* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download document 2150
Photosynthetic reaction centre wikipedia , lookup
Interactome wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Point mutation wikipedia , lookup
Drug design wikipedia , lookup
Peptide synthesis wikipedia , lookup
Genetic code wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Western blot wikipedia , lookup
Metalloprotein wikipedia , lookup
Enzyme inhibitor wikipedia , lookup
Protein structure prediction wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Biosynthesis wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Biochemistry wikipedia , lookup
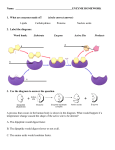
MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel 7.013 Solutions to Enzymes/Protein Structure a) i) leu-thr-phe-ala-ser (unchanged) GENERIC ENZYME: ii) trp-tyr-lys + ala-phe iii) 2 lys + 2 arg Schematics of enzymes: (side chains shown in black) ACTIVE SITE - peptide bond cleavage gly gly gly gly ile - asp gly protease A val phe protease C protease B RECOGNITION POCKET specific substrate regognition Schematics of substrates: glycine, alanine phenylalanine, tryptophan, tyrosine lysine, arginine peptide backbone + small long & + charged large & hydrophobic b) Matching them up: protease A - large open pocket. Could be Lys/Arg or Phe/Trp/Tyr. protease B - large open pocket with (-) charge at bottom. Therefore, Lys/Arg, which means that protease A must cut after Phe/Trp/Tyr protease C - small pocket. Gly, Ala. c) Change the asp in the bottom of the pocket in protease B to a lys or arg. d) It might still bind lys or arg, but if the space in the pocket were constrained, there might not be enough room because glu is longer than asp. e) The others are required to hold the essential ones in place. The also provide the correct folding to bring these amino acids in close proximity and orientation. S S GLY ALA VAL LEU ILE SER THR ASP ASN GLU GLN LYS ARG HIS TRP PHE TYR PRO CYS MET G A V L I S T D N E Q K Figure by MIT OCW. R H W F Y P C M AA sidechains, 7.013 CG 2004, 1 f) Protease A is a protein, therefore other protease A molecules can cleave it and thereby inactivate it. Having casein around decreases the chance that a protease A molecule will cleave a protease A molecule, because it will be more likely to cleave the more numerous casein molecules. g) The protease enzyme must be able to bind to the target amino acids. If they are buried inside the target protein, the protease can't "see" them and therefore can't cut at them. Eventually, the structure of the target protein gets so broken down that the inside amino acids are exposed to the protease. h) To protect protease A from degradation by other protease A molecules, bury all the phe/trp/tyr inside the protein so that they cannot be recognized and cleaved. i) The graph should show an exponential rise – the more enzyme, the faster enzyme will be produced. Eventually, the enzyme will be degraded as in part (e). This is sketched below. activ ity tim e AA sidechains, 7.013 CG 2004, 2