* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Supplementary Figure Legends
Minimal genome wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Epigenetics in stem-cell differentiation wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Gene expression profiling wikipedia , lookup
Oncogenomics wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
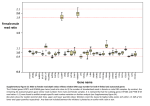
1 Supplementary Figure S1. 2 Unsupervised clustering analysis using Firehose of RNA-seq data from TCGA breast 3 cancer transcriptomes, identifying 7 sample clusters. 4 5 Supplementary Figure S2. 6 (A) GO analysis (by DAVID) of the 100 genes whose expression is most highly correlated 7 with that of Ki-67, as determined by COEXPRESdb. 8 (B) Correlation between cyclin-annotated genes and Ki-67 across colorectal cancer 9 subtypes. 10 (C) Histogram of the number of Ki-67 significantly correlated genes in TCGA breast 11 tumours with respect to Spearman correlation values (all correlation adjusted P < 0.05). 12 The upper right bar plots report the proportion and number of cell cycle genes among 13 genes above a certain correlation coefficient. 14 (D) Correlation between cancer-annotated genes and Ki-67 across TCGA breast cancer 15 subtypes. 16 17 Supplementary Figure S3. Cell cycle dynamics of Ki-67 expression are similar in 18 non-transformed cells and cancer cell lines. 19 A. Immunofluorescence analysis of Ki-67 expression and localisation in asynchronous 20 individual human fibroblasts at different cell cycle phases defined using cell cycle markers 21 cyclin D1, cyclin E1 and cyclin B1. 22 23 B. Left: Graphs showing Ki-67 immunofluroescence staining intensity at different cell cycle 24 stages in asynchronous individual cells, in HDF, U2OS and HCT-116 cells. Pixel intensity 25 was quantified using ImageJ software. The legend shows the criteria for attributing cell 26 cycle phases A-F (Sen: Senescent). Error bars show standard deviation. 50 cells were 1 27 analysed in each case. Right: Example source data for quantification of cell cycle 28 regulation of Ki-67 expression in asynchronous cells, showing a panel of HDF. Top: 29 Hoechst staining. Cells quantified are indicated by red lables corresponding to the cell 30 cycle position, according to the legend, Top left. Middle: Cyclin A immunofluorescence; 31 Bottom: Ki-67 immunofluorescence. 32 33 Supplementary Figure S4. Cell cycle arrest by serum starvation or DNA damage is 34 complete. 35 Analysis of DNA replication by incorporation of EdU following 72 hours of the indicated 36 treatments; cells were labelled with EdU for 24 hours. Gate shows percentage of cells 37 showing any level of incorporation of EdU. 38 39 Supplementary Figure S5. 40 Time course of senescence induction by bleomycin, as assessed by beta-galactosidase 41 assay. Bar, 10μm 42 43 Supplementary Figure S6. 44 (A) Correlation between drug response and Ki-67, CCNA2, CCNB2, CCND1, and CCND3 45 expression across all CCLE cell lines and drugs; smoothing was applied. Number is the 46 correlation coefficient between expression of each gene and drug sensitivity. 47 (B) Same as (A) but restricted to Irinotecan; no smoothing. 48 (C) Same as (A) but restricted to Topotecan; no smoothing. 49 2