* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Macromolecular Interaction
Protein folding wikipedia , lookup
Protein structure prediction wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Protein purification wikipedia , lookup
Protein mass spectrometry wikipedia , lookup
Western blot wikipedia , lookup
List of types of proteins wikipedia , lookup
Intrinsically disordered proteins wikipedia , lookup
Bimolecular fluorescence complementation wikipedia , lookup
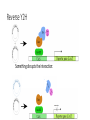
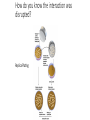
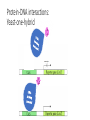
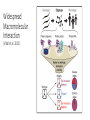
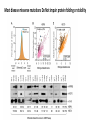
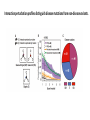
MacromolecularInteraction AdvancedGenetic YangLi Feb.272017 Proteinsformcomplexes/interactwithother macromolecules Howcanyouidentifyprotein-protein interactions? Howdoyouidentifyprotein-protein interactions? Biochemical/biophysicalmethods • Affinitypurification– massspectrometry • Co-immunoprecipitation • Proteinmicroarrays • Förster resonanceenergytransfer Genetic/molecularmethods • Yeast-twohybridsystem • Split-ubiquitinsystem • Bacterial-two-hybridsystem Y2H:usingthesplitGal4-UASsystemtogo “fishing”forprotein-proteininteractions ThisishowGal4UASnormallyworks: UAS=Upstreamactivationsequence Gal4=atranscriptionfactormadeupofaDNAbindingandanactivation domain TheDNAbindingdomainandactivation domainsofGAL-4canbeseparated TheDNAbindingdomain(BD)alonecannotactivate transcription The“bait”(i.e.yourproteinofinterest)isfusedtothe BD TheDNAbindingdomainandactivation domainsofGAL-4canbeseparated Theactivationdomain(AD)alonecannotbindtheUAS The“prey”isfusedtotheAD Y2H:usingthesplitGal4-UASsystemtogo “fishing”forprotein-proteininteractions Ifthe“prey”bindstothe“bait”theADandBDare broughttogetherandactivatetranscriptionofthe reporter andaninteractionisdiscovered. Y2H:usingthesplitGal4-UASsystemtogo “fishing”forprotein-proteininteractions Testmanydifferent“prey” e.g.cDNAlibrary CreatingacDNAlibrary Reporters • Auxotroph->prototrophselection • Aminoacidbiosynthesis(His,Leu,Ade,Ura) • Nucleicacidbiosynthesis • Colordetectionscreen • LacZ:blue/whitescreen • GFP Yeasttwo-hybridassayofdifferentLexAfusionproteins(baits)withvariousPDZ domains(preys) HostOrganism • Yeast • • • • • Eukaryotic Hospitableinternalenvironment Genomesequenced,techniquestomanipulate Onlyabletodetectinteractionsinthenucleus Someproteinsaretoxictoyeast • E.coli • Highertransformationefficiency • Donotneedanuclearlocalizationsignal • Canusestrainswithoutinterferingmethyltransferase activity • MammalianCells BenefitsofY2H • Candetecttransientinteractionsnotfoundinco-IP • Semi-quantitative(2colorsystem) • Canbeusedtostudyknowninteractions • Modifyspecificresidues,observewhetherinteractionismaintained • Immediateidentificationofthegenethatencodestheproduct WeaknessesofY2H • Falsepositives • Proteinsmaynotbeexpressedtogetherinreality • Unnaturalconcentrations • Non-specificinteractions • Falsenegatives • Interactionsmustoccurinthenucleus(andproteinsmustbesoluble)tobe detected • FusiontoADorBDmayblockproteininteraction • Yeastmaylackchaperonesforproperfolding RelatedScreens • Reverseyeast-twohybrid • Detectwhenaninteractionisdisrupted • Yeastthree-hybrid • DetectProtein-RNAinteractions • Yeastone-hybrid • DetectProtein-DNAinteractions ReverseY2H Somethingdisruptstheinteraction: 1 Howdoyouknowtheinteractionwas disrupted? ReplicaPlating Howdoyouknowtheinteractionwas disrupted? URA3geneand5-fluoro-oroticacid(5-FOA) Yeastsurviveswhenproteininteractionisdisrupted Protein-RNAinteractions: Yeastthree-hybrid Gal4BD-Protein1-RNA-Protein2-Gal4AD Protein-DNAinteractions: Yeast-one-hybrid Widespread Macromolecular Interaction (Vidaletal.2015) Experimental Workflow LUMIERAssay MostdiseasemissensemutationsDoNotimpairproteinfoldingorstability Interactionperturbationprofilesdistinguishdiseasemutationsfromnon-diseasevariants. Protein-DNAinteractionperturbation