* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download A bioinformatika elméleti alapjai 4
Survey
Document related concepts
List of types of proteins wikipedia , lookup
Gene regulatory network wikipedia , lookup
Bottromycin wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Non-coding DNA wikipedia , lookup
Whole genome sequencing wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Genome evolution wikipedia , lookup
Molecular evolution wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Protein domain wikipedia , lookup
Protein structure prediction wikipedia , lookup
Transcript
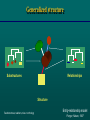
This course is sponsored by the International Centre for Genetic Engineering and Biotechnology Welcome Bioinformatics: Computational approaches to biological information Organizer: Sándor Pongor Leonardo Marino-Ramirez, Christoph W. Sensen, Laurent Falquet, Sándor Pongor Teaching staff: Stefan Grabuschnig, János Juhász Secretariat: Elisabetta Lippolis Chiara Alberti Giorgia Danelon Computer system manager: Dario Palmisano Diego Soldano Trieste, 26-30 June, 2017 Computational approaches to biological information Trieste, May 23 - 27, 2016 Theoretical intro: Sándor Pongor Sequence database searching, theory and practice (Leonardo Marino) Multiple alignment, tree building (Christoph Sensen) Next Generation Sequencing (Laurent Falquet) Genome annotation (Christoph Sensen) Chip-seq, RNA-seq (Leonardo Marino-Ramirez) BIOINFORMATICS INFORMATICS Model, description and visualization The subjects: Molecular structures MARTKQTARK STGGKAPRKQ LATKAARKSA Sequences CIPKWNRCGPKMDGVPCCEPYTCTSDYYGNCS Extended sequences (e.g. disulphide-topologies) Diagrams (hydrophobicity plots, helical circles) Domain-cartoons (sec. str. cartoons) 3D structures 3D cartoons Core data-types tassfvvswvsasdtvsgfrvey elseegdepqyldlpstatsvni pdllpgrkytvnvyeiseegeqn lilstsqttapdappdptvdqvd dtsivvrwsrprapitgyrivys psvegsstelnlpetansvtlsd lqpgvqynitiyaveenqestpv fiqqettgvprsdkvppprdlqf vevtdvkitimwtppespvtgyr vdvipvnlpgehgqrlpvsrntf aevtglspgvtyhfkvfavnqgr eskpltaqqatkldaptnlqfin etdttvivtwtpprarivgyrlt vgltrggqpkqynvgpaasqypl rnlqpgseyavslvavkgnqqsp rvtgvfttlqplgsiphyntevt ettivitwtpaprigfklgvrps qggeaprevtsesgsivvsgltp gveyvytisvlrdgqerdapivk SEQUENCES 3-D GENOMES TEXT A structural model Relationships Substructures Structure Entity-relationship model Pongor, Nature, 1987 Core data groups -GAA- CONSENSUS STRUCTURES TREES NETWORKS A structural model Relationships Substructures Structure Entity-relationship model Pongor, Nature, 1987 Generalized structure Relationships Substructures Structure Susbstructures, relations, rules = onthology Entity-relationship model Pongor, Nature, 1987 Core operations Simplification + annotation Comparison Aggregation Annotation: providing sg with notes, adding notes to sg SEQUENCES Model: Chemical structure Description: Series of characters Simplified and/or extended visualization IFPPVPGP Domain A Domain B SEQUENCES Domain A Domain B 001-200 DOMAIN PROTEASE A 205-230 DOMAIN TRANSMEMBRANE 250-350 DOMAIN SIGNAL BINDING TABULAR DESCRIPTION: FEATURE TABLE, PTT TABLE Leonardo Marino ANNOTATING GENOME SEQUENCES Gene 1 Christoph Sensen Gene 2 Genome annotation .ptt table RNAseq, CHIPseq: MAPPING READS TO REFERENCE GENES OR GENOMES ~ NUMERICAL ANNOTATION Leonardo Marino SIMPLIFICATION OF 3D STRUCTURES Model: 3D chemical structures Description: 3D coordinates Simplified and/or extended visualization (xi, yi, zi)n Domain A Some molecules are more equal then others… …”This figure is purely diagrammatic. The two ribbons symbolize the the phosphate-sugar chains, and the horizontal rods the pairs of the bases holding the chains together. The vertical line marks the fibre axis” Protein visualization Input: atomic 3D coordinates and sequence. Structures As Database Records Identification Name of protein Organism Function Cross-references ... Domain structure Sec. structure Disulphides …. ANNOTATIONS CIPKWNRCGPKMDGVPCCEPYTCTSDYYGNC Sequence (structure) qfinetdttvivtwtpprarivgyrltvgllseeg depqyldlpstatsvnipdllpgrkytvnvyeise egeqnlilstsqttapdappdptvdqvddtsivvr wsrprapitgyrivyspsvegsstelnlpetansv tlsdlqpgvqynitiyaveenqestpvfiqqettg vprsdkvppprdlqfvevtdvkitimwtppespvt gyrvdvipvnlpgehgqrlpvsrntfaevtglspg vtyhfkv Database record, fields SEQUENCE OR STRUCTURE Core operations 2 Comparison The concept of similarity I Shared parts Shared context ...easier if modular The concept of similarity II …Easy for humans, hard for computers Similarity in bioinformatics: Important properties Quantitative: we need a similarity score and a method to calculate significance Alignment (finding matches between sequences, between structures, etc.) Aggregation (adding small similarities together). Similarity scores and significance: A score is a number. Higy score is high similarity. No inherent „scale”. A score can be scaled if we know the probabilities of random similarities. This gives significance: what is the probability of finding this number by chance? The smaller the better Alignment Finding the best match between two sequences Finding exact matches is easy. In biology we need approximate matches, and that is difficult. The result: 1)A similarity score (number), with significance 2) An alignment pattern RGD RGD...W Substructure identity ~ similarity ”The similarity of objects can be best described as partial identities of components and relationships Erich Goldmeier, The similarity of perceived forms, 1936 Which alignment is better? The one with a higher score The one with a „nicer” motif.. Core Operations 3 Aggregation Why do we need aggregation? Biological objects are large and complex (genomes, proteomes, metagenomes, pathway data, etc.) Often, measuring instruments can only collect data on small pieces (next generation sequencing reads, peptide spectra in proteomics) Computational analysis of small fragments is accurate. Why do we need aggregation? (in other words) Only simple objects can be easily located by similarity, say we easily find a 3 amino acid motif in a sequence or in a 3D structure. Unfortunately, most objects in bioinformatics are COMPLICATED, like genomes, proteomes, metagenomes, pathways, even ordinary protein or gene sequences. There is one general trick: We divide a complex object into simple parts (like characteristic motifs), identify individual parts by simple numerical means, and then AGGREGATE the results. Not elegant, but works, even with very complex problems. Aggregating local sequence similarities Sequence 1 Sequence 2 Are these two sequences related by evolution? (are they homologous?) Only probabilistic answers... We need aggregate scores, i.e. probabilities for finding combinations by chance... Leonardo Marino BLAST Examples for aggregation in bioinformatics Single proteins, genes: constructing protein/gene similarity from local similarities (BLAST) Inferring homolgy. Proteomics: Constructing protein similarities from peptide fragment similarities. Inferring protein presence. Genomics1: Aggregating a long sequence from short reads (next generation sequencing). Inferring a genome. Genomics2: Putting protein similarities together into pathways. Metagenomics: Inferring a microbial community from species similarities. The human mind is good at aggregating noisy signals Edgar Rubin’s vase (~1915, Copenhagen) Kanizsa’s Triangle (~1955, Trieste) The human mind is good at aggregating noisy signals according to structures Contour recognition principles In bioinformatics, computers do this in an abstract space of data, and without human intuition. Filtering, search space reduction is useful when designing bioinformatics tools. Psychology of vision. SUMMARY: Core data types tassfvvswvsasdtvsgfrvey elseegdepqyldlpstatsvni pdllpgrkytvnvyeiseegeqn lilstsqttapdappdptvdqvd dtsivvrwsrprapitgyrivys psvegsstelnlpetansvtlsd lqpgvqynitiyaveenqestpv fiqqettgvprsdkvppprdlqf vevtdvkitimwtppespvtgyr vdvipvnlpgehgqrlpvsrntf aevtglspgvtyhfkvfavnqgr eskpltaqqatkldaptnlqfin etdttvivtwtpprarivgyrlt vgltrggqpkqynvgpaasqypl rnlqpgseyavslvavkgnqqsp rvtgvfttlqplgsiphyntevt ettivitwtpaprigfklgvrps qggeaprevtsesgsivvsgltp gveyvytisvlrdgqerdapivk A structural model Relationships Substructures Structure Entity-relationship model Pongor, Nature, 1987 SUMMARY: Core operations Simplification + annotation Comparison Aggregation Models are human constructs... THIS IS NOT A PIPE! Models are human constructs... THIS IS NOT A MOLECULE Bioinformatics: Computational approaches to biological information Organizer: Sándor Pongor Leonardo Marino-Ramirez, Christoph W. Sensen, Laurent Falquet, Sándor Pongor Teaching staff: Stefan Grabuschnig, János Juhász Secretariat: Elisabetta Lippolis Chiara Alberti Giorgia Danelon Computer system manager: Dario Palmisano Diego Soldano Trieste, 26-30 June, 2017