* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download LOF1 and Interacting Transcription Factors in Plant Development
Protein adsorption wikipedia , lookup
Synthetic biology wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Histone acetylation and deacetylation wikipedia , lookup
Eukaryotic transcription wikipedia , lookup
Gene expression wikipedia , lookup
Expression vector wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Gene regulatory network wikipedia , lookup
List of types of proteins wikipedia , lookup
RNA polymerase II holoenzyme wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Paracrine signalling wikipedia , lookup
Transcription factor wikipedia , lookup
Two-hybrid screening wikipedia , lookup
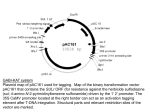
Student Publications Student Scholarship Summer 2016 LOF1 and Interacting Transcription Factors in Plant Development and Crop Yield Zoe Yeoh '18, Gettysburg College Elizabeth Luscher, University of California, Riverside Patricia Springer, University of California, Riverside Follow this and additional works at: http://cupola.gettysburg.edu/student_scholarship Part of the Biology Commons, Cell Biology Commons, and the Plant Biology Commons Share feedback about the accessibility of this item. Yeoh, Zoe; Luscher, Elizabeth; and Springer, Patricia, "LOF1 and Interacting Transcription Factors in Plant Development and Crop Yield" (2016). Student Publications. 506. http://cupola.gettysburg.edu/student_scholarship/506 This is the author's version of the work. This publication appears in Gettysburg College's institutional repository by permission of the copyright owner for personal use, not for redistribution. Cupola permanent link: http://cupola.gettysburg.edu/student_scholarship/ 506 This open access poster is brought to you by The Cupola: Scholarship at Gettysburg College. It has been accepted for inclusion by an authorized administrator of The Cupola. For more information, please contact [email protected]. LOF1 and Interacting Transcription Factors in Plant Development and Crop Yield Abstract Transcription factors (TFs) help ensure proper gene expression in developing tissues, and thus play a role in plant development and plant architecture. LATERAL ORGAN FUSION1, or LOF1, is a TF expressed in the organ boundaries of Arabidopsis thaliana. lof1 mutants have fused axillary branches and cauline leaves, which indicates importance in boundary development. Because transcription factors are known to act in complexes, we wanted to discover what other proteins interact with LOF1. We executed a yeast-2-hybrid (Y2H) screen that identified several TFs as potential interactors: WHIRLY 3 (WHY3), MYB DOMAIN PROTEIN32 (MYB32), HOMEOBOX-LEUCINE ZIPPER PROTEIN4 (HB4), and LIGHT RESPONSE BTB2 (LRB2). WHIRLY1 (WHY1) and HOMEOBOX ARABIDOPSIS THALIANA3 (HAT3) are thought to be redundant with WHY3 and HB4, respectively, and are included in our study. To gain evidence that the interactions between the potential protein interactors and LOF1 is biologically relevant in planta, we will characterize T-DNA insertion lines in which the genes that encode these interactors are disrupted. Our goal is three-fold: genotype the T-DNA lines to identify homozygous mutants; characterize the phenotypes of these mutants and compare to known phenotypes; and create double- and triple-mutants between lof1 and the other TFs. Because the boundary region is involved in determining leaf angle and leaf angle affects planting density, changes in leaf angle have the potential to impact crop yield. In the future, we may be able to apply the knowledge we obtain in the model plant Arabidopsis thaliana to crop species in order to improve crop yield. Keywords molecular biology, plant research, PCR, LOF1, transcription factors Disciplines Biology | Cell Biology | Plant Biology Comments Research funded by the National Science Foundation REU Program. Presented at the 2017 Emerging Researchers National Conference in Washington, D.C., the 2016 University of California-Riverside Center for Plant Cell Biology Research Experience for Undergraduates Symposium, and at the Undergraduate Research on the Cycle (UROC) during Gettysburg College's Year of Food (2016-2017). This poster is available at The Cupola: Scholarship at Gettysburg College: http://cupola.gettysburg.edu/student_scholarship/506 LOF1 and Interacting Transcription Factors in Plant Development and Crop Yield Zoe C. Yeoh1, Elizabeth Luscher2, Patricia Springer2 1Department of 2Center for Biology, Gettysburg College, Gettysburg, PA Plant and Cell Biology, University of California, Riverside, CA Genotyping T-DNA Insertion Lines Introduction Transcription factors (TFs) help ensure proper gene expression in developing tissues, playing a role in plant development and plant architecture. LATERAL ORGAN FUSION1, or LOF1, is a TF expressed in organ boundaries of Arabidopsis thaliana. Because TFs are known to act in complexes, we want to discover what other proteins interact with LOF1. A yeast-2-hybrid (Y2H) screen identified several potential protein interactors (listed below). To explore the role of these proteins in LOF1-related processes in planta, we obtained T-DNA insertion lines of the genes that encode these proteins. Because the boundary region is involved in determining leaf angle and leaf angle affects planting density, understanding how boundary proteins interact to affect leaf angle has the potential to improve crop yield. lof1 mutant phenotype lof1 mutants have fusion between the axillary branch and cauline leaf and abnormal boundary cell division and growth. A T-DNA Primer Plants were genotyped using Polymerase Chain Reaction (PCR) to identify homozygous mutants. 1. Gene-specific primers amplify wild-type product if wild-type allele is present; T-DNA primer and one gene-specific primer amplify T-DNA product if insertion allele is present. 2. Comparison of amplified PCR products from both reactions indicates the genotype of the plants tested (example). T-DNA Insertion Reverse Primer Forward Primer hat3_SALK_105877 plants 7-12 Wild-type primers T-DNA Primers •HOMEOBOX ARABIDOPSIS THALIANA 3 (HAT3) hat3-1 hat3-3 hat3-2 - 152* - 36** 12** 305* hb4-1 - 136* 1125** Create double- and triplemutants between LOF1 and other TFs (future) •LIGHT RESPONSE BTB2 (LRB2) lrb2-1 lrb2-2 1895** 2220* •MYB DOMAIN PROTEIN 32 (MYB32) Lee and Springer 2009 myb32 Interacting Transcription Factors • TFs identified in the Yeast-TwoHybrid (blue). TFs that are thought to be genetically redundant with the Y2H identified TFs (black) -WHIRLY 3 (WHY3) and WHIRLY 1 (WHY1) -MYB DOMAIN PROTEIN 32 (MYB32) -HOMEOBOX-LEUCINE ZIPPER PROTEIN 4 (HB4) and HOMEOBOX ARABIDOPSIS THALIANA3 (HAT3) -LIGHT RESPONSE BTB 2 (LRB2) •WHIRLY 1 (WHY1) why1-3 95* Characterize transcript levels using RT-PCR Confirm insertion sites 894** lof1-1 Genotype to identify homozygous mutants Characterize organ fusion; compare to known phenotypes D Wild type Obtain Salk Institute T-DNA insertion lines hb4-2 671** C Experimental Design •HOMEOBOX-LEUCINE ZIPPER PROTEIN 4 (HB4) lrb2-3 lof1-1 Intensity of band correlates to the level of mRNA transcript accumulated. Lack of a band from lrb2-2 suggests that it is likely a null allele. ACTIN Hetero- Homozygous Wild mutant type zygote •Insertions sites were confirmed via sequencing of PCR products containing genomic DNA flanking left border. hat3-4 Semi-quantitative RT-PCR was used to determined the level of transcript in homozygous mutants. LRB2 T-DNA Insertion Site Identification B Wild type Gene Transcript levels in homozygous mutants why1-2 121* Future Directions •Create double and triple mutants between lof1 and the homozygous TDNA mutants identified in this study •Resulting phenotypes will give support for LOF1-interacting TFs’ involvement in LOF1 processes •Gain a better understanding of how LOF1 functions •May be able to apply this knowledge to crop species to improve crop yield Acknowledgements •WHIRLY 3 (WHY3) why3 This work was supported by the National Science Foundation REU grant 1461297 to the UC-Riverside Center for Plant Cell Biology. -94* *Insertion sites were sequenced. **Insertion sites were identified through TAIR. Figures are not drawn to scale. Numbers correspond to bp, beginning from transcription start site. Figure Legend: = Exon = Intron = 5’ UTR References: Lee, D. K., M. Geisler and P. S. Springer (2009). Development 136(14): 2423-2432. Contact Info: = 3’ UTR = T-DNA Insertion Site Zoe Yeoh, [email protected] 300 North Washington Street, Gettysburg, PA 17325