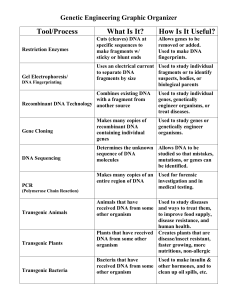
DNA Technology Tools Graphic Organizer KEY
... fragments or to identify fragments by size suspects, bodies, or biological parents ...
... fragments or to identify fragments by size suspects, bodies, or biological parents ...
Genetic Engineering Short Notes
... short specific DNA sequences and that cut the DNA there 4. Plasmid- small, circular DNA molecules that can replicate independantly of the main chromosome 5. Vector- something used to carry the gene of interest into another cell ...
... short specific DNA sequences and that cut the DNA there 4. Plasmid- small, circular DNA molecules that can replicate independantly of the main chromosome 5. Vector- something used to carry the gene of interest into another cell ...
DNA Analysis of Various Mouse Organs
... extracted DNA obtained from various mouse organs • The DNA was tagged with Ethidium bromide to illustrate the difference in DNA concentrations between organs. • Gel electrophoresis allowed for visualization of DNA from the varying organ tissues. ...
... extracted DNA obtained from various mouse organs • The DNA was tagged with Ethidium bromide to illustrate the difference in DNA concentrations between organs. • Gel electrophoresis allowed for visualization of DNA from the varying organ tissues. ...
Molecuar Structure of DNA Questions
... 2. What two parts of the DNA nucleotide make up the “backbone” of the macromolecule? ...
... 2. What two parts of the DNA nucleotide make up the “backbone” of the macromolecule? ...
Biotechnology and Gel Electrophoresis
... In DNA Fingerprinting, the DNA of an organism is cut up into fragments using restriction enzymes producing a large number of fragments of DNA Because no two individuals have identical DNA, no two individuals will have the same length fragments This technique allows us to identify families because th ...
... In DNA Fingerprinting, the DNA of an organism is cut up into fragments using restriction enzymes producing a large number of fragments of DNA Because no two individuals have identical DNA, no two individuals will have the same length fragments This technique allows us to identify families because th ...
Review Sheet NYS Regents Lab Activity #1 Relationships and Biodiversity
... pigments blue, yellow, and pink, scattered bundles, no difference in the amino acid sequences, and the same DNA banding pattern. 4. The evidence that should receive the most emphasis when determining the relatedness would be the genetic sequence, as many things can look similar structurally (converg ...
... pigments blue, yellow, and pink, scattered bundles, no difference in the amino acid sequences, and the same DNA banding pattern. 4. The evidence that should receive the most emphasis when determining the relatedness would be the genetic sequence, as many things can look similar structurally (converg ...
5echap12guidedreading
... 3. How does the rapid reproduction of bacteria make them a good choice for cloning a foreign gene? ...
... 3. How does the rapid reproduction of bacteria make them a good choice for cloning a foreign gene? ...
DNA Technology
... • A way to identify an individual based on their unique sequences of DNA • Many sequences are the same in humans • Look for regions with greatest diversity (polymorphisms) – Often are noncoding (intron) regions – Short tandem repeats (STRs) used now • 4 nucleotide sequences • Vary in length between ...
... • A way to identify an individual based on their unique sequences of DNA • Many sequences are the same in humans • Look for regions with greatest diversity (polymorphisms) – Often are noncoding (intron) regions – Short tandem repeats (STRs) used now • 4 nucleotide sequences • Vary in length between ...
Unit 4: Genetics
... 3) Elongation of DNA strand. Cycle is repeated multiple times, which results in an exponential gain of the target DNA sequence. Courtesy of Madeleine Ball ...
... 3) Elongation of DNA strand. Cycle is repeated multiple times, which results in an exponential gain of the target DNA sequence. Courtesy of Madeleine Ball ...
Parallel Computing with DNA
... Researchers also investigated how the ability to repeatedly recombine DNA strands can be used to construct “computational devices” that work massively parallel and are highly energy efficient. It is not obvious how such devices can be programmed to solve computational problems for which currently co ...
... Researchers also investigated how the ability to repeatedly recombine DNA strands can be used to construct “computational devices” that work massively parallel and are highly energy efficient. It is not obvious how such devices can be programmed to solve computational problems for which currently co ...
Genetic Engineering
... • Hundreds of useful bacterial strains have been produced • Bacteria can even digest oil ...
... • Hundreds of useful bacterial strains have been produced • Bacteria can even digest oil ...
Who am I?
... What is cloning? Clones are identical copies of living things. Humans have cloned a lot of things already. ...
... What is cloning? Clones are identical copies of living things. Humans have cloned a lot of things already. ...
Webquests_files/Genes and DNA SWQ
... The four nucleotides Difference between dominant and recessive alleles ...
... The four nucleotides Difference between dominant and recessive alleles ...
Name - OnCourse
... 3. The “backbones” of the DNA molecule is made up of two components, what are these? c. _______________________________ d. _______________________________ 5. There are four different bases that make up the “rungs.” What are the names of those bases? a. _______________________________ b. ____________ ...
... 3. The “backbones” of the DNA molecule is made up of two components, what are these? c. _______________________________ d. _______________________________ 5. There are four different bases that make up the “rungs.” What are the names of those bases? a. _______________________________ b. ____________ ...
DNA Test Review
... 3. If a DNA molecule has the sequence TACGAACCC, what would be the complimentary mRNA sequence? 4. The process by which a DNA molecule is copied is called _____. 5. What is a codon? 6. What are the types of RNA? 7. Messenger RNA is formed in the process of _____. 8. What happens during translation a ...
... 3. If a DNA molecule has the sequence TACGAACCC, what would be the complimentary mRNA sequence? 4. The process by which a DNA molecule is copied is called _____. 5. What is a codon? 6. What are the types of RNA? 7. Messenger RNA is formed in the process of _____. 8. What happens during translation a ...
DNA TECHNOLOGY
... DNA is broken up (restriction enzyme) & sorted by size using gel electrophoresis ...
... DNA is broken up (restriction enzyme) & sorted by size using gel electrophoresis ...
analysis
... b) Each of the four reaction mixtures will have a different dideoxynucleotide (ddGTP, ddATP, ddCTP, or ddTTP) C. Electrophoresis 1. Denature the DNA before electrophoresis 2. Each reaction mixture will be electrophoresed in a separate lane through a ...
... b) Each of the four reaction mixtures will have a different dideoxynucleotide (ddGTP, ddATP, ddCTP, or ddTTP) C. Electrophoresis 1. Denature the DNA before electrophoresis 2. Each reaction mixture will be electrophoresed in a separate lane through a ...
Aim # 29: NYS Lab Relationships and
... pink, scattered bundles, no difference in the amino acid sequences, and the same DNA banding pattern. 4. The evidence that should receive the most emphasis when determining the relatedness would be the genetic sequence, as many things can look similar structurally (convergent evolution), but would b ...
... pink, scattered bundles, no difference in the amino acid sequences, and the same DNA banding pattern. 4. The evidence that should receive the most emphasis when determining the relatedness would be the genetic sequence, as many things can look similar structurally (convergent evolution), but would b ...
Gel electrophoresis of nucleic acids

Nucleic acid electrophoresis is an analytical technique used to separate DNA or RNA fragments by size and reactivity. Nucleic acid molecules which are to be analyzed are set upon a viscous medium, the gel, where an electric field induces the nucleic acids to migrate toward the anode, due to the net negative charge of the sugar-phosphate backbone of the nucleic acid chain. The separation of these fragments is accomplished by exploiting the mobilities with which different sized molecules are able to pass through the gel. Longer molecules migrate more slowly because they experience more resistance within the gel. Because the size of the molecule affects its mobility, smaller fragments end up nearer to the anode than longer ones in a given period. After some time, the voltage is removed and the fragmentation gradient is analyzed. For larger separations between similar sized fragments, either the voltage or run time can be increased. Extended runs across a low voltage gel yield the most accurate resolution. Voltage is, however, not the sole factor in determining electrophoresis of nucleic acids.The nucleic acid to be separated can be prepared in several ways before separation by electrophoresis. In the case of large DNA molecules, the DNA is frequently cut into smaller fragments using a DNA restriction endonuclease (or restriction enzyme). In other instances, such as PCR amplified samples, enzymes present in the sample that might affect the separation of the molecules are removed through various means before analysis. Once the nucleic acid is properly prepared, the samples of the nucleic acid solution are placed in the wells of the gel and a voltage is applied across the gel for a specified amount of time.The DNA fragments of different lengths are visualized using a fluorescent dye specific for DNA, such as ethidium bromide. The gel shows bands corresponding to different nucleic acid molecules populations with different molecular weight. Fragment size is usually reported in ""nucleotides"", ""base pairs"" or ""kb"" (for thousands of base pairs) depending upon whether single- or double-stranded nucleic acid has been separated. Fragment size determination is typically done by comparison to commercially available DNA markers containing linear DNA fragments of known length.The types of gel most commonly used for nucleic acid electrophoresis are agarose (for relatively long DNA molecules) and polyacrylamide (for high resolution of short DNA molecules, for example in DNA sequencing). Gels have conventionally been run in a ""slab"" format such as that shown in the figure, but capillary electrophoresis has become important for applications such as high-throughput DNA sequencing. Electrophoresis techniques used in the assessment of DNA damage include alkaline gel electrophoresis and pulsed field gel electrophoresis.For short DNA segments such as 20 to 60 bp double stranded DNA, running them in Polyacrylamide gel (PAGE) will give better resolution(native condition). Similarly, RNA and single stranded DNA can be run and visualised by PAGE gels containing denaturing agents such as Urea. PAGE gels are widely used in techniques such as DNA foot printing, EMSA and other DNA-protein interaction techniques.The measurement and analysis are mostly done with a specialized gel analysis software. Capillary electrophoresis results are typically displayed in a trace view called an electropherogram.