,1983 40. Collins SJ, Gallo RC, Gallagher RE: Continuous growth and differentiation of human myeloid leukemia cells in suspension culture. Nature 270:347,1977 41. Metz R, Ziff E: CAMP stimulates the C/EBP-related transcription factor rNFIL-6 to translocate to the nucleus and induce c-fos transcription. Genes Dev 5:1754,1991 42. Davis RL, Weintraub H, Lassar AB: Expression of a single transfected cDNA clone converts fibroblasts to myoblasts. Cell 51:987, 1987 43. Clerc RG, Corcoran LM, LeBowitz JH, Baltimore D, Sharp PA: The B-cell specific Oct-2 protein contains POU box- and homeo-box type domains. Genes Dev 2:1570,1988 44. Tsai SF, Martin DI, Zon L, D’Andrea AD, Wong GG, Orkin SH: Cloning of cDNA for the major DNA-binding protein of the erythroid lineage through expression in mammalian cells. Nature 339:446, 1989 45. Martin DI, Zon LI, Mutter G, Orkin SH: Expression of an erythroid transcription factor in megakaryocytic and mast cell lineages. Nature 344:444, 1990 46. Courtois G, Morgan JG, Campbell LA, Fourel G, Crabtree G: Interaction of a liver-specific nuclear factor with the fibrinogen and al-antitrypsin promoters. Science 238:688, 1987 47. Baumhueter S, Mendel DB, Conley PB, Kuo CJ,Turk C, Graves MK, Edwards CA, Courtois G, Crabtree GR: HNF-1 shares three sequence motifs with the POU domain proteins and is identical to LF-B1 and APF. Genes Dev 4:372,1990 48. Sterneck E, Muller C, Katz S, Leutz A. Autocrine growth induced by kinase type oncogenes in myeloid cells requires AP-1 and NF-M, a myeloid specific, CiEBP-like factor. EMBO J 11:115, 1992 49. Mueller CR, Maire P, Schibler U: DBP, a liver-enriched transcriptional activator, is expressed late in ontogeney and its From www.bloodjournal.org by guest on August 11, 2017. For personal use only. C/EBP EXPRESSION IN MYELOID CELLS tissue specificity is determined posttranscriptionally. Cell 61:279, 1990 50. Freytag SO, Geddes TJ: Reciprocal regulation of adipogenesis by Myc and C/EBPa. Science 256:379,1992 51. Leutz A, Beug H, Graf T Purification and characterization of cMGF, a novel chicken myelomonocytic growth factor. EMBO J 3:3191,1984 52. Nishizawa M, Nagata S: Regulatory elements responsible for inducible expression of the granulocyte colony-stimulating factor gene in macrophages. Mol Cell Biol10:2002,1990 53. Natsuka S, Akira S, Nishio Y ,Hashimoto S, Sugita T, Isshiki H, Kishimoto T: Macrophage differentiation-specific expression of NF-IM, a transcription factor for interleukin-6. Blood 79:460,1992 54. Hromas R, Zon L, Friedman AD: Hematopoietic transcription factors and the origins of leukemia. Crit Rev Hematol Oncol 12167,1992 55. Sheiness D, Gardinier M: Expression of a proto-oncogene (proto-myb) in hematopoietic tissues of mice. Mol Cell Biol4:1206, 1984 56. Ness SA, Markness A, Graf T The v-myb oncogene product 1735 binds to and activates the promyelocyte-specific mim-1 gene. Cell 59:1115,1989 57. Mucenski ML, McClain K, Kier AB, Swerdlow SH, Schreiner CM, Miller TA, Pietryga DW, Scott WJ, Potter SS: A functional c-myb gene is required for normal murine fetal hepatic hematopoiesis. Cell 65:677, 1991 58. Skalnick DG, Strauss EC, Orkin SH: CCAAT-displacement protein as a repressor of the myelomonocytic-specific gp91-phox gene promoter. J Biol Chem 266:16736,1991 59. Bavisotto L, Kaushansky K, Lin N, Hromas R: Anti-sense oligonucleotides from the stage-specific myeloid zinc finger gene, MZF-1, inhibits granulopoiesis in vitro. J Exp Med 174:1097, 1991 60. Hromas R, Collins SJ, Hickstein D, Raskind W, Deaven LL, O’Hara P, Hagen FS, Kaushansky K A retinoic acid-responsive human zinc finger gene, MZF-1, preferentially expressed in myeloid cells. J Biol Chem 266:14183,1991 61. Kreider BL, Benzara R, Rovera G, Kadesch T: Inhibition of myeloid differentiation by the helix-loop-helix protein Id. Science 255:1700,1992 From www.bloodjournal.org by guest on August 11, 2017. For personal use only. 1992 80: 1725-1735 A novel temporal expression pattern of three C/EBP family members in differentiating myelomonocytic cells LM Scott, CI Civin, P Rorth and AD Friedman Updated information and services can be found at: http://www.bloodjournal.org/content/80/7/1725.full.html Articles on similar topics can be found in the following Blood collections Information about reproducing this article in parts or in its entirety may be found online at: http://www.bloodjournal.org/site/misc/rights.xhtml#repub_requests Information about ordering reprints may be found online at: http://www.bloodjournal.org/site/misc/rights.xhtml#reprints Information about subscriptions and ASH membership may be found online at: http://www.bloodjournal.org/site/subscriptions/index.xhtml Blood (print ISSN 0006-4971, online ISSN 1528-0020), is published weekly by the American Society of Hematology, 2021 L St, NW, Suite 900, Washington DC 20036. Copyright 2011 by The American Society of Hematology; all rights reserved.