* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Oxidative Stress and Repair
Zinc finger nuclease wikipedia , lookup
Protein mass spectrometry wikipedia , lookup
Protein purification wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Protein structure prediction wikipedia , lookup
Protein–protein interaction wikipedia , lookup
List of types of proteins wikipedia , lookup
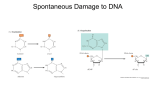
Oxidative Stress and Repair • Oxidative DNA modifications Faiez Al Nimer • Oxidative lipid modifications Huang Liyue • Oxidative protein modifications Sanyal Nikhilesh • Oxidative DNA and protein damage repair Melanie Neely Willis Oxidative DNA damage Both nuclear and mitochondrial DNA is attacked by ROS but mtDNA is affected more because it does not contain histones it is in proximity to ROS generation and has limited antioxidant repair system Strand breaks Reactive oxygen species Oxidized base adducts DNA-DNA and DNA-protein cross links Oxidized base adducts: Hydroxyl radical Evans et al, Reviews in mutation research, 2004 8-OH-Gua and Thymin glycol Guanine: lowest oxidation potential most oxidized base Cell viability and disease For 8-OH-Gua: Results in C→A and G→T transversions, leading to mutations Affects transcription factor binding and gene expression Interferes with methylation of i.e. oncogenes For thymin glycol: T→C transitions have been noted Blocks replication DNA oxidation: Neurodegenerative diseases 8-OHDG staining p62 promoter oxidation Lovell et al, J Neurochem, 1999 Du et al, Neurobiol of disease,2009 DNA oxidation: Neurodegenerative diseases p62 expression Absence of p62 induces cell death p62 protects cells from oxidative stress via the antioxidant response element (ARE) Du et al, Neurobiol of disease,2009 Oxidative Stress and Repair • Oxidative DNA modifications Faiez Al Nimer • Oxidative lipid modifications Huang Liyue • Oxidative protein modifications Sanyal Nikhilesh • Oxidative DNA and protein damage repair Melanie Neely Willis Lipid modifications by reactive oxygen species (ROS) • Overproduction of ROS damages cellular components, including lipids, leading to decline in physiological function and cell death Reaction of ROS with lipids lead to • Lipid oxidation • Lipid peroxidation Lipid Oxidation • Initiation – A radical is formed • Propagation – Radicals react and transfer their unpaired electron to other compounds • Termination – Two radicals combine to stop the reaction Adibhatla RM; Hatcher JF. Antioxid Redox Signal. 2010 Lipid peroxidation Initiation LH OH or M=O L L + O2 Propagation LOO + LH LOOH + metal Commonly measured decomposition products LOO L + LOOH Hydroperoxide Alkoxy radical LO LOO Peroxy radical Alkanes Malondialdehyde 4 hydroxynananol 8-isoprostanes Adibhatla RM; Hatcher JF. Antioxid Redox Signal. 2010 Oxidative Stress and Repair • Oxidative DNA modifications Faiez Al Nimer • Oxidative lipid modifications Huang Liyue • Oxidative protein modifications Sanyal Nikhilesh • Oxidative DNA and protein damage repair Melanie Neely Willis Effects of Reactive oxygen species on Protein backbone • Oxidation of proteins often targets them for degradation (20S proteasome and other proteases). • Several age-related disorders are characterised by Protein oxidation. E.g. Parkinson’s disease, Amyotropic Lateral sclerosis. • Product of the oxidation (Peptide alkoxyl radical) is susceptible to cleavage by diamide or a-amidation pathways. Redox Biochemistry, Wiley Interscience, 2008; p186 Oxidation of amino acid residue side chain Redox Biochemistry, Wiley Interscience, 2008; p188 Oxidation of amino acid residue side chains : Tyrosine • Tyrosine oxidation can lead to dityrosine formation. • This leads to protein crosslinking. • Useful Biomarker for Oxidative damage and aging. Tyrosine Dityrosine Larios J M et al. J. Biol. Chem. 2001;276:17437-17441 • Increased levels of dityrosine is found in rat hearts where Matrix metalloproteinase-2 (MMP-2) was induced by cytokines. Qun Gao C et al. Cardiovasc Res 2003;57:426-433 Oxidation of amino acid residue side chains : Cysteine Sulfonic acid Sulfinic acid Sulfenic acid R’ - Glutathione or another protein Green J and Paget MS (2004) Nat Rev Microbiol. 2(12):954-66 Beneficial effects of ROS on Proteins : e.g. DJ-1 H2O2 oxidises the Cys106 (-SH) to sulfinic acid (-SOOH) Cys106 oxidation is controlled by its neighbouring environment. Oxidised Cys106 Partially oxidised Cys106 (Sulfenic acid) Blackinton et al et al. J. Biol. Chem. 2009;284:6476-6485 Beneficial effects of ROS on Proteins : e.g. DJ-1 Cys106 is essential for maintaining nuclear morphology. Shown here are M17neuroblastoma cells Cys106 helps increase Cell viability under mitochondrial stress Blackinton et al et al. J. Biol. Chem. 2009;284:6476-6485 Oxidative Stress and Repair • Oxidative DNA modifications Faiez Al Nimer • Oxidative lipid modifications Huang Liyue • Oxidative protein modifications Sanyal Nikhilesh • Oxidative DNA and protein damage repair Melanie Neely Willis DNA & Protein Repair Enzymes Oxidation of pyrimidine and purine bases leads to DNA damage. Oxidation of methionine and cysteine residues can lead to protein damage and aggregation. DNA damage is repaired by several enzymes in the Base Excision Repair pathway (BER). Proteins are repaired by methionine sulfoxide reductases (MsrA, MsrB) and sulfiredoxins. Base Excision Repair 1. Removal of the damaged base from the sugar by a damage-specific glycosylase 2. Processing of the apyrmidinic-apurinic (AP) site by an AP endonuclease and deoxyribosylphosphate lyase. 3. Replacement of an undamaged nucleotide by DNA polymerase. 4. Repairing nicks in the DNA backbone by DNA ligase. Crystal Structure Shows DNA Bending Lukianova OA, David SS. A role for iron-sulfur clusters in DNA repair. Curr Opin Chem Biol. 2005;9:145–151. Reversibility of Prx Hyperoxidation Before exposure After 10 minutes exposure to H202 Recovery for 4 hours 10 more minutes of exposure to H202 Recovery for 8 hours Woo HA, Chae HZ, Hwang SC, Yang KS, Kang SW, Kim K, Rhee SG.Science. 2003 Apr 25;300(5619):592-4. Mechanism of reduction of Prx-SO2− by Srx. Jönsson T J et al. J. Biol. Chem. 2009;284:33305-33310