Michael T. Woodside “OBSERVING THE FOLDING AND MISFOLDING OF SINGLE PROTEIN
... prion protein molecules that allow us to follow the change in structure of the protein as it folds in real time, by applying tension across the protein with optical tweezers. The prion protein is responsible for "mad cow" disease, through the action of an incorrectly folded structure that is infecti ...
... prion protein molecules that allow us to follow the change in structure of the protein as it folds in real time, by applying tension across the protein with optical tweezers. The prion protein is responsible for "mad cow" disease, through the action of an incorrectly folded structure that is infecti ...
S4. Computational Molecular Modeling- Pre
... DNA can cause an abnormal phenotype. Terms/phrases: DNA mutation, normal allele, mutant allele, gene, primary protein structure, secondary protein structure, tertiary protein structure, transcription, translation, protein function, normal phenotype, mutant phenotype, protein function. Part 2: Inform ...
... DNA can cause an abnormal phenotype. Terms/phrases: DNA mutation, normal allele, mutant allele, gene, primary protein structure, secondary protein structure, tertiary protein structure, transcription, translation, protein function, normal phenotype, mutant phenotype, protein function. Part 2: Inform ...
Introduction to databases
... pattern results in terms of predicted function. Explain why these small motifs are so evolutionarily conserved that they can be used to predict what a protein’s function is? ...
... pattern results in terms of predicted function. Explain why these small motifs are so evolutionarily conserved that they can be used to predict what a protein’s function is? ...
Proteomics techniques used to identify proteins
... Identification of regulatory proteins from human cells using 2D-GE and LC-MS/MS Victor Paromov Christian Muenyi William L. Stone ...
... Identification of regulatory proteins from human cells using 2D-GE and LC-MS/MS Victor Paromov Christian Muenyi William L. Stone ...
LOYOLA COLLEGE (AUTONOMOUS), CHENNAI – 600 034
... 6. Luciferase is used to degrade the excess nucleotide bases. 7. Operons are not found in prokaryotes. 8. Genes are upregulated in cancers. 9. Nitrosylation is very important for cell cycle progression. 10. Metabolomics and metagenomics are same. III Complete the following ...
... 6. Luciferase is used to degrade the excess nucleotide bases. 7. Operons are not found in prokaryotes. 8. Genes are upregulated in cancers. 9. Nitrosylation is very important for cell cycle progression. 10. Metabolomics and metagenomics are same. III Complete the following ...
Carbon Based Compounds
... Lipids are a diverse group of hydrophobic (water fearing) molecules Proteins have many structures, resulting in a wide range of functions ...
... Lipids are a diverse group of hydrophobic (water fearing) molecules Proteins have many structures, resulting in a wide range of functions ...
Notes
... – arranged in specific sequence – linked by PEPTIDE BONDS – range in length from a few to 1000+ ...
... – arranged in specific sequence – linked by PEPTIDE BONDS – range in length from a few to 1000+ ...
Chapter 6 questions
... 1. Identify the body's working proteins. 2. Identify the body's structural proteins. 3. What do proteins contain that carbohydrates and lipids do not? 4. _______________ are the building blocks of proteins. 5. What is an essential amino acid? How many are there? 6. What are proteins made of? Illustr ...
... 1. Identify the body's working proteins. 2. Identify the body's structural proteins. 3. What do proteins contain that carbohydrates and lipids do not? 4. _______________ are the building blocks of proteins. 5. What is an essential amino acid? How many are there? 6. What are proteins made of? Illustr ...
Chapter 6: Protein 1. Identify the body's working proteins.
... 3. What do proteins contain that carbohydrates and lipids do not? 4. _______________ are the building blocks of proteins. 5. What is an essential amino acid? How many are there? 6. What are proteins made of? Illustrate an example. 7. Globular shaped proteins are __________ proteins and are _________ ...
... 3. What do proteins contain that carbohydrates and lipids do not? 4. _______________ are the building blocks of proteins. 5. What is an essential amino acid? How many are there? 6. What are proteins made of? Illustrate an example. 7. Globular shaped proteins are __________ proteins and are _________ ...
Interaction Site Evolution
... COMPUTER SCIENCE - Interaction Site Evolution DNA is the blueprint for generating strings of amino acids which fold into proteins. The interactions these proteins form with each other are primary components of organismal physiology. Proteins assume very specific shapes, and the amino acids on their ...
... COMPUTER SCIENCE - Interaction Site Evolution DNA is the blueprint for generating strings of amino acids which fold into proteins. The interactions these proteins form with each other are primary components of organismal physiology. Proteins assume very specific shapes, and the amino acids on their ...
Abstract
... maps for many protein domains. Inferred contacts by mfDCA can be utilized as a reliable guide in high accuracy computational predictions of domain structure. Our results capture clear signals beyond intradomain residue contacts, for instance, interdomain interactions in macro molecular assemblies an ...
... maps for many protein domains. Inferred contacts by mfDCA can be utilized as a reliable guide in high accuracy computational predictions of domain structure. Our results capture clear signals beyond intradomain residue contacts, for instance, interdomain interactions in macro molecular assemblies an ...
Proteins for Growth and Repair
... Complete Proteins from Animal Foods: Complete proteins contain 9 of the 22 amino acids that are essential to life and must be added to the diet. They are found in animal foods like meat, fish, milk cheese and eggs. Vegans and vegetarians can get shortages of the essential amino acids lysine and thre ...
... Complete Proteins from Animal Foods: Complete proteins contain 9 of the 22 amino acids that are essential to life and must be added to the diet. They are found in animal foods like meat, fish, milk cheese and eggs. Vegans and vegetarians can get shortages of the essential amino acids lysine and thre ...
ProteinChipâ technology is one of the most exciting advancements
... Ciphergen Biosystems, Inc., Fremont, CA, USA ProteinChip technology is one of the most exciting advancements in protein analysis in the last 5 years. The Protein Biology SystemTM (PBS) combines the power of mass analysis with chromatography surfaces on an integrated platform. The PBS can easily be ...
... Ciphergen Biosystems, Inc., Fremont, CA, USA ProteinChip technology is one of the most exciting advancements in protein analysis in the last 5 years. The Protein Biology SystemTM (PBS) combines the power of mass analysis with chromatography surfaces on an integrated platform. The PBS can easily be ...
IB2.14.3 Building a protein
... All the basic structural material of the human body is made of proteins. Skin, muscles, bone, cartilage, ligaments and cell membranes all contain a lot of protein. In addition, other proteins do important jobs in cells. All protein molecules contain the elements: Carbon Oxygen Hydrogen Nitro ...
... All the basic structural material of the human body is made of proteins. Skin, muscles, bone, cartilage, ligaments and cell membranes all contain a lot of protein. In addition, other proteins do important jobs in cells. All protein molecules contain the elements: Carbon Oxygen Hydrogen Nitro ...
Daniel Kaganovich Molecular Mechanism of
... folding quality control system, which includes chaperones that enhance protein folding and regulate protein aggregation. From basic findings in simple cellular models, we develop animal models of neural function and neurodegenerative disease. Our goal is to understand some of the ways in which neuro ...
... folding quality control system, which includes chaperones that enhance protein folding and regulate protein aggregation. From basic findings in simple cellular models, we develop animal models of neural function and neurodegenerative disease. Our goal is to understand some of the ways in which neuro ...
Amino acid sequence fingerprints in divergent evolution of
... are preserved, the resulting protein function and activity can remain without any observable modifications. The essential amino acid residues are mostly parts of the so-called conserved sequence regions that cover the isolated segments belonging to the active site of the protein. In the case the pro ...
... are preserved, the resulting protein function and activity can remain without any observable modifications. The essential amino acid residues are mostly parts of the so-called conserved sequence regions that cover the isolated segments belonging to the active site of the protein. In the case the pro ...
TIGR_ISS
... Visually inspect alignments, look for conserved active sites, look for (generally) at least 35% identity across the full lengths of both proteins. If matches are not full length, look to see if there are recognized functional domains in the area where the match occurs. Decide how much information ca ...
... Visually inspect alignments, look for conserved active sites, look for (generally) at least 35% identity across the full lengths of both proteins. If matches are not full length, look to see if there are recognized functional domains in the area where the match occurs. Decide how much information ca ...
Document
... Objective: To know the major steps in protein synthesis and the RNAs and proteins involved in this process. To understand the mechanism by which proteins are targeted to specific cimpartments. I. Genetic code A. Three nucleotides make one codon B. "Universal" C. Degenerate D. Commaless II. Translati ...
... Objective: To know the major steps in protein synthesis and the RNAs and proteins involved in this process. To understand the mechanism by which proteins are targeted to specific cimpartments. I. Genetic code A. Three nucleotides make one codon B. "Universal" C. Degenerate D. Commaless II. Translati ...
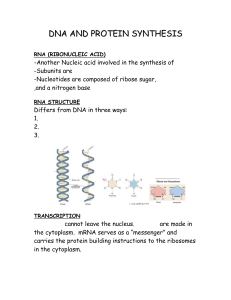
DNA AND PROTEIN SYNTHESIS
... bases in mRNA into the amino acids of a protein. 1 Codon = 3 nucleotides on mRNA 1 Codon = Some codons are redundant (can be used again to give the same amino acid) RESULT OF TRANSLATION ...
... bases in mRNA into the amino acids of a protein. 1 Codon = 3 nucleotides on mRNA 1 Codon = Some codons are redundant (can be used again to give the same amino acid) RESULT OF TRANSLATION ...
Protein: How Cows and Carrots Become People 1. Your body can
... 6. In the video, Jake compares proteins to pasta. What characteristic do they share other than the fact that we eat both proteins and pasta? ...
... 6. In the video, Jake compares proteins to pasta. What characteristic do they share other than the fact that we eat both proteins and pasta? ...
IFITM3 Peptide PRODUCT DATA SHEET Bioworld Technology CO., Ltd.
... IFITM3 (interferon induced transmembrane protein 3), also known as 1-8U or IP15, is a multi-pass membrane protein that belongs to the IFITM (interferon inducible transmembrane) family of proteins. IFITM proteins are induced by type I and type II interferons and contain multiple interferon (IFN)-stim ...
... IFITM3 (interferon induced transmembrane protein 3), also known as 1-8U or IP15, is a multi-pass membrane protein that belongs to the IFITM (interferon inducible transmembrane) family of proteins. IFITM proteins are induced by type I and type II interferons and contain multiple interferon (IFN)-stim ...
Answers to Biological Inquiry Questions – Brooker et al ARIS site
... one cell expresses a high-affinity receptor and another cell a low-affinity receptor, the two cells would respond to the signaling protein at different concentrations. Likewise, the different receptors may be linked with different second messenger molecules generated within the cell. These messenger ...
... one cell expresses a high-affinity receptor and another cell a low-affinity receptor, the two cells would respond to the signaling protein at different concentrations. Likewise, the different receptors may be linked with different second messenger molecules generated within the cell. These messenger ...
Protein: A polymer of amino acids Amino Acid Structure
... – Coiled or folded shape held together by hydrogen bonds ...
... – Coiled or folded shape held together by hydrogen bonds ...
Slide ()
... (ORFs) coding for latent proteins, reactivation proteins, and structural proteins. Host genes that help the virus evade immune surveillance and inhibit apoptosis have been acquired from chromosomes through a process of molecular piracy. These genes include vFLIP, vBcl-2, v-cyclin, interferon respons ...
... (ORFs) coding for latent proteins, reactivation proteins, and structural proteins. Host genes that help the virus evade immune surveillance and inhibit apoptosis have been acquired from chromosomes through a process of molecular piracy. These genes include vFLIP, vBcl-2, v-cyclin, interferon respons ...
Protein moonlighting

Protein moonlighting (or gene sharing) is a phenomenon by which a protein can perform more than one function. Ancestral moonlighting proteins originally possessed a single function but through evolution, acquired additional functions. Many proteins that moonlight are enzymes; others are receptors, ion channels or chaperones. The most common primary function of moonlighting proteins is enzymatic catalysis, but these enzymes have acquired secondary non-enzymatic roles. Some examples of functions of moonlighting proteins secondary to catalysis include signal transduction, transcriptional regulation, apoptosis, motility, and structural.Protein moonlighting may occur widely in nature. Protein moonlighting through gene sharing differs from the use of a single gene to generate different proteins by alternative RNA splicing, DNA rearrangement, or post-translational processing. It is also different from multifunctionality of the protein, in which the protein has multiple domains, each serving a different function. Protein moonlighting by gene sharing means that a gene may acquire and maintain a second function without gene duplication and without loss of the primary function. Such genes are under two or more entirely different selective constraints.Various techniques have been used to reveal moonlighting functions in proteins. The detection of a protein in unexpected locations within cells, cell types, or tissues may suggest that a protein has a moonlighting function. Furthermore, sequence or structure homology of a protein may be used to infer both primary function as well as secondary moonlighting functions of a protein.The most well-studied examples of gene sharing are crystallins. These proteins, when expressed at low levels in many tissues function as enzymes, but when expressed at high levels in eye tissue, become densely packed and thus form lenses. While the recognition of gene sharing is relatively recent—the term was coined in 1988, after crystallins in chickens and ducks were found to be identical to separately identified enzymes—recent studies have found many examples throughout the living world. Joram Piatigorsky has suggested that many or all proteins exhibit gene sharing to some extent, and that gene sharing is a key aspect of molecular evolution. The genes encoding crystallins must maintain sequences for catalytic function and transparency maintenance function.Inappropriate moonlighting is a contributing factor in some genetic diseases, and moonlighting provides a possible mechanism by which bacteria may become resistant to antibiotics.